Open Access Article
Open Access ArticleComment on “Natural and synthetic flavonoid derivatives as new potential tyrosinase inhibitors: a systematic review” by R. Obaid, E. Mughal, N. Naeem, A. Sadiq, R. Alsantali, R. Jassas, Z. Moussa and S. Ahmed, RSC Advances, 2021, 11, 22159
Hubert Wojtasek
Institute of Chemistry, Opole University, Ul. Oleska 48, 45-052 Opole, Poland. E-mail: Hubert.Wojtasek@uni.opole.pl; Fax: +48 77 452 7101; Tel: +48 77 452 7122
First published on 14th February 2022
Abstract
A review article has been published recently (RSC Advances, 2021, 11, 22159–22198) describing flavonoids as inhibitors of tyrosinase. However, many compounds included in this review have been previously shown to act as substrates of this enzymes or antioxidants reducing tyrosinase-generated o-quinones. Products of their oxidation absorb light in a range different than dopachrome, the oxidation product of L-tyrosine or L-dopa, whose concentration is measured spectrophotometrically in the standard enzymatic assay to monitor the activity of this enzyme. This effect is interpreted as enzyme inhibition, which, in fact, is only apparent and results from inadequate methodology.
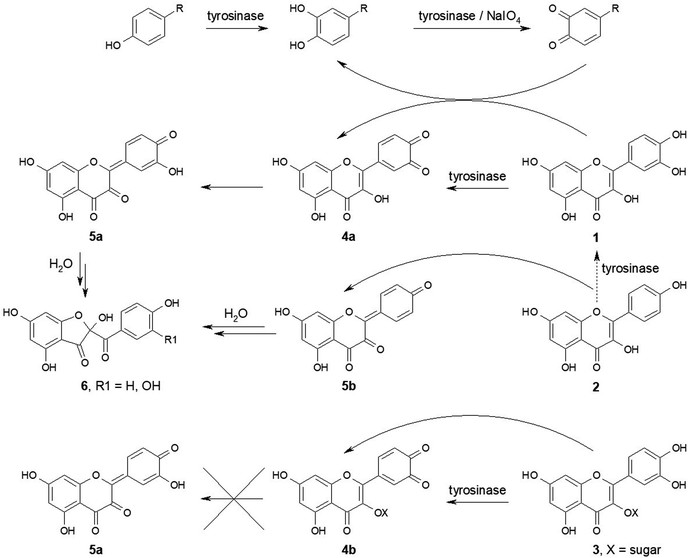
RSC Advances has recently published a review article describing flavonoids as inhibitors of tyrosinase.1 This article, however, overlooked important aspects of interactions of this group of compounds with this enzyme. It is surprising that such interactions were omitted, because just two months earlier RSC Advances published a paper describing oxidation of four flavonoids – apigenin, luteolin, kaempferol, and quercetin by tyrosinase and their promotional effect on mushroom browning.2 Studies in this area were initiated in the 1990s by Isao Kubo and co-workers, who discovered that flavonoids isolated from American plants strongly inhibited tyrosinase.3–5 The active compounds were identified as quercetin, kaempferol, luteolin and several of their derivatives.3–5 Subsequently mechanisms of inhibition by quercetin and kaempferol were studied in more detail.6–8 Later, however, oxidation of quercetin by polyphenol oxidase from broad bean (Vicia faba)9 and by mushroom (Agaricus bisporus) tyrosinase10,11 was demonstrated. The reaction pathway was elucidated and the product of these reactions was identified as a relatively stable protocatechuate derivative – 2-(3,4-dihydroxybenzoyl)-2,4,6-trihydroxy-3(2H)-benzofuranone12 (Fig. 1). We later demonstrated that both quercetin and kaempferol (as well as several other flavonoids) were not only oxidized by tyrosinase, although in most cases at rates much smaller than model substrates, such as L-tyrosine or L-dopa, but they reacted with o-quinones generated from catechols by oxidation with either tyrosinase or chemical oxidants, such as sodium periodate13 (Fig. 1, curved arrows). Such reactions were also described in the paper published recently in RSC Advances.2 Oxidation of flavonoids by three major groups of plant oxidoreductases – laccases, polyphenol oxidases, and peroxidases has also been well described in an excellent review.14 Our paper13 was intended to warn all researchers trying to find inhibitors of tyrosinase that the standard assay used to monitor the activity of this enzyme (oxidation of L-tyrosine or L-dopa to dopachrome) does not give reliable results with compounds possessing strong reducing properties, such as flavonoids. They reduce the enzymatically-generated dopaquinone, thus preventing formation of dopachrome, whose concentration is measured spectrophotometrically in this assay. This effect is then interpreted as enzyme inhibition, which, in fact, is only apparent and results from inadequate methodology.
Many papers cited in this review1 describe flavonoids containing a catechol group in either ring A or B, or a conjugated system of hydroxy groups in position 3 and either 2′ or 4′ as inhibitors of tyrosinase. They should, however, be substrates of this enzyme or should undergo oxidation by tyrosinase-generated o-quinones, as we described for quercetin, kaempferol, morin and catechin.13 Compounds, which should show such properties, or for which they have already been demonstrated, are listed below, with their numbers given in parenthesis:
malvidin (1), peonidin (2), pelargonidin (3), delphinidin (5), cyanidin 3-O-glucoside (7), delphinidin 3-O-glucoside (8), baicalein (57), luteolin (58), 6,7-dihydroxy-2-phenyl-4H-chromen-4-one (63), 7,8,4′-trihydroxyflavone (64), dihydromyricetin (74), taxifolin (75), compound 77 (which is labeled as dihydroxykaempferol, but whose structure actually corresponds to dihydroquercetin), compound 80 (which is labeled as chlorophorin, but whose structure actually corresponds to 2,3,4,5-tetrahydroxycyclohexyl derivative of dihydrofisetin), proanthocyanidins (81), 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-3-methoxychroman-4-one (84), 2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxychroman-4-one (87), 2-(3,4-dihydroxyphenyl)-3,5-dihydroxy-7-[2,3,4-trihydroxy-5-(hydroxymethyl)cyclohexyl]oxychroman-4-one (89), 7,3′,4′-trihydroxyisoflavone (92), 7,8,4′-trihydroxyisoflavone (93), 3′-hydroxygenistein (95), 6-hydroxydaidzein (96), calycosin (97), 7,8-dihydroxy-3-(4-hydroxyphenyl)-4H-chromen-4-one (106, which is identical with compound 93), 7,8-dihydroxy-3-(3-methoxyphenyl)-4H-chromen-4-one (107), 3-(3,4-dihydroxyphenyl)-8-hydroxy-7-methyl-4H-chromen-4-one (108), 7,8-dihydroxy-3-(3-hydroxyphenyl)-4H-chromen-4-one (109), 7,8-dihydroxy-3-(4-methoxyphenyl)-4H-chromen-4-one (110), 7,8-dihydroxy-3-(2-methoxyphenyl)-4H-chromen-4-one (111), 6,7-dihydroxy-3-(4-methoxyphenyl)-4H-chromen-4-one (112), 6,7-dihydroxy-3-(4-hydroxyphenyl)-4H-chromen-4-one (113, which is identical with compound 96), quercetin (not quercitin, 139), kaempferol (140), rutin (142), 2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-4H-chromen-4-one (143, which is identical with compound 139), 2-(3,4-dihydroxyphenyl)-3,5-dihydroxy-7-methoxy-4H-chromen-4-one (144), 6-hydroxykaempferol (145), 6-hydroxygalangin (146).
I don't think it would be necessary to demonstrate that such reactions occur in the case of all these compounds. It should have been done when they were studied as inhibitors of tyrosinase. Cyclohexane rings in flavonols 80 and 89 are surprising. Since the most common natural derivatives of flavonoids are their glycosides, one would expect pyranose rings of sugars in these compounds. Unfortunately, it was not possible to verify these structures, because I could not find compound 80 in ref. 172 and compounds 82–90 in ref. 28.
The Authors state that “The flavonol's structure, 3-hydroxy-4-keto moiety, is essential in copper chelation” (p. 22183). This statement has been repeated in many publications describing flavonols as inhibitors of tyrosinase. However, it was demonstrated more than two decades ago that coordination of Cu2+ ion by the 3-hydroxy-4-keto moiety of flavonol does not occur, in contrast to Fe3+.15 Instead, flavonoids may coordinate Cu2+ ion by a catechol group in ring B, but this binding is rapidly followed by their oxidation.16 This oxidation is greatly enhanced by the presence of the 3-hydroxy group in flavonols. It was first demonstrated for quercetin and kaempferol, which were oxidized much more rapidly than flavonoids without this functional group – rutin (quercetin-3-O-rutinoside, with the 3-OH group blocked by glycosylation) and luteolin (a flavone, not containing the 3-OH group). It was therefore postulated that the 3,4′-dihydroxy system of flavonols was preferentially oxidized by Cu2+ ions.16 Details of the oxidation of quercetin by Cu2+ ions may be found in a subsequent publication.17 Such reactions were demonstrated for other flavonols, such as fisetin, also containing a catechol group in ring B (3′-OH, 4′-OH, as in quercetin), myricetin, containing a triol moiety (3′-OH, 4′-OH, 5′-OH), and morin, containing two phenolic groups not forming a catechol (2′-OH, 4′-OH).15 These results confirmed that the conjugated 3,4′-dihydroxy system of flavonols was oxidized under such conditions. Products of these reactions with quercetin and kaempferol were then identified as the 2-substituted 2,4,6-trihydroxy-3(2H)-benzofuranone derivatives,15 the same as that obtained later by oxidation of quercetin with tyrosinase.12
Unfortunately, many researchers searching for inhibitors of tyrosinase are only familiar with articles reporting this function of compounds they are studying, e.g. flavonoids, but are not aware of subsequent papers demonstrating that they are substrates of this enzyme. This may be exemplified by a statement from p. 22187: “Kubo et al. assumed that the chelation mechanism by flavonols may be attributed to the free 3-OH group”, which is followed by several citations, including one of Kubo's earlier publications.7 However, subsequent papers describing oxidation of quercetin and fisetin by this enzyme, in which this concept was abandoned,12,18 are not mentioned.
The review1 also contains other errors. The bibliography contains only 178 positions, yet in Table 10 ref. 180 appears. Some references appear twice or even three times in the bibliography. Examples that I have found are listed below:
D. Sohretoglu, S. Sari, B. Barut and A. Ozel, Bioorg. Chem., 2018, 81, 168–174 (ref. 15, 79, and 129)
K. Bagherzadeh, F. Shirgahi Talari, A. Sharifi, M. R. Ganjali, A. A. Saboury and M. Amanlou, J. Biomol. Struct. Dyn., 2015, 33, 487–501 (ref. 52(d) and 75)
E. J. Land, C. A. Ramsden and P. A. Riley, Tohoku J. Exp. Med., 2007, 212, 341–348. (ref. 63 and 155)
Y. Kakumu, K. Yamauchi and T. Mitsunaga, Holzforschung, 2019, 73, 637–643. (ref. 72 and 162)
R. R. Arroo, S. Sari, B. Barut, A. Ozel, K. C. Ruparelia and D. Sohretoglu, Phytochem. Anal., 2020, 31, 314–321. (ref. 83 and 128)
N. Guo, C. Wang, C. Shang, X. You, L. Zhang and W. Liu, Int. J. Biol. Macromol., 2018, 118, 57–68. (ref. 93 and 166)
N. Kishore, D. Twilley, A. Blom van Staden, P. Verma, B. Singh, G. Cardinali, D. Kovacs, M. Picardo, V. Kumar and N. Lall, J. Nat. Prod., 2018, 81, 49–56. (ref. 101 and 163)
Conflicts of interest
There are no conflicts to declare.References
- R. J. Obaid, E. U. Mughal, N. Naeem, A. Sadiq, R. I. Alsantali, R. S. Jassas, Z. Moussa and S. A. Ahmed, RSC Adv., 2021, 11, 22159–22198 RSC.
- Y. Lu, Y. Xu, M.-T. Song, L.-L. Qian, X.-L. Liu, R.-Y. Gao, R.-M. Han, L. H. Skibsted and J.-P. Zhang, RSC Adv., 2021, 11, 13769–13779 RSC.
- I. Kubo, Y. Yokokawa and I. Kinst-Hori, J. Nat. Prod., 1995, 58, 739–743 CrossRef CAS PubMed.
- I. Kubo and Y. Yokokawa, Phytochemistry, 1992, 31, 1075–1077 CrossRef CAS.
- I. Kubo, I. Kinst-Hori, K. Ishiguro, S. K. Chaudhuri, Y. Sanchez and T. Ogura, Bioorg. Med. Chem. Lett., 1994, 4, 1443–1446 CrossRef CAS.
- I. Kubo and I. Kinst-Hori, J. Agric. Food Chem., 1999, 47, 4121–4125 CrossRef CAS PubMed.
- I. Kubo, I. Kinst-Hori, S. K. Chaudhuri, Y. Kubo, Y. Sanchez and T. Ogura, Bioorg. Med. Chem., 2000, 8, 1749–1755 CrossRef CAS PubMed.
- Q. X. Chen and I. Kubo, J. Agric. Food Chem., 2002, 50, 4108–4112 CrossRef CAS PubMed.
- M. Jimenez and F. Garcia-Carmona, J. Agric. Food Chem., 1999, 47, 56–60 CrossRef CAS PubMed.
- L. G. Fenoll, P. A. Garcia-Ruiz, R. Varon and F. Garcia-Canovas, J. Agric. Food Chem., 2003, 51, 7781–7787 CrossRef CAS PubMed.
- D. P. Makris and J. T. Rossiter, Food Chem., 2002, 77, 177–185 CrossRef CAS.
- I. Kubo, K. Nihei and K. Shimizu, Bioorg. Med. Chem., 2004, 12, 5343–5347 CrossRef CAS PubMed.
- B. Gasowska-Bajger and H. Wojtasek, J. Agric. Food Chem., 2016, 64, 5417–5427 CrossRef CAS PubMed.
- L. Pourcel, J.-M. Routaboul, V. Cheynier, L. Lepiniec and I. Debeaujon, Trends Plant Sci., 2006, 12, 29–36 CrossRef.
- G. Jungbluth, I. Rühling and W. Ternes, J. Chem. Soc., Perkin Trans. 2, 2000, 1946–1952 RSC.
- J. E. Brown, M. Khodr, R. C. Hider and C. A. Rice-Evans, Biochem. J., 1998, 330, 1173–1178 CrossRef CAS.
- A. Gulsen, B. Turan, D. P. Makris and P. Kefalas, Eur. Food Res. Technol., 2007, 225, 435–441 CrossRef.
- I. Kubo, T. Nitoda and K. Nihei, Molecules, 2007, 12, 1045–1056 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2022 |