Exploring the effect of Ru(II) arene complexes on cytotoxicity upon co-ligand variation and loading on amine-functionalized silica nanoparticles†
Srijita
Pal
,
Pragti
,
Amardeep
Kumar
and
Suman
Mukhopadhyay
 *
*
Department of Chemistry, School of Basic Sciences, Indian Institute of Technology Indore, Simrol, Khandwa Road, Indore, 453552, India. E-mail: suman@iiti.ac.in; Tel: +91 731660 3328
First published on 31st March 2025
Abstract
To overcome the undesirable side effects and acquired resistance associated with platinum-based chemotherapeutics, scientists are searching for alternative strategies involving novel metal-based compounds with improved pharmacological properties. Ruthenium complexes have emerged as prospective candidates to combat side effects and improve the selectivity of anticancer agents. In this work, a benzimidazole-based chelating ligand, HL (4-(1H-naphth[2,3-d]imidazol-2-yl)-1,3-benzenediol) with O and N donor atoms, was synthesized and used for complexation with ruthenium to obtain three Ru(II) arene complexes represented by [Ru(η6-p-cym)(L)Cl], [Ru(η6-p-cym)(L)(PPh3)]+ and [Ru(η6-p-cym)(L)(PTA)]+ (where p-cym = p-cymene, PPh3 = triphenylphosphine and PTA = 1,3,5-triaza-7-phosphaadamantane). The synthesized complexes were characterized using spectroscopic techniques. UV-Vis absorption spectroscopy and LC-MS were used to study the stability of the complexes in biological medium. Their lipophilicity was studied by calculating the partition coefficient in n-octanol and water. The complexes showed significant binding with biomolecules like albumin proteins and nucleic acids. All the complexes were found to be cytotoxic, with complex [Ru(η6-p-cym)(L)PPh3]PF6 exhibiting the highest anticancer activity. The mechanism of anticancer activity was attributed to the ability of the complexes to induce apoptosis and generate reactive oxygen species (ROS). The complexes also exhibited antimetastatic properties. Furthermore, complex [Ru(η6-p-cym)(L)PPh3]PF6 was loaded onto amine-functionalized mesoporous silica nanoparticles which led to an increase in its cytotoxic activity.
1. Introduction
The discovery of cisplatin in the 1960s was a breakthrough in medicinal chemistry and cancer therapy. Cisplatin binds to N7 of guanine of DNA, causing inhibition of the cell division process and ultimately leading to apoptosis. Despite being effective against various cancers, it suffers from several side effects like nephrotoxicity, ototoxicity, and nausea, which occur because of lack of selectivity. The problems of toxicity and selectivity can be resolved using nanotechnology, but the problem of resistance acquired by cancer cells toward platinum-based drugs requires a novel solution. Thus, switching to alternative metal ions with different mechanistic pathways, greater target specificity and fewer side effects has become an urgent need.1–3 Ruthenium(II) complexes because of their similar ligand exchange kinetics to platinum(II) complexes (similar to the cell division rate) and lesser toxicity seem to be promising candidates.4 Also, since ruthenium can mimic iron in binding to biomolecules, it can be selectively transferred to cancer cells having overexpressed transferrin proteins on their membranes.5 A few ruthenium(III) drugs, namely NAMI-A (imidazolium trans-[tetrachloro(dimethylsulfoxide)(1H-imidazole)ruthenium(III)]), KP1019 (indazolium trans-[tetrachlorobis(1H-indazole)ruthenium(III)]), and NKP-1339, the sodium salt of KP1019, have attracted a lot of attention as promising anticancer drugs but these drugs are not very stable and can undergo facile chemical transformations. These complexes are reduced to ruthenium(II) in the hypoxic acidic environment of tumor cells.6,7RAPTA complexes, which stand for ruthenium arene PTA complexes, have gained significant attention over the last few years.8 The hydrophobic arene ring facilitates the transportation of the drug molecule through the hydrophobic lipid bilayer of the cell membrane, and PTA (1,3,5-triaza-7-phosphaadamantane), a water-soluble ligand, enhances the aqueous solubility of the complex.9 Also, by varying the other ligands, one can modulate the overall reactivity of the complex. In addition to binding to DNA, these piano-stool complexes can also bind to serum proteins involved in transporting the drug molecules to the desired targets. They can also act as multitargeted drugs by targeting certain enzymes and peptides that are overexpressed in cancer cells.10 The choice of ligand plays a major role in designing anticancer drugs. The dicarboxylate chelating ligand in carboplatin confers greater stability to the complex, resulting in fewer side effects than cisplatin.11 Thus, the biological properties of ruthenium complexes can be modulated through appropriate choice of ligands.12 The imine group in Schiff bases can form intermolecular hydrogen bonds with intracellular active components, resulting in various pharmacological applications. Ligands with extensive π-conjugation like phenanthroline, anthracene, naphthalene, and pyrene derivatives render fluorescent nature to the complexes, making them suitable for monitoring the probable mechanism of anticancer activity and cellular distribution.13 Furthermore, benzimidazole derivatives, due to their similarity in structure to natural molecules, are known to possess several biological activities, such as antimicrobial, antifungal, and anticancer activities.14 To increase the therapeutic performance of drugs, various nanocarrier-based delivery systems have been developed, which are known for targeted delivery and stimuli active nature. Mesoporous silica nanoparticles (MSNs), because of their several properties like tunable pore size, large surface area, biocompatibility, high stability, facile synthesis, and functionalization, prove to be an excellent candidate for delivering drugs.15 Their ability to transport insoluble hydrophobic antitumor drugs and enable sustained release make them an efficient vehicle for drug delivery.16 In particular, amine-functionalized MSNs (A-MSNs) are known to have higher loading capacity due to the formation of weak intermolecular forces between the amine group and the polar groups present in the drug molecule.17 They are also known for improved cellular uptake of cationic ruthenium complexes arising due to the strong interaction between the positively charged surface of A-MSNs and the negatively charged phospholipids of the cell membrane.18 The efficacy of anticancer platinum-based drugs is also known to improve upon using MSNs.19
This work involves the synthesis of a benzimidazole-based fluorescent ligand (HL) possessing inherent biological activity followed by its complexation with ruthenium to yield three ruthenium(II) arene complexes with p-cymene as the arene group. The objective of this work is to investigate the effect of co-ligands on anticancer activity and study the structure–activity relationship. Thus, the co-ligands have been varied between chloride (a good labile group that should facilitate the ligand exchange reactions as in cisplatin), triphenylphosphine (a hydrophobic group expected to facilitate the transportation of the ruthenium complex through the hydrophobic lipid bilayer of the cell membrane) and PTA (a water-soluble ligand which increases the overall water solubility of the complex). All the complexes were studied for their cytotoxicity on HeLa and DU145 cells, and the complex possessing the highest anticancer activity was further loaded on amine-functionalized MSNs (A-MSNs) to explore if its therapeutic potential gets enhanced upon incorporation into a drug delivery system.
2. Results and discussion
2.1 Synthesis and characterization
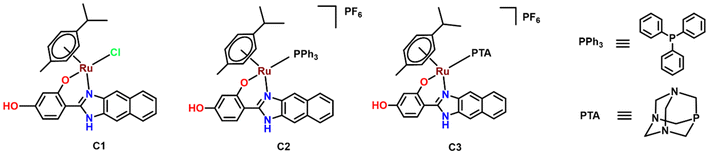
Three piano-stool Ru(II) arene complexes have been synthesized and characterized. The complexes have pseudo-octahedral geometry, with the arene moiety occupying three coordination sites and the bidentate chelating ligand L occupying two other coordination sites. N and O act as the donor sites for the chelating ligand. p-Cymene was chosen as the arene group because of its better structure–activity relationship (SAR) arising from its increased solubility.20 The benzimidazole-based chelating ligand HL was chosen because of its inherent biological activity attributed to its structural similarity to the purine bases of DNA and the ability of its imine bond to form intermolecular H-bonds with intracellular components.14 An additional phenyl ring was added to increase π-conjugation in the system rendering inherent fluorescence to the complex and a hydroxy group was introduced with the expectation of H-bonding interactions with the amine groups on A-MSNs.13HL was designed as a chelating ligand as chelating ligands are known to confer better chemical stability and thus lower toxicity.11 The co-ligands were varied between chloride (Cl−), PPh3, and PTA. Cl− was chosen because of its good leaving group nature which allows easy dissociation followed by coordination of the metal ion with biomolecules, thereby enhancing its structure–activity relationship (SAR).20 PPh3 was chosen because of its hydrophobic nature which is expected to facilitate the transportation of the drug through the cell membrane. In contrast, PTA, because of its water-soluble property, is expected to increase the aqueous solubility of the drug. The effect of global charge on the Ru(II) complex was also investigated by involving a neutral Ru(II) complex (C1) and two cationic Ru(II) complexes (C2 and C3) in the study. For the cationic complexes, hexafluorophosphate was employed as a counterion, and NH4PF6 was utilized as the precursor. C1 was synthesized by the reaction of the Ru p-cymene dimer and the ligand (HL). The chloride ligand was substituted with PPh3 or PTA, followed by the addition of the necessary counter anions to obtain complexes C2 and C3, respectively (Scheme 1). MSN was synthesized using the surfactant CTAB (cetyltrimethyl ammonium bromide) and the silica precursor TEOS (tetraethyl orthosilicate). APTES ((3-aminopropyl)triethoxysilane) was used to functionalize the surface of MSN with NH2 groups and complex C2 was loaded onto it. The synthetic procedures along with the reaction schemes are provided in the ESI.†All the synthesized complexes were characterized using ESI-MS spectra. The ligand HL shows a base peak corresponding to [H2L]+, while for complex C1, the peak is obtained due to the [RuL(p-cym)]+ fragment after losing one chloride ion. Complexes C2 and C3 show peaks corresponding to the cationic complex part. The NMR data matched well with the predicted structures. The 1H NMR peaks in the δ 5.25–6.05 ppm region correspond to the p-cymene aromatic protons, while the aromatic proton peaks of the ligand (L) appear at δ 8.27–6.45 ppm. The peaks at δ 2.8, 1.7–2.09, and 0.9–1.2 ppm are attributed to the methyl and isopropyl groups of the p-cymene ring. The highly deshielded broad peaks beyond 10 ppm correspond to OH and NH protons.21 The PTA protons are observed in the region δ 4.73–4.26 ppm, and the peaks for the PPh3 aromatic ring protons appear in the region δ 7.44–7.15 ppm (merging with the aromatic range of ligand L).9
The FTIR spectra of complexes C1, C2, and C3, and the ligand (Fig. S9†) exhibited a band at around 1600 cm−1 corresponding to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N functionality.22 The bands for N–H and O–H stretching frequencies were found to overlap in the region of 3200–3460 cm−1. The band at 1230–1240 cm−1 corresponds to the aromatic C–O stretching frequency. The presence of characteristic bands at 526 and 540 cm−1 in the IR spectra of C2 and C3 corresponds to the Ru–P stretching frequency, signifying successful complexation with ruthenium.23 Bands at 831 and 821 cm−1 in case of complexes C2 and C3 arise due to the P–F stretching frequency of the counter anion.24
N functionality.22 The bands for N–H and O–H stretching frequencies were found to overlap in the region of 3200–3460 cm−1. The band at 1230–1240 cm−1 corresponds to the aromatic C–O stretching frequency. The presence of characteristic bands at 526 and 540 cm−1 in the IR spectra of C2 and C3 corresponds to the Ru–P stretching frequency, signifying successful complexation with ruthenium.23 Bands at 831 and 821 cm−1 in case of complexes C2 and C3 arise due to the P–F stretching frequency of the counter anion.24
MSN, A-MSN, and drug-loaded A-MSN (C2@A-MSNs) were characterized using IR spectroscopy, FE-SEM, TGA, and BET. The FT-IR spectra of MSN, A-MSN, and C2@A-MSNs show bands at 1072 cm−1, 794 cm−1, and 456 cm−1 corresponding to the silanol groups (Si–O bond). The bare silanol groups on the surface of pristine MSN can be confirmed by bands at 3364 cm−1 and 1642 cm−1 corresponding to the deformation vibration of interlayer water and the OH stretching vibration of the silanol groups, respectively.25 The amine functionalization of MSN was confirmed by the bending vibration mode of N–H which appeared at 1560 cm−1.18 The loading of complex C2 on A-MSNs was confirmed by the imine band which appeared at 1604 cm−1. The shift of the broad band corresponding to the O–H stretching frequency from 3300 cm−1 (free complex) to 3029 cm−1 (C2@A-MSNs) represents H-bonding interactions that may occur between the NH2 groups of A-MSNs and the free OH group present in complex C2 (Fig. S10†).17
The mesoporous nature of the MSNs and A-MSNs was characterized using N2 adsorption–desorption analysis (Fig. S12†). The decrease in BET surface area from 752.617 m2 g−1 to 68.326 m2 g−1 and in pore diameter from 1.49202 nm to 1.32461 nm indicates the successful introduction of amine groups on the MSN surface. The surface area further decreased to 47.093 m2 g−1 in C2@A-MSNs as C2 was loaded onto the nanoparticles. This was further confirmed by the decrease in pore volume from 2.807 × 10−1 cc g−1 (MSNs) to 1.951 × 10−2 cc g−1 (A-MSNs) to 1.923 × 10−2 cc g−1 (C2@A-MSNs).18 As observed from the TGA analysis, the drug-loaded system is thermally stable up to 200 °C as there is no significant weight loss except for the removal of water molecules and other solvents adsorbed on the surface (Fig. S11†).26–29 The amorphous nature of the nanoparticles was confirmed by SEM images (Fig. S13†). The loading of complex C2 onto A-MSNs causes the surface to appear rough. The powder X-ray diffraction pattern shows peaks corresponding to the hexagonal mesoporous pattern in MSN and A-MSNs.17 The broad nature of the peaks also justifies the amorphous nature of the silica nanoparticles (Fig. S14†).
The absorption and emission spectra of all the complexes and the ligand were recorded in DMSO. In the UV-Vis spectra, two bands were observed for all the complexes and the ligand. The band at around 275–280 nm corresponded to the π → π* transition, and the one at 340–360 nm corresponded to the n → π* transition. An additional band was also observed only in the case of metal complexes in the range of 390–400 nm, which corresponds to the MLCT transition (Fig. S15a†).30
It is important to note that all the synthesized complexes and the ligand were found to be fluorescent. Their fluorescence spectra were recorded by exciting at a wavelength of 360 nm. Complex C1 exhibited the highest fluorescence intensity, followed by complex C2 and complex C3 (Fig. S16†). The bands corresponding to the free C2 complex were observed when the absorbance spectrum of C2@A-MSNs was recorded, indicating the successful loading of the drug onto amine-functionalized MSNs. The shift of the n → π* transition to a lower wavelength in the case of C2@A-MSNs further confirmed the involvement of H-bonding interactions between A-MSNs and the polar groups of C2 (Fig. S15b†).31 SEM-EDX for the drug-loaded nanoparticles was also recorded, indicating the presence of Ru in C2@A-MSNs, which confirmed the successful loading of C2 onto A-MSNs (Fig. S17a–c†).
2.2 Stability study
The stability of the complexes in biological medium was explored for the prospective application of the compounds as anticancer drugs. The complexes were dissolved in a 1% DMSO/PBS solution mixture, and the UV-Vis spectra were recorded at 0, 6, 12, and 24 hours. The spectra showed no significant change or shift in band positions, indicating that the complexes are stable in the biological medium. Additionally, LC-MS data were recorded over a period of 48 hours and the main peak remained unaltered, confirming the integrity of the molecule in the biological medium (Fig. S19a–c†).Since the tumor microenvironment is acidic in nature and the drug release study is performed at an acidic pH, it is crucial to check whether complex C2 can maintain its integrity at a lower pH. Thus, the stability of complex C2 at an acidic pH of 5.4 was monitored over 0, 24, 48, and 72 hours using absorption spectroscopy. No shift in the absorbance band indicated that the complex does not degrade in an acidic pH for up to 72 hours (Fig. S18†).
2.3 Lipophilicity study
Lipophilicity refers to the ability of a compound to pass through the lipid bilayer of the cell membrane and enter the cell. It is an important parameter regulating the cellular uptake of drugs. The lipophilicity of the synthesized and loaded complexes was determined from the partition coefficient PO/W in n-octanol and water by the shake-flask method. UV-Vis spectroscopy was used to determine the concentration of the complex in n-octanol (organic phase) and water (aqueous phase), and the partition coefficient was calculated. Complex C2 and C2@A-MSNs with the triphenylphosphine group was found to have the highest partition coefficient values, making them the most lipophilic among all the tested complexes. This was probably due to the presence of the hydrophobic phenyl rings, which facilitate interaction in the non-polar medium.21 Table S2† lists the partition coefficient values for the three complexes.2.4 Protein binding study
 | ||
| Fig. 1 Fluorometric titration spectra of BSA (10 μM) with (a) ligand (HL), (b) complex C1, (c) complex C2, and (d) complex C3 at 298 K. | ||
The binding parameters were calculated using the Stern–Volmer equation and the Scatchard equation (Fig. S21 and S22†). All the complexes exhibited significant interactions with proteins. The bimolecular quenching constants for all the complexes and the ligand are on the order of 1013, indicating static quenching of fluorescence (Table S2†).13 Complex C1 shows a higher KSV value because of the presence of a chloride ligand, which dissociates easily, leading to subsequent binding of other biomolecules to Ru.
2.5 Nucleic acid binding study
2.6 Drug loading and pH-stimulated release study
Mesoporous silica nanoparticles (MSNs) because of their tunable properties have emerged as an efficient way of loading drugs and selectively delivering them to the desired target. Cytotoxicity of cisplatin was enhanced when it was loaded on mesoporous silica materials (MCM-41 and SBA-15).19 Doxorubicin when loaded on MSNs coated with polydopamine (PDA) and polyethylene glycol (PEG) exhibited higher cellular uptake and improved anticancer efficacy compared to the free drug.34 The antibiotic drug rifampicin when encapsulated by MSNs resulted in better treatment efficiency than the free drug due to greater uptake by macrophages.35 Since loading of drugs on MSNs significantly improves their efficiency compared to the free drug, the next target was to load the Ru(II) complex C2 and see if there is any enhancement in anticancer activity. Determining the drug loading capacity of nanoparticles is crucial to study the drug activity and release. The loading capacity was found to be 40.4% (40 mg/100 mg) for C2@A-MSNs which was calculated using the weight % method (eqn (1)†).36 Cancer cells have a hypoxic and acidic environment compared to normal cells. Thus, PBS buffers of acidic pH (5.4) and neutral pH (7.4) were used to investigate the selective release of drugs, specifically in cancer cells and normal cells. The drug release profile shows that in the case of neutral pH, after 4 hours, no detectable amount of the drug is released, and after 96 hours, only 30% of the drug is released. However, at acidic pH, a small amount of the drug is released in the initial hours, and 65% is released after 96 hours (Fig. 2). The greater amount of drug release in acidic pH may be attributed to the protonation of amine groups, causing the cleavage of H-bonding interactions between A-MSNs and the complex (Fig. 3).18,26The integrity of the released drug C2 was characterized by mass spectrometry. The appearance of a peak corresponding to the free complex C2 in the mass spectrum after 96 hours at acidic pH confirmed the identity of the released complex (Fig. S28†).
2.7. Cell viability assay
An in vitro MTT assay was performed to determine whether the synthesized complexes were cytotoxic against cancer cells. MTT is a tetrazolium yellow-coloured salt that is reduced to formazan – a purple-coloured insoluble product in the presence of NADPH, an enzyme present only in live cells. Cancer cell lines HeLa and DU145 were treated with different concentrations (0–100 μM) of the synthesized Ru(II) complexes, and their IC50 values were calculated using the cell viability curve. All the complexes had potent cytotoxic activity against the above cell lines, and they were more responsive to HeLa cells (Fig. 4). Among the three complexes, complex C2 was found to have the lowest IC50 value (17.78 μM in the case of HeLa and 27.42 μM in the case of DU145) and, hence, the highest cytotoxicity which was attributed to the hydrophobic triphenylphosphine group which leads to enhanced hydrophobic interactions with biomolecules. These data are also in accordance with the results obtained from the lipophilicity study, which indicated the highest cellular uptake in case of complex C2. It is noteworthy to mention that all the complexes are active against cisplatin-resistant DU145 cells. The IC50 value further decreased upon loading complex C2 on A-MSNs, indicating an enhancement in the cytotoxic profile. This was probably due to better cellular internalization occurring due to the interaction between negatively charged phospholipids and the positively charged surface of A-MSNs.18 Also, the free ligand has a higher IC50 value than the ruthenium complexes indicating the advantage of the metal complexes over the free ligand. The complexes were also tested on normal cell lines HEK 293 and slight selectivity was observed for HeLa cell lines. Complex C2 and C2@A-MSNs exhibited decent selectivity for the cancer cell lines over HEK 293 (Fig. 4 and S29†). The IC50 values for all the complexes and drug-loaded A-MSNs are listed in Table 1. | ||
| Fig. 4 Histogram representing the IC50 values (μM) of the ligand, three complexes and C2@A-MSNs for cancer cell lines (a) HeLa, (b) DU145 and (c) normal HEK 293 cells. | ||
| IC50 values (in μM) | |||
|---|---|---|---|
| Compounds | HeLa | DU145 | HEK 293 |
| Ligand HL | 30.79 ± 0.56 | 41.13 ± 0.53 | 32.28 ± 0.45 |
| Complex C1 | 19.21 ± 0.43 | 30.11 ± 0.83 | 33.73 ± 0.51 |
| Complex C2 | 17.78 ± 0.48 | 27.42 ± 0.58 | 34.59 ± 0.59 |
| Complex C3 | 20.26 ± 0.36 | 31.46 ± 0.74 | 36.21 ± 0.44 |
| C2@A-MSNs | 14.01 ± 0.43 | 19.83 ± 0.75 | 31.83 ± 0.39 |
| Cisplatin | 17.40 ± 0.38 | 40.22 ± 0.59 | 23.18 ± 0.28 |
2.8 Apoptosis study using a dual staining method
Since, C2 and C2@A_MSNs were found to be the most potent among all the candidates, they were further studied for apoptosis, ROS generation and metastasis inhibition activity. The dual staining assay was performed using two DNA-binding dyes to study whether the complexes are potent of inducing apoptosis and to investigate the morphological changes associated with it. Acridine orange (AO) is a membrane-permeable dye capable of staining both viable and non-viable cells, and ethidium bromide (EtBr) is a membrane-impermeable dye, thereby capable of staining only non-viable cells with ruptured membranes. Upon binding to DNA, AO emits bright green fluorescence, while EtBr emits bright red fluorescence.37 The confocal images show that C2 and C2@A-MSNs are capable of inducing apoptosis. Morphological changes like ruptured cell membranes and fragmented nuclei were observed, which are characteristic of apoptosis (Fig. 5). The apoptosis results for C1 and C3 are shown in Fig. S30.† | ||
| Fig. 5 Confocal images of HeLa cells stained with AO and EtBr after treatment with complex C2 and C2@A-MSNs. | ||
2.9 ROS generation: DCFH DA staining
The generation of reactive oxygen species (ROS) is an efficient way of killing cancer cells by producing oxidative stress inside the cells, which causes damage to intracellular substances. The generation of ROS in quantities greater than what can be scavenged by cellular enzymes like superoxide dismutase (SOD) causes oxidative damage to the cells. Additionally, because of the rapidly multiplying nature of cancer cells, they require a greater amount of oxygen than normal cells. Thus, converting oxygen into reactive oxygen species like superoxide anion, hydrogen peroxide, and OH˙ can hamper cell division, thereby causing cell death.38 Thus, the synthesized complexes were investigated for their ability to generate ROS using the DCFH-DA staining method. DCFH-DA (dichloro-dihydro-fluorescein diacetate) is a dye that, in the presence of ROS, gets oxidized to DCF, which emits green fluorescence.39 Thus, it can be used for the detection of ROS. The microscope images clearly show green fluorescence only in the case of treated cells and not in the control, suggesting that C2 and C2@A-MSNs can generate ROS, which are responsible for the green fluorescence (Fig. 6). Furthermore, from the fluorescence intensity graph (Fig. S31†), it is noticed that complex C2 and C2@A-MSNs can generate ROS to a greater extent.2.10 Metastasis inhibition
Metastasis refers to the spread of primary cancer cells to surrounding tissues and distant organs. The ability of cancer cells to migrate to non-malignant parts of the body and proliferate there makes it difficult to control the disease, making metastasis a dominant reason for mortality.40 Thus, inhibiting the migration of malignant cells plays a major role in anticancer activity. To investigate the ability of the complexes to inhibit the cell migration process, a wound-healing assay was performed on HeLa cell lines. The cells were treated with the complexes at their IC50 values and the wound coverage was monitored for 24 hours. Compared to the control, where no drug was used, C2 and C2@A-MSNs exhibited decreased wound coverage, indicating lesser cell migration to the scratched wound region (Fig. 7). Among the three free complexes, C2 exhibited the maximum potential in inhibiting cell migration with a wound closure ratio of 35% and the antimetastatic property was further enhanced upon treatment with C2@A-MSNs (Fig. S32 and S33†). | ||
| Fig. 7 Wound healing motility assay of HeLa untreated and treated cell with free complex C2 and C2@A-MSNs. Images were taken at 0 and 24 h. | ||
3. Conclusion
Three ruthenium(II) arene complexes with p-cymene as the arene group and a bidentate benzimidazole-based chelating ligand with N and O as donor sites have been synthesized. The co-ligands have been varied between chloride, PPh3 (triphenylphosphine) and PTA (1,3,5-triaza-7-phosphaadamantane). The complexes were characterized using several spectroscopic techniques like ESI-MS, IR, NMR and UV-Vis spectroscopy and their molecular formulas are found to be [Ru(η6-p-cym)(L)Cl], [Ru(η6-p-cym)(L)PPh3]PF6, and [Ru(η6-p-cym)(L)PTA]PF6. Both the ligand and the complexes were found to be fluorescent in nature. The complexes were studied for their lipophilic behaviour and complex C2 was found to be the most lipophilic due to the increased hydrophobicity of the phenyl rings. All the complexes are stable in biological medium for up to 48 hours. Tryptophan quenching experiments suggested that the complexes significantly bind to albumin proteins like BSA and HSA. As is evident from absorption and emission spectroscopy, the complexes also show potent binding to CT-DNA and RNA and the mode of binding was found to be intercalation. However, this does not provide any evidence of DNA being the target molecule for exhibiting cytotoxicity. A cytotoxicity study was performed on HeLa and DU145 cells and all the complexes showed anticancer activity with better results on HeLa cells. Among all the three complexes, complex C2 exhibited the lowest IC50 value and the highest anticancer activity due to the presence of the hydrophobic triphenylphosphine group which led to better interaction with the biomolecules as compared to chloride or PTA. The hydrophobic nature of the phenyl rings also increased the lipophilicity of the complex. The overall cationic charge of the complex also enhanced the solubility of the complex when compared to its neutral counterpart C1. Since complex C2 was the most potent anticancer drug among the three complexes, it was loaded on amine-functionalized mesoporous silica nanoparticles to see whether these drug delivery nanocarriers can enhance its therapeutic performance. As observed from the cytotoxicity study, the anticancer activity was improved upon loading and their selectivity for cancer cells also increased. The enhancement in their therapeutic performance was attributed to the interactions between the negatively charged phospholipids and the positively charged surface of the nanoparticles which led to better cellular internalization. Complex C2 and C2@A-MSNs were also capable of inducing apoptosis, generating ROS and exhibiting anti-metastatic activity on HeLa cell lines. In conclusion, the above synthesized complexes can act as potent anticancer agents and further studies can be conducted with in vivo tumor models.Data availability
Synthetic procedures and experimental details of the in vitro biological study, detailed characterisation of the complexes and materials, and spectroscopic data can be found in the ESI.†Conflicts of interest
There are no conflicts to declare.Acknowledgements
The instrument facilities provided by SIC, IIT Indore, are gratefully acknowledged. The authors Prgati and Amardeep Kumar thank UGC and CSIR respectively for the fellowship. We gratefully acknowledge DST-FIST [Project No.- SR/FST/CSII/2017/22(C)] for providing a 500 MHz NMR facility.References
- S. Ghosh, Bioorg. Chem., 2019, 88, 102925 CrossRef CAS PubMed.
- R. Oun, Y. E. Moussa and N. J. Wheate, Dalton Trans., 2018, 47, 6645–6653 RSC.
- S. H. van Rijt and P. J. Sadler, Drug Discovery Today, 2009, 14, 1089–1097 CrossRef CAS PubMed.
- J. Reedijk, Metallomics, 2012, 4, 628–632 CrossRef CAS PubMed.
- S. Y. Lee, C. Y. Kim and T.-G. Nam, Drug Des., Dev. Ther., 2020, 14, 5375–5392 CrossRef CAS PubMed.
- E. Alessio and L. Messori, Molecules, 2019, 24, 1995 CrossRef CAS PubMed.
- Pragti, B. K. Kundu, C. Sonkar, R. Ganguly and S. Mukhopadhyay, Polyhedron, 2021, 207, 115379 CrossRef CAS.
- R. Pettinari, F. Marchetti, F. Condello, C. Pettinari, G. Lupidi, R. Scopelliti, S. Mukhopadhyay, T. Riedel and P. J. Dyson, Organometallics, 2014, 33, 3709–3715 CrossRef CAS.
- P. Mandal, N. Malviya, B. K. Kundu, S. S. Dhankhar, C. M. Nagaraja and S. Mukhopadhyay, Appl. Organomet. Chem., 2018, 32, e4179 CrossRef.
- Pragti, B. K. Kundu and S. Mukhopadhyay, Coord. Chem. Rev., 2021, 448, 214169 CrossRef CAS.
- R. G. Kenny and C. J. Marmion, Chem. Rev., 2019, 119, 1058–1137 CrossRef CAS PubMed.
- H. A. Sahyon, A. A. El-Bindary, A. F. Shoair and A. A. Abdellatif, J. Mol. Liq., 2018, 255, 122–134 CrossRef CAS.
- Pragti, B. K. Kundu, S. N. Upadhyay, N. Sinha, R. Ganguly, I. Grabchev, S. Pakhira and S. Mukhopadhyay, Dalton Trans., 2022, 51, 3937–3953 RSC.
- N. Shrivastava, M. J. Naim, M. J. Alam, F. Nawaz and S. Ahmed, Arch. Pharm., 2017, 350, e201700040 CrossRef PubMed.
- Y. Li, G. Deng, X. Hu, C. Li, X. Wang, Q. Zhu, K. Zheng, W. Xiong and H. Wu, Nanomedicine, 2022, 17, 1253–1279 CrossRef CAS PubMed.
- B. Siddiqui, A. ur. Rehman, I. Haq, A. A. Al-Dossary, A. Elaissari and N. Ahmed, Int. J. Pharm.: X, 2022, 4, 100116 CAS.
- R. Kotcherlakota, A. K. Barui, S. Prashar, M. Fajardo, D. Briones, A. Rodríguez-Diéguez, C. R. Patra and S. Gómez-Ruiz, Biomater. Sci., 2016, 4, 448–459 RSC.
- M. M. Carmona, Q. P. Ho, J. Morand, A. García, E. Ortega, L. C. S. Erthal, E. R. Hernandez, M. D. Santana, J. Ruiz, M. V. Regí and Y. K. Gun'ko, Inorg. Chem., 2020, 59(14), 10275–10284 CrossRef PubMed.
- Z. Tao, B. Toms, J. Goodisman and T. Asefa, ACS Nano, 2010, 4, 789–794 CrossRef CAS PubMed.
- M. Martínez-Alonso, N. Busto, F. A. Jalón, B. R. Manzano, J. M. Leal, A. M. Rodríguez, B. García and G. Espino, Inorg. Chem., 2014, 53, 11274–11288 CrossRef PubMed.
- P. Porwal, S. Nayek, S. Singh, A. Sonawane, I. Grabchev, R. Ganguly and S. Mukhopadhyay, Dalton Trans., 2023, 52, 7104–7118 RSC.
- B. K. Kundu, Pragti, Reena, S. M. Mobin and S. Mukhopadhyay, New J. Chem., 2019, 43, 11483–11492 RSC.
- J. G. Małecki and A. Maroń, Transition Met. Chem., 2012, 37, 727–734 CrossRef.
- A. O. Adeloye, T. O. Olomola, A. I. Adebayo and P. A. Ajibade, Int. J. Mol. Sci., 2012, 13, 3511–3526 CrossRef CAS PubMed.
- P. Qin, Y. Yang, W. Li, J. Zhang, Q. Zhou and M. Lu, Anal. Methods, 2018, 11, 105–112 RSC.
- B. K. Kundu, Pragti, W. A. Carlton Ranjith, U. Shankar, R. R. Kannan, S. M. Mobin, A. Bandyopadhyay and S. Mukhopadhyay, ACS Appl. Bio Mater., 2022, 5, 190–204 CrossRef CAS PubMed.
- M. Almaghrabi, A. Alqurshi, S. A. Jadhav, F. Mazzacuva, A. Cilibrizzi, B. Raimi-Abraham and P. G. Royall, Thermochim. Acta, 2023, 730, 179616 CrossRef CAS.
- H. Zhang, C. Li, J. Guo, L. Zang and J. Luo, J. Nanomater., 2012, 2012, 217412 CrossRef.
- M. Mladenović, I. Morgan, N. Ilić, M. Saoud, M. V. Pergal, G. N. Kaluđerović and N. Ž. Knežević, Pharmaceutics, 2021, 13, 460 CrossRef PubMed.
- R. M. Ramadan, W. M. Elsheemy, N. S. Hassan and A. A. A. Aziz, Appl. Organomet. Chem., 2018, 32, e4180 CrossRef.
- G. J. Brealey and M. Kasha, J. Am. Chem. Soc., 1955, 77, 4462–4468 CrossRef CAS.
- E. N. Hoogenboezem and C. L. Duvall, Adv. Drug Delivery Rev., 2018, 130, 73–89 CrossRef CAS PubMed.
- C. S. Devi, B. Thulasiram, S. Satyanarayana and P. Nagababu, J. Fluoresc., 2017, 27, 2119–2130 CrossRef CAS PubMed.
- Y. Duo, Y. Li, C. Chen, B. Liu, X. Wang, X. Zeng and H. Chen, RSC Adv., 2017, 7, 39641–39650 RSC.
- S. Subramaniam, N. Thomas, H. Gustafsson, M. Jambhrunkar, S. P. Kidd and C. A. Prestidge, Antibiotics, 2019, 8, 39 CrossRef CAS PubMed.
- G. Lv, L. Qiu, G. Liu, W. Wang, K. Li, X. Zhao and J. Lin, Dalton Trans., 2016, 45, 18147–18155 RSC.
- K. Liu, P. Liu, R. Liu and X. Wu, Med. Sci. Monit. Basic Res., 2015, 21, 15–20 CrossRef PubMed.
- H. Nakamura and K. Takada, Cancer Sci., 2021, 112, 3945–3952 CrossRef CAS PubMed.
- H. Kim and X. Xue, J. Visualized Exp., 2020, 160 DOI:10.3791/60682.
- C. Sonkar, S. Sarkar and S. Mukhopadhyay, RSC Med. Chem., 2022, 13, 22–38 RSC.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4dt03536d |
| This journal is © The Royal Society of Chemistry 2025 |