Recent advances in visual detection for cancer biomarkers and infectious pathogens
Meng-Meng
Pan
,
Yi-Fan
Wang
,
Le
Wang
,
Xu
Yu
 * and
Li
Xu
*
* and
Li
Xu
*
Tongji School of Pharmacy, HuaZhong University of Science and Technology, Wuhan 430030, China. E-mail: xuyu@hust.edu.cn; xulpharm@mails.tjmu.edu.cn
First published on 3rd November 2020
Abstract
It is an urgency to detect infectious pathogens or cancer biomarkers using rapid, simple, convenient and cost-effective methods in complex biological samples. Many existing approaches (traditional virus culture, ELISA or PCR) for the pathogen and biomarker assays face several challenges in the clinical applications that require lengthy time, sophisticated sample pre-treatment and expensive instruments. Due to the simple and rapid detection manner as well as no requirement of expensive equipment, many visual detection methods have been considered to resolve the aforementioned problems. Meanwhile, various new materials and colorimetric/fluorescent methods have been tried to construct new biosensors for infectious pathogens and biomarkers. However, the recent progress of these aspects is rarely reviewed, especially in terms of integration of new materials, microdevice and detection mechanism into the visual detection systems. Herein, we provide a broad field of view to discuss the recent progress in the visual detection of infectious pathogens and cancer biomarkers along with the detection mechanism, new materials, novel detection methods, special targets as well as multi-functional microdevices and systems. The novel visual approaches for the infectious pathogens and biomarkers, such as bioluminescence resonance energy transfer (BRET), metal-induced metallization and clustered regularly interspaced short palindromic repeats (CRISPR)-based biosensors, are discussed. Additionally, recent advancements in visual assays utilizing various new materials for proteins, nucleic acids, viruses, exosomes and small molecules are comprehensively reviewed. Future perspectives on the visual sensing systems for infectious pathogens and cancers are also proposed.
1. Introduction
Infectious pathogens such as viruses and bacteria have caused many deaths and disastrous effects all over the world and have become one of the major threats to the public health.1 Until July 2020, the outbreak of the pneumonia epidemic (SARS-Cov-2) has infected more than 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400
400![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 people and caused more than 640
000 people and caused more than 640![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths, and the statistics of both are still increasing rapidly. Rapid diagnosis of the highly infectious pathogens is crucial for monitoring the spread of pathogens and reducing the possibility of the large-scale outbreak. The massive screening of potentially infectious people even without any symptoms, adopting some quarantine measures and treatment are significant and can avoid more human infections and prevent the virus pandemic. Meanwhile, the early and accurate diagnosis of cancer can significantly reduce the morbidity and extend the survival time of patients.2,3 In previous studies, 90% of the cancer patients died due to cancer metastasis. From the occurrence of first cancer cell to the cancer cell primary metastasis, the time taken was often about 12 years. During this early stage before the primary metastasis, the majority of cancers might be cured with the effective treatment.4,5 However, it is difficult to diagnose cancer at the primary stage due to no symptoms and imaging features of the small solid tumor. “Liquid biopsy” might be a possible way to diagnose cancer at its early stage by measuring the special cancer biomarkers (circulating tumor cells, exosomes or cell-free DNA) in the whole blood. Consequently, there is a critical requirement for the rapid detection and identification of cancer biomarkers in the complex biological samples for accurate diagnosis of cancer at the primary stage.6–8
000 deaths, and the statistics of both are still increasing rapidly. Rapid diagnosis of the highly infectious pathogens is crucial for monitoring the spread of pathogens and reducing the possibility of the large-scale outbreak. The massive screening of potentially infectious people even without any symptoms, adopting some quarantine measures and treatment are significant and can avoid more human infections and prevent the virus pandemic. Meanwhile, the early and accurate diagnosis of cancer can significantly reduce the morbidity and extend the survival time of patients.2,3 In previous studies, 90% of the cancer patients died due to cancer metastasis. From the occurrence of first cancer cell to the cancer cell primary metastasis, the time taken was often about 12 years. During this early stage before the primary metastasis, the majority of cancers might be cured with the effective treatment.4,5 However, it is difficult to diagnose cancer at the primary stage due to no symptoms and imaging features of the small solid tumor. “Liquid biopsy” might be a possible way to diagnose cancer at its early stage by measuring the special cancer biomarkers (circulating tumor cells, exosomes or cell-free DNA) in the whole blood. Consequently, there is a critical requirement for the rapid detection and identification of cancer biomarkers in the complex biological samples for accurate diagnosis of cancer at the primary stage.6–8
Thus far, many approaches have been developed to detect the infectious viruses9–11 and cancer biomarkers,12–16 from the traditional virus culture, immunological methods such as enzyme-linked immunosorbent assay (ELISA) to molecular diagnostic techniques such as polymerase chain reaction (PCR) assays.17,18 However, these methods face several challenges in the practical applications, such as lengthy operation, low sensitivity, expensive instruments, and sophisticated sample pre-treatment.19 Thus, it is highly urgent to develop fast and sensitive methods to detect the infectious viruses and cancer biomarkers during the early stage of infection and cancer occurrence, so that immediate procedures can be adopted to control virus spread and cancer metastasis.20–22
It is urgent to develop simple and use-friendly microdevices to detect the infectious pathogens and cancer biomarkers, which is significant for massive screening for the fast control of the outbreak of the pathogen infection and/or detect cancer at the early stage. Although, with many efforts, microdevices such as microfluidic chips may be designed and fabricated into a small device that could realize the sample preparation process; however, several detection methods also require large and expensive equipment and are difficult to be miniaturized. Visual detection may provide a possible solution to achieve some of the abovementioned criteria and solve the abovementioned problems and is highly desirable for developing microdevices for the detection of disease-specific biomarkers, especially in the resource-limited areas.23 Among them, visual sensing systems could be achieved using a smartphone or without additional equipment, which have attracted attention due to the characteristics of fast speed, low cost and simplicity. Thus, selection of the proper integration of visual methods and microdevices enables the creation of high-sensitive sensor systems that can meet the need of the society. In the past several years, many rapid and significant sensing systems based on visual methods have been achieved for the detection of infectious pathogens and cancer-related biomarkers. For instance, some researchers tried to integrate the technologies to construct a multiple detection microdevice based on visual assay and signal amplification for the purpose of multiplex detection with high sensitivity. Meanwhile, many systems were flexible for the point-of-care testing, which could be achieved in both laboratory and practical applications, greatly easing the issue of tight resources and abundant samples in the resource-limited areas.
Although several papers reviewed the colorimetric assays in the sensing of infectious pathogens and biomarkers,9,24 most of them focus on one kind of special material for the colorimetric detection or single detection approach, but regardless of the microdevice or sensor system. Few papers reviewed the recent novel advances to construct new sensing systems with some new materials (e.g. nanozymes), new mechanisms (e.g. bioluminescence resonance energy transfer (BRET)),25 enzyme-induced metallization and clustered regularly interspaced short palindromic repeats (CRISPR/Cas-based signal amplification),26 and new systems by integrating the signal amplification technology (e.g. PCR, loop-mediated isothermal amplification (LAMP)) and visual methods (smart phone as a detector) into new flexible devices such as microfluidic chips, paper-based microdevices and/or lateral flow-based test strips.27–29
Herein, we provide a broad view of scope to discuss the recent visual sensing systems from the reaction mechanism, new materials and novel detection methods, and special targets to develop the whole multifunctional microdevices and systems, and focus on high-quality articles on related applications in recent years. We compare the advantages and disadvantages of various detection methods and introduce some new materials and technologies into the visual biosensors, which is significant to develop fast, low-cost, simple operation and sensitive biosensors to meet the clinical requirements. The novel colorimetric and fluorescence visual approaches for the biomarkers, such as metal-induced metallization, BRET, and the CRISPR-based biosensors are also discussed. Recent advancements in visual assays utilizing various novel materials and methods for protein, nucleic acid, exosome, virus and small molecules are comprehensively reviewed, which would provide some help for the researches who will enter the field in the future. In addition, future perspectives and challenges of the visual biosensors are also proposed in this review.
2. Mechanism and methods of visual detection
Generally, three basic principles, namely, immune sandwich assay, fluorescence resonance energy transfer (FRET) and the competitive immunoassay are involved in visual detection.9 Among them, the immune sandwich assay is the most widely applied for the detection of infectious pathogens and cancer biomarkers. To be simple, the infectious pathogens (e.g. virus or bacterial) or the cancer biomarkers are first specially recognized by their corresponding target substances (e.g. antibodies, aptamers, or peptides) on the surfaces of the magnetic beads, paper-based devices or microfluidic chips. After washing with the buffer, the second recognition elements (e.g. enzymes/nanozymes/fluorescent probe-labeled antibodies, aptamers or peptides) are introduced into the system to generate a sandwich complex. The enzymes/nanozymes catalyzed the substrate to obtain the color change for visual detection. For the visual fluorescence assay, the fluorescence was observed under UV light. Various signal amplification approaches could be applied in the visual detection systems in order to enhance the detection signal and get a low detection limit.30,31 Another principle is the FRET, the recognition of the target induced the conformational change of the acceptor and donor, which might change the fluorescence emission and the fluorescence change could be simply observed by the naked eyes or using a smartphone.31 Finally, the competitive immunoassay was sometimes used for the small molecule detection, which is commonly based on the competition between the target and a labeled analyte, which might induce the signal change.32 The visual signal changes are commonly built on the colorimetric or fluorescence change, which could be observed by the naked eyes or by simple image equipment. In order to distinguish the common fluorescence measurement methods in the cuvette, here we mainly focus on the visual fluorescence detection.2.1 Catalytic color reaction by the enzyme or nanozyme
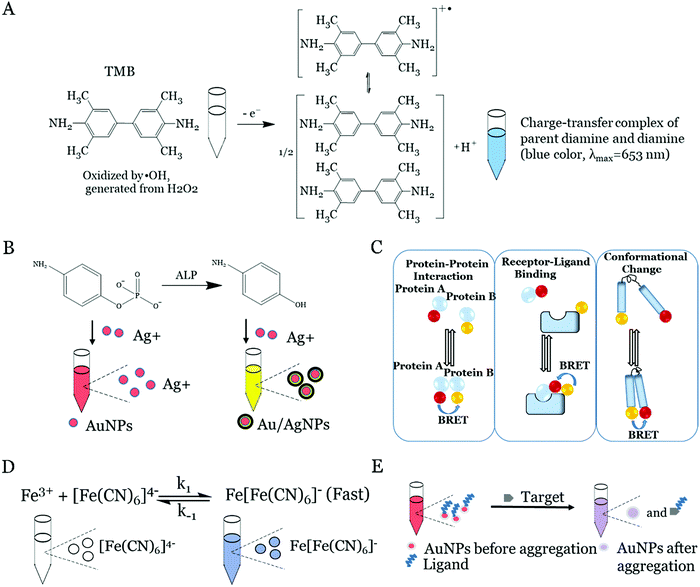
Natural enzymes such as horseradish peroxidase (HRP) are widely used to catalyze the enzymatic oxidation of the colorless substrates to generate colored signals, which can be used to detect the present targets.33 The most commonly used substrates for the catalytic color reaction by enzymes include 3,3′,5,5′-tetramethylbenzidine (TMB), P-phenylenediamine and 3,3′-diaminobenzidine.34,35 In recent years, the iron, carbon, noble metal or metal–organic framework (MOF)-based nanomaterials were found to possess a similar or higher catalytic activity to the natural enzyme. Fig. 1A depicts the enzymatic oxidation of TMB in the presence of the natural enzyme or nanozyme and H2O2 in the system.23 In this catalytic reaction, the nanozyme decomposed the O–O bond in H2O2 to obtain OH˙, and OH˙ oxidized the substrate of TMB to generate a blue colored complex product in a short time. The blue colored product turned yellow after the addition of some acids into the solution. This yellow colored product was a two-electron oxidation product (Fig. 1A), which was very stable in the acid solution.23 The measurement of the signal of the product could determine the targets in the sample as the intensity of the colored product was proportional to the amount of the enzyme in the solution, which was, in turn, proportional to the concentration of the targets in the samples. In addition, the color change of these catalytic reactions can be observed by the naked eyes for visual detection, imaged using a smartphone or quantitatively detected by the UV spectrum.2.2 Enzyme-induced metallization
The metallization-based visual approach for target detection has been developed by combining the enzyme-induced metallization with the preferred metal nanoparticle-induced metal deposition reaction. The overall principle of this strategy is that the enzyme (e.g. alkaline phosphatase (ALP)) could trigger the conversion of inactive substrates (e.g. p-aminophenyl phosphate monohydrate (p-APP)) to active reducing substances (e.g. p-aminophenol (p-AP)). After this, the active reducing substance can induce the (e.g. silver) metallization on the preferred metal nanoparticles (e.g. gold nanoparticles (AuNPs)) to form core/shell nanoparticles, which results in an obvious color change (e.g. red to yellow, Fig. 1B).36,37 The presence of preferred metal nanoparticles can efficiently enhance the metal deposition reaction by the activated reducing substance. Thanks to the enzymatic reaction and the excellently high molar extinction coefficient of the metal nanoparticles (e.g. AuNPs@AgNPs), the ultra-low concentrated targets can be detected in this system. The similar strategies based on the enzyme-induced metallization are successfully employed for the DNA, virus and bacterial detection.36,38,392.3 Fluorescence and bioluminescence resonance energy transfer
The visual fluorescence approaches for target detection can directly record the fluorescence signal intensity of the labelled fluorescent materials (e.g. various fluorescent dyes, semiconductor quantum dots, carbon dots) using a spectrophotometer or camera or observed by the naked eyes under UV light. In most cases, the fluorescence intensity of the labeled tags is proportional to the concentration of the targets, and thus, the intensity of fluorescence can quantify the targets. Meanwhile, several indirect fluorescence visual methods such as fluorescence derivatization, fluorescence quenching and FRET have been developed.40 These mechanisms are based on the change in the fluorescence intensity or comply with the fluorescence emission peak change. For example, in the fluorescence quenching method, the analyte can interact with a fluorescent probe to reduce its fluorescence intensity, thereby using the quantitative relationship between the degree of fluorescence quenching and the concentration of the analyte could achieve a sensitive detection of the targets.Besides, the BRET is applied to construct the visualized biosensors. The BRET is superior to the conventional fluorescence methods in terms of avoiding an external excitation light and phototoxicity, but a high signal-to-noise ratio.41,42 The principle is that luciferase oxidizes luciferin releasing energy in the form of photons. The nonradiative energy transfer would occur when luciferase is brought in close proximity to a suitable acceptor with a favorable geometric orientation (Fig. 1C).42 After this, the bioluminescence intensity is decreased and a new fluorescence emission peak occurs with the increase in fluorescence intensity as the concentration of the target increases. The BRET can also be used to study the protein–protein interaction, receptor–ligand binding and conformational change.
2.4 Metal coordination mechanism
Metal coordination sometimes could produce a color change which might be used to construct the visual detection-based biosensors. The coordination bonds between the metal and the ligand might affect the energy of the electrons before and after complexation. Since the different energies are absorbed by the corresponding metal coordination complex, the color change occurs and is observed. For example, the reaction of the Fe3+ and the ferrocyanide ([FeII(CN)6]4−) could form a blue colloidal solution of Prussian blue. If Fe3+ is excess, the precipitate of the Prussian blue will be produced, as shown in the following equation (Fig. 1D):43| K+ + Fe3+ + [FeII(CN)6]4− → KFeIII [FeII(CN)6] |
| Fe3+ + 3KFeIII [FeII(CN)6] → FeIII [FeIIIFeII(CN)6]3↓ + 3K+ |
Meanwhile, a similar reaction between of Fe2+ and ferricyanide ([FeIII(CN)6]3−) can also generate a blue solution of Prussian blue.44
| K+ + Fe2+ + [FeIII(CN)6]3− → KFeIII [FeII(CN)6] |
The color change could be attributed to Prussian blue that is alternately connected to the cyanide bridge through Fe3+ and Fe2+ to form a mixed valence complex. The transfer of the electrons from Fe3+ to Fe2+ induces the energy change and causes the blue color due to the mixed valence complex absorbing the orange-red light and reflecting the blue color as a result. As the concentration of (Fe(CN)6)4− and Fe3+ or (Fe(CN)6)3− and Fe2+ increases, the visible blue color turned deeper.45,46
2.5 Surface plasmon resonance (SPR) of noble metal nanoparticles
The collective motion of the longitudinal oscillation of free electrons on the metal surface is called plasma oscillation. When the frequency of incident light waves is the same as the plasmon oscillation frequency of the metal surface, there will be SPR and a strong resonance absorption peak or scattering peak will appear on the spectrum.47–49 The resonance frequency is related to the density of electrons, the shape and size, as well as the charge distribution. Therefore, the SPR effect can be adjusted by changing the size or aggregation of the metal nanoparticles. Taking the AuNPs as an example, the SPR peaks of AuNPs are red-shifted as the size increases from 30 to 100 nm. In addition, the nanoparticle aggregation (typically AuNPs or AgNPs aggregation) is also commonly used to generate colorimetric biosensors.50–52 The ligands, e.g. DNA aptamers, are first modified to the nanoparticles, which can protect the nanoparticles from aggregation under a high salt condition. In the presence of targets, the weaker binding equilibrium between the ligands and the nanoparticles is broken, and meanwhile, the stronger binding between the targets and ligands takes place. Under these circumstances, the fast displacement of ligands from the surface of the nanoparticles would result in the aggregation of nanoparticles, thus leading to the color change. Typically for AuNPs in Fig. 1E, the color of the solution is changed from red to blue, which can be monitored by the naked eyes or absorption spectroscopy. We summarized the advantages and disadvantages of the different visual detection methods in Table 1.| Visual detection method | Advantage | Disadvantage | Ref. |
|---|---|---|---|
| Naked eye | No equipment required; low cost; simple operation; convenience for POCT | Low sensitivity; ambient light sources interference; single target detection; semi-quantitative analysis | 68, 70 and 71 |
| UV | High stability; low cost; simple operation | UV equipment; poor sensitivity; single excitation light; single target detection | 59, 67 and 35 |
| Fluorescence | High sensitivity; broad detection range; indirect and direct detection methods; multiplexed detection | Fluorescence equipment; UV light irradiation; autofluoresence | 75 and 77 |
| Bioluminescence | Low background; high sensitivity; no light source needed; convenience for POCT | Low reagents’ stability; FRET condition | 53 and 76 |
3. Visual detection of cancer biomarkers and infectious pathogens
3.1 Visual detection of protein biomarkers
The detection of the low-abundance target proteins in the clinical samples in a fast and low-cost manner is significant for the early cancer diagnosis, which might improve the survival time of the patients.54,55 Although many methods such as electrochemistry, electrochemiluminescence, and surface-enhanced Raman scattering (SERS) have been developed,56,57 most of them need complex sample preparation processes or expensive instruments. The visual sensing systems with portable instruments might solve the problem and be applied to the protein detection.The detection of protein based on colorimetric reactions with natural enzymes still gained much interest among the researchers due to the high catalytic property of the enzyme. For instance, the HRP and Gox were widely used to sensitively detect cancer proteins such as α-fetoprotein (AFP), carcinoembryonic antigen (CEA), and prostate-specific antigen (PSA).58 In order to increase the concentration of enzymes in the detection system, an ultrasensitive visual immunoassay was demonstrated for the human soluble epoxide hydrolase (sEH) protein detection based on self-assembled polymeric horseradish peroxidase (donated as PolyHRP).59 The sandwich ELISA by the PolyHRP as a tracer retained the characteristics of the nature enzyme, i.e. high activity, low cost, and long-term stability, and showed 141-fold higher sensitivity than that of the HRP-labeled anti-hemagglutinin as a tag. The limit of detection (LOD) for the sEH by the PolyHRP reached 0.05 ng mL−1, which was more than 57-fold lower than that of the conventional ELISA by using the HRP labeled anti-HA as a tag.
Although the natural enzyme for the colorimetric reaction has several advantages for the construction of the protein detection biosensors, the expensive cost, low stability and difficulty to produce hamper its applications. In recent years, alternatively, the nanoparticle mimic nanozyme-based colorimetric reaction has been used for the visual detection of protein. The nanoparticle-based nanozymes including iron, carbon, noble metal or MOF-based nanomaterials possess high catalytic activity as natural enzymes, which are widely used for protein detection because of their high stability, low cost, and easy preparation.60–64
Magnetic nanoparticle (Fe3O4) was the first reported and widely applied nanozyme.65 However, the relatively low catalytic activity limits its applications as a highly sensitive biosensor. Various other metal- or carbon-based nanozymes were used to catalyze the colorimetric reaction for the visual detection of proteins such as AuNPs and Pd nanoparticles. Particularly, the porous core–shell nanozymes merited with large specific surface area and synergistic catalytic activity could enhance the detection sensitivity and enlarge the range of visual detection of the proteins. Based on this strategy, Gao et al. synthesized AuNPs coated with ultrathin Pt of sub-10 atomic layers (Au@PtNPs) for the colorimetric detection of proteins in a lateral flow assay manner (LFA).66 The immune sandwich assay was performed on the LFA and the antibody-modified Au@PtNPs were used as a tag for the plasmonic or catalytic detection. The plasmonic and catalytic dual functions of the mimic nanozyme made the Au@PtNPs be applied in different fields such as low-sensitivity model using the plasmonic property or the high-sensitivity model using the catalytic property of the nanozyme. In the high-sensitivity model, this LFA based on the Au@PtNPs could detect the PSA with an LOD of 3.1 pg mL−1, which improved the sensitivity by two orders of magnitude (Fig. 2A).
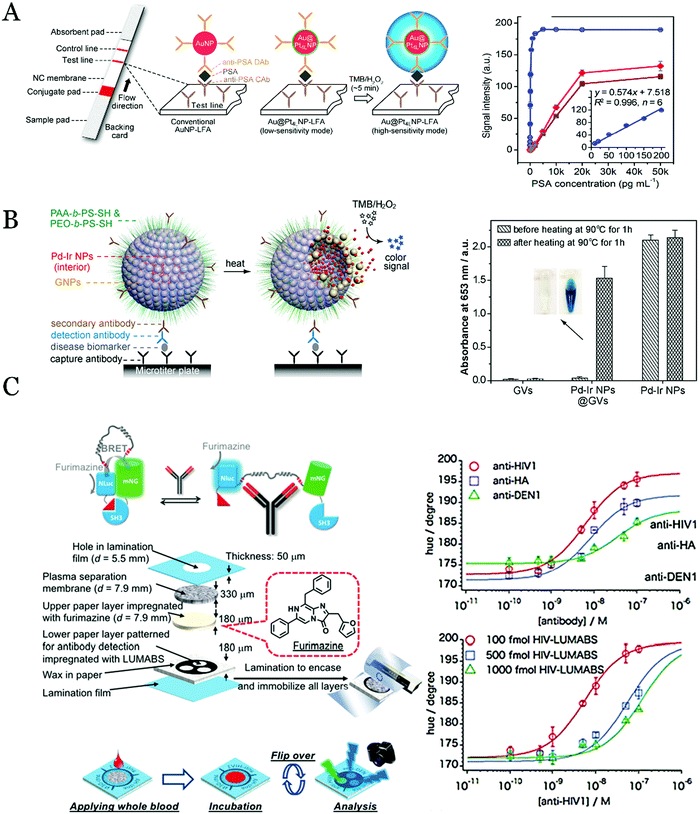 | ||
| Fig. 2 (A) Scheme of the principles of AuNP-LFA and Au@Pt4L NP-LFA for the detection of the target protein at low- (by the observation of the color of AuNPs) and high-sensitivity modes (Au@Pt4L NP-LFA catalyzed the TMB oxidization). Reproduced with permission.66 Copyright 2017, Nano Lett. (B) Schematic illustration of utilizing Pd-Ir NPs@GVs to construct the biosensors for the protein detection. Reproduced with permission.67 Copyright 2017, ACS Nano. (C) Scheme of the BRET detection of three kinds of protein biomarkers on the 3D paper devices. Reproduced with permission.72 Copyright 2018, Angew. Chem., Int. Ed. | ||
In order to enhance sensitivity, increasing the concentration of the nanozymes might be a possible solution. By using this strategy, a signal amplification method for protein detection based on self-assembled gold vesicle encapsulated with Pd-Ir nanoparticles was developed (Fig. 2B).67 The sandwich immunoassay was used to detect the protein, and the gold vesicle encapsulated with Pd–Ir nanoparticles was used to catalyze the colorimetric reaction instead of using the natural enzyme. Thanks to the superior catalytic efficiency of the enzyme mimics and the increased amount of the nanozyme in the detection system, this system showed an enhanced detection sensitivity for the visual assay of protein. Using this method, the LOD for human prostate surface antigen (PSA) was as low as fg mL−1, which shows more than 103 times higher sensitivity than the conventional natural enzyme-based colorimetric assay.
The MOF-coordinated metal ions are also used as mimic nanozymes for the visual detection of proteins.68 The metal ions could increase the catalytic activity of the MOF. Thanks to the large surface area, adjustable porous structure and high efficiency of encapsulation of the MOF, the biosensors using the MOF-metal ions showed high sensitivity for protein detection. In one case, an immunoglobulin antibody-encapsulated MOF nanozyme (RIgG@Cu-MOF) was constructed and used as a label tag for immunoglobulin detection with a visual manner. The RIgG@Cu-MOF was immobilized on a microplate to indicate mouse IgG by forming a double-antibody sandwich complex. The Cu-MOF moiety acted as a catalyst to catalyze the oxidization of TMB to obtain a blue precipitate in the presence of H2O2. The LOD was 0.34 ng mL−1, which was 3 times lower than that using the HRP-labeled IgG as a tag.
On the contrary, some special metal ions might decrease the catalytic activity of the mimic nanozyme, which would be used as an alternative approach to construct the visual biosensors. For example, He and co-workers reported a visual assay for the heparinase based on a ternary system of Hg2+–heparin–osmium nanoparticles (OsNPs).69 In the presence of Hg2+, the oxidase-like activity of heparin–OsNPs to the TMB oxidation was boosted to 76-fold higher than that of using heparin–OsNPs. In this sensor, the heparin on OsNPs was hydrolyzed to small fragments by the heparinase, which weakened the catalytic oxidase activity of Hg2+–heparin–OsNPs. By monitoring the TMB oxidization and the color change, the quantitative determination of heparinase was completed.
Albeit fluorescence detection shows high sensitivity, the requirement of the UV light irradiation and the fluorescent label materials limits its broad applications in the POC testing. Meanwhile, the high background of the auto-fluorescence by the external excitation light might interfere the detection, and the prolonged exposure in the UV light might quench the fluorescence or cause damage to the sample. Herein, the BRET-based visual sensing systems were developed to detect protein biomarkers in recent years. The property of the self-illumination of the bioluminescence reduces the requirement of the UV light irradiation and the background luminescence. Based on this method, a paper-based microfluidic device was used to detect proteins (Fig. 2C).72 The visual detection system combined luciferase (NLuc) with the mNeonGreen fluorescent protein (mNG) to form a BRET complex. In the presence of target proteins (anti-HIV1, anti-HA, and anti-DEN1), the structural change of the complex inhibited the occurrence of the BRET. By measurement of the BRET, the target proteins could be quantified by a simple camera imaging method. These advantages made this approach a great possibility to create a POC platform to detect the infectious pathogens in the resource-limited areas. Further, Li et al.73 established an amplified protein detection strategy by the assembly of the DNA-template BRET system. By using the hybridization chain reaction (HCR) to construct the DNA-template and rationally incorporate luciferase and green fluorescent protein to the DNA template, a highly sensitive method for the detection of human α-thrombin and PSA was developed. The signal of the BRET was observed using a spectrophotometer or a smartphone with the LOD in the pico-molar range. The POC testing based on the BRET possesses high response values, simple steps, and low volume of samples. However, the optical properties of luciferase will be affected in clinical practice while the biomarkers bind to the corresponding antibodies, which limits the wide application of BRET. Therefore, finding the suitable antibodies may resolve the problem of the BRET in the near future.
3.2 Visual detection of nucleic acid biomarkers
The visual fluorescence sensing systems have also been used for RNA biomarker detection by combining the logic gate or new fluorescent dyes. In one case, a DNA nanodevice was constructed to detect the miRNA-21 with the polyethylenimine-modified MnO2 nanosheets and fluorescent nucleic acid probes.75 In this sensor, a double-stranded DNA was attached to the MnO2 nanosheets via the electrostatic interaction with 4-imidazoleacetic acid hydrochloride-modified polyethylenimine on the nanosheets, while a single-stranded DNA was adsorbed onto MnO2 and worked as a fuel for the strand displacement reaction. The triggers of the GSH and miRNA-21 were composed of an ‘AND’ logic gate. The fuel DNA was released by the dissolution of the MnO2 nanosheets with the GSH and then the miRNA-21 triggered toehold-mediated strand displacement reaction to produce the visual fluorescence signal. With the logic gate, the false-positive signals could be efficiently reduced by more than 50%. For the mRNA detection, Luo et al.76 invented a benzothiazole-based fluorescent probe that could turn on the fluorescence when combined with the special RNA fragment containing the G-quadruplex structure of the viral RNA (hepatitis C virus). The probe was constituted by a methylbenzothiazole as an electron acceptor and N,N-diethylaniline as an electron donor. In the presence of the G-quadruplex structure, the fluorescence of probe turned on. This strategy could be used to monitor the individual virus infecting the cells by real-time detection and could observe the subcellular distribution in live cells.
Besides, the FRET strategy was resorted to detect miRNA-122 based on up-conversion nanoparticle (UCNP) and DNA self-assembly.77 The UCNP with captured DNA was applied as the energy donor, and the other TAMRA-labeled DNA was treated as the energy accepter. The sandwich structure was formed in the presence of miRNA-122 and then the FRET occurred. Compared with long-chain miRNA-122, the distance between the energy donor and receptor was reduced by generating a DNA hybridization sandwich complex with two short DNAs to capture miRNA-122, which improved the efficiency of the FRET.
The fast PCR technology can shorten the detection time to tens of minutes, but it still requires the accurate temperature-controlled instruments.81 Recently, various isothermal nucleic acid amplification technologies including in LAMP, HCR, helicase-dependent amplification (HDA), rolling circle amplification (RCA), and recombination polymerase amplification (RPA) have been proposed to overcome the shortcomings of PCR and supposed to build the visual nucleic acid detection approaches.82,83 For example, Lee et al.84 reported a visual method based on LAMP amplification to specifically detect V. parahaemolyticus. The DNA sequences including the HRP-like DNA mimic fragments were combined with the LAMP-amplified products, which inhibited the single-stranded DNA fragment to generate the G-quadruplex structure. The catalytic activity of the HRP-like DNA mimic enzyme to TMB was also inhibited and this could be used to detect V. parahaemolyticus. Suitable complementary sequences and use of HRP enzyme was the key issue in this study, which could avoid the false-positive results.
Except for LAMP, various other isothermal amplification methods such as PRA, HCR and RCA were also adopted to generate visual biosensors for the nucleic acid detection. For example, the RPA was adopted to construct the visual nucleic acid detection method for infectious pathogens.85 In this system, the forward primers with nicking endonuclease recognition regions and G-quadruplex sequence were used for RPA. After the RPA process, the amplicon was cut by Nt. Bst NBI nicking enzyme, G-quadruplex fragments were left off, and the amplicon with nick was extended by polymerases for cycles. Under this strategy, LODs of Salmonella spp. and genetically modified maize MON810 were 4 CFU mL−1 and 0.1% transgenic components, respectively. It should be noted that the TiO2 nanoparticles were used to enhance the RPA technology efficiency by adsorbing the component of RPA and increasing the local concentration.
Besides, Tian and co-workers used the target-triggered HCR strategy for amplification to detect miRNA with a visual manner.86 The miRNA was partially hybridized with the hairpin capture probes (HCP) on Fe3O4 and then the rest part of the DNA was used to initiate HCR amplification to generate a long channel with the nicked double-helix. After that, a novel plane-embedded molecular Cu(II) complex was inserted into the nicked double-helix in the long DNA chain to catalyze the TMB reaction and realize a high sensitive detection of the miRNA by the visual method. In addition, other isothermal amplification methods were also combined with the visual detection for the RNA assay. For instance, a system that integrated zinc finger protein (ZFP)-based BRET and RCA was demonstrated for the detection of miRNAs in the human body fluids (Fig. 3A).87 The blue-emitting nanoLuc luciferase as a donor and green fluorescent protein mNeonGreen as an acceptor were fused with different ZFPs. The hybridization between amplicons of RCA and assistant DNA provided a large number of binding sites of ZFPs for the BRET reaction, which could improve the detection sensitivity.
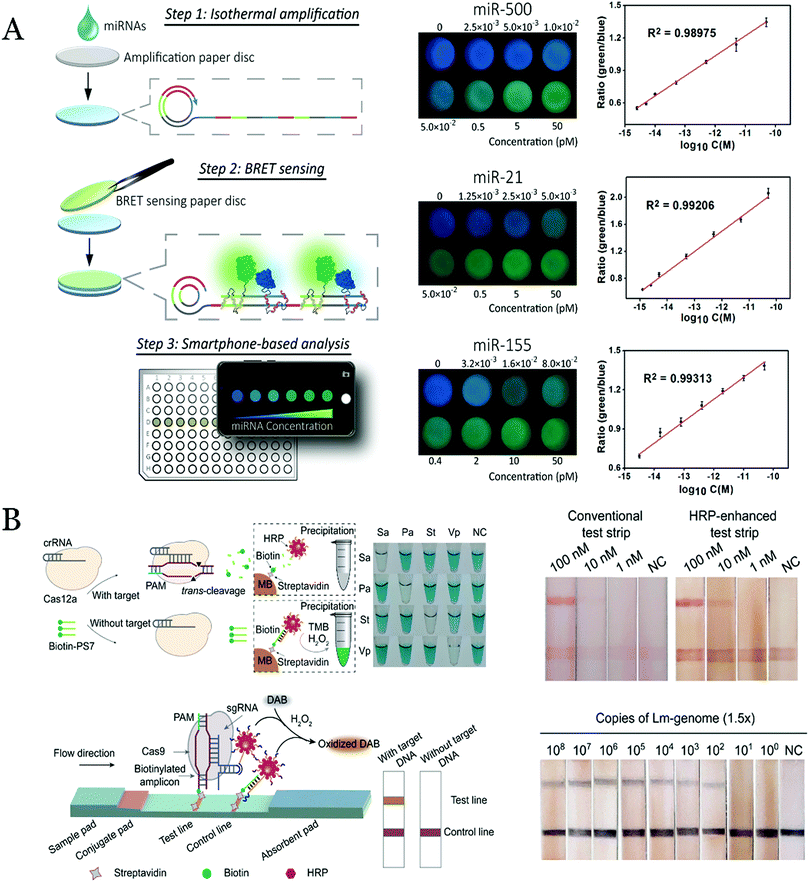 | ||
| Fig. 3 (A) Principle of the paper-based BRET system for the miRNA detection with RCA amplification and smartphone imaging. Reproduced of permission.87 Copyright 2019, Anal. Chem. (B) Principle of the utilization of CRISPR/Cas12a/DNA–HRP–AuNP and CRISPR/Cas9/DNA–HRP–AuNP complexes to develop biosensors for target nucleic acid detection. Reproduced with permission.91 Copyright 2020, J. Am. Chem. Soc. | ||
In order to further improve the detection performance, the cascade amplification technology was applied to build the visual biosensors based on enzyme-induced metallization. In our group, we developed a cascade amplification method to detect the nucleic acid based on the HCR amplification and enzyme-induced metallization.39 The target was captured on the DNA-modified magnetic beads, and then the HCR was processed to amplify the detection signal. By inducing the SA-ALP enzyme into the amplification products, the enzyme could convert the substrate into reduced products. After that, the reduced products induced the Ag shell deposition on the AuNPs core to generate the plasmonic color change from wine red to yellow or black with the increase in concentration of the enzyme. With this cascade amplification technology, the detection of virus nucleic acid with an LOD of 10 pg mL−1 was achieved.
Recently, the novel CRISPR technology has been attempted to develop visual biosensors for nucleic acid detection with signal amplification.88,89 By combining the CRISPR technology, Wang et al.90 successfully developed the sensors for nucleic acid detection with the DNA–AuNP bioprobes. The AuNPs were modified with poly(A) fragments directly to the AuNPs instead of the thiol-modified DNA due to the affinity of the poly(A) to the AuNPs. In one case, the CRISPR/Cas 12a system was activated in the presence of the target DNA, and the non-specific cleavage of single-stranded DNA occurred. After that, the DNA–HRP–AuNPs with biotin modification could not be captured by the streptavidin-modified magnetic beads, and thus, the colorimetric TMB reaction was suppressed due to the lack of the enzyme in the catalysis systems. In another case, the CRISPR/Cas9/DNA–AuNPs complexes were formed by special sequence of DNA–AuNPs complementary to the hairpin structure in CRISPR/Cas9. The complex combined with the targeted DNA modified with biotin, and retained on test line with streptavidin on the surface in LFA (Fig. 3B). With the similar principle, they also construct sensing systems to specifically recognize the targeted DNA of Staphylococcus aureus, Pseudomonas aeruginosa, Salmonella typhimurium, Vibrio parahaemolyticus, and African swine fever virus (ASFV).91
3.3 Visual detection of viruses and exosomes
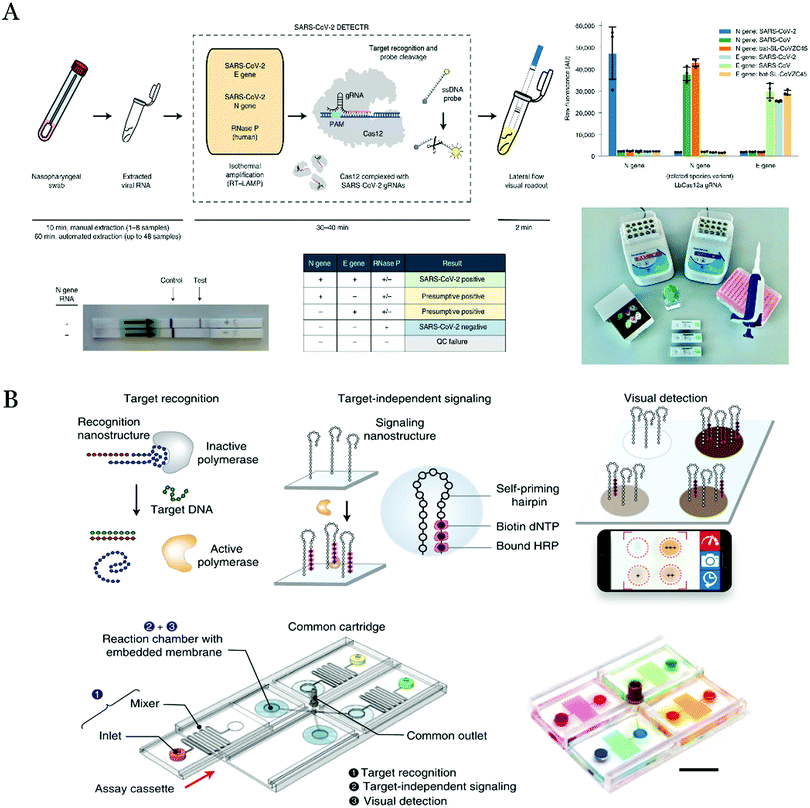 | ||
| Fig. 4 (A) Schematic of the mechanism for the detection of SARS-CoV-2 based on LAMP amplification and CRISPR/Cas12-mediated amplification on the LFA. Reproduced with permission.94 Copyright 2020, Nat. Biotechnol. (B) Scheme of detection of multiple pathogen nucleic acids on the enVision system. Reproduced with permission.96 Copyright 2018, Nat. Commun. | ||
Except for SARS-CoV-2, other highly infectious viruses, such as Ebola virus (EBOV), Zika and HIV are also huge threats to humanity. The effective early diagnosis and treatment can reduce the outbreak and save the lives, especially in the resource-limited areas. Therefore, the simple, rapid and convenient detection systems are urgent demands. For this purpose, an LFA-based test strip combined with a nanosphere (RNs@Au) was developed to rapidly, sensitively, and quickly detect EBOV in resource-limited areas.95 The dual-signal reporter RNs@Au self-assembled the AuNPs on the polymer beads encapsulated with QDs. Thus, the result could be determined by naked eye observation or fluorescence quantification. Furthermore, by inducing the biotin–streptavidin multivalent network to amplify the detection signals, the LOD for the EBOV could reduce 1 order of magnitude. This strategy was simple and successfully applied for the detection of spiked samples by a smartphone.
The massive parallel detection of multiplexed pathogens with limited equipment and samples is also challenged. The combining of the microfluidic chips with visual detection was a brilliant idea. For instance, Ho et al. designed visual identification of nucleic acids (enVision) system for the simultaneous detection of different subtypes of HPV virus genes on a microfluidic chip (Fig. 4B).96 Specifically, a DNA aptamer nanostructure combined with the Taq DNA polymerase which inhibited the activity of the enzyme. In the presence of the target DNA, the DNA hybridization triggered the aptamer release from the aptamer-Taq polymerase enzyme complex and then the enzyme was activated to elongate a self-priming DNA nanostructure for the subsequent visual detection. In this strategy, only the target DNA probe needed to be adapted for targeting different pathogen nucleic acids, which eased to detect various infection pathogens. Moreover, this microfluidic chip might be altered to accommodate more chambers and integrate modules for the sample preparation. This strategy may enhance the current technology, enabling array-type visual detection of multiple biomarkers realized in the clinical diagnosis.
As a new target, the cancer-related exosome can be measured on visual detection-based biosensors. Based on TMB colorimetric reaction, Zhou et al.98 developed a visual sensor with hairpin-like structure for detecting exosomes with the naked eyes. The hemin/G-quadruplex complex acted as a mimic horseradish catalase and was bound to the exo-specific mucin 1 (MUC1) aptamer, which oxidized 2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) in the presence of H2O2 and generated a blue product. Xia et al.99 reported a method that combined tubular nanomaterial with aptamer (specific binding to protein CD63 in exosomes) to detect exosome. The aptamer instead of antibodies on the surface of s-SWCNT enhanced peroxidase activity, thereby improving sensitivity.
To improve the sensitivity, the colorimetric reaction with the HCR amplification was used to detect exosome.100 The exosomes were captured on magnetic beads and the cholesterol-modified DNA initial primer was inserted into the lipid membrane of the captured exosomes. The end of the DNA primer acted as an initiator to induce the HCR amplification. After the amplified products hybridized with the alkaline phosphatase enzyme-modified DNA probe, the enzyme was fixed onto the magnetic beads. The enzyme could convert the substrate to produce a reducing agent of ascorbic acid, and then the silver ions were reduced by ascorbic acid on the Au nanorod and the enzyme-induced metallization induced a plasmonic color change of the solution. The similar idea was reported by combining the HCR amplification and the HRP enzyme-catalyzed TMB reaction to detect the exosomes.101
Besides, the combination of microfluidic chip and enzyme-catalyzed TMB reaction was applied for exosome visual detection.28 In this system, the ZnO nanowire-coated three-dimensional scaffold microfluidic chip was used to enrich the exosomes. The nanowire structure on the chip surface provided a large surface area and a certain exclusion effect, which resolved and significantly promoted the capture of exosomes at high flow rates. Besides the single detection manner, the results could be guaranteed by dual detection methods. For example, the antibody-modified screen-printed electrode microchip was developed to detect the exosomes with two visual and electrochemical method.102 The ferrous oxide nanocubes loaded with Au was used as a signal tag, which could catalyze the TMB to produce the color products, and the products could also be measured by electrochemistry. The linear dynamic range for the exosome was from 103 to 107 exosomes mL−1 with the LOD of 103 exosomes per mL.
In addition, the visual fluorescence approaches were used for the exosome detection. An immuno-biochip modified with antibodies against exosome was reported to specifically capture and detect the tumor-derived exosomes. Lipoplexes modified with a molecular beacon was used to quantify exosome.103 Compared with the conventional PCR testing (100 μL, 24 h), the consumption of reagent volume by the immune-biochip was only 30 μL, and the result could be obtained within 4 h. Similarly, Oh et al.104 designed a visual biosensor based on graphene oxide (GO)-aided molecular beacons to detect the miRNAs in the exosome. This study demonstrated a simple method to detect the inside contents of the exosome, demonstrating the possibility of monitoring the expression of miRNAs in individual cells.
3.4 Visual detection of small molecules
Various small molecules in the human blood were associated with some chronic disease and they should be carefully monitored. For example, reactive oxygen species (ROS) as significant signaling molecules are related with cancer and many other diseases. ROS was considered as second messengers for regulating physiological functions. Specifically, high ROS, such as hypochlorite (OCl−) and hydroxyl radical (˙OH), can directly cause serious damage to living cells by oxidizing nucleic acids, proteins, etc. Therefore, the gold nanoclusters (AuNCs) was facilely synthesized,105 and the surfaces were modulated by quaternary ammonium and oligopeptides for promoting cellular uptake and selectively targeting high ROS. The fluorescence was changed due to high ROS tending to attack the AuNCs. Similarly, Ran and coworkers106 prepared AuNCs for ratiometric sensing of the high ROS with FITC-modified hyaluronic acid as a control signal. The excellent performance was exhibited for imaging high ROS (ClO−, ˙OH and ONOO−) through the aggregation-induced emission properties of AuNCs, dual-emission signal of fluorescence, and high ROS-responsive cleavage of the hyaluronic acid. The LOD for ClO−, ˙OH and ONOO− were 0.2 μM, 0.13 μM and 0.26 μM, respectively. Except for AuNCs, several fluorescent probes could target ClO− in different sites of the cells, such as lysosomes and mitochondria, providing a development for multilocal and multicolor imaging.107Except for the ROS detection, by using the nanozyme, Chen and co-workers108 demonstrated an approach to detect the vitamin B1 (VB1) by using the polyethylene glycol (PEG)-MoO3−x nanozyme with sulfite oxidase activity. Due to the oxygen vacancies in the structure, the developed nanozyme provided good catalytic activity of MoO3−x by PEG modification and an LOD for the VB1 of 0.46 μg mL−1 was achieved.
Visual fluorescence biosensors were also utilized for detecting small molecules. A fluorescent probe by combining the resorufin with 7-nitrobenzofuran was developed to detect cysteine in a paper-based device.109 In the presence of cysteine, the probe reacted with it to cleave the C–O bond, and then resorufin was released. The color of the solution was changed from light yellow to red. Otherwise, a water-soluble smart 19F probe (1-ICG NPs) by the self-assembly of 19F-containing polymers and indocyanine green molecules (ICG, near-infrared absorbing dye) was designed to detect glutathione (GSH) (Fig. 5A).110 The high concentration of the GSH in the tumor microenvironment triggered the cleavage of the S–S bond and turned on the 19F signals. The photothermal treatment induced the 1-ICG NPs dissociating to smaller nanoparticles, which boosted the signals of 19F.
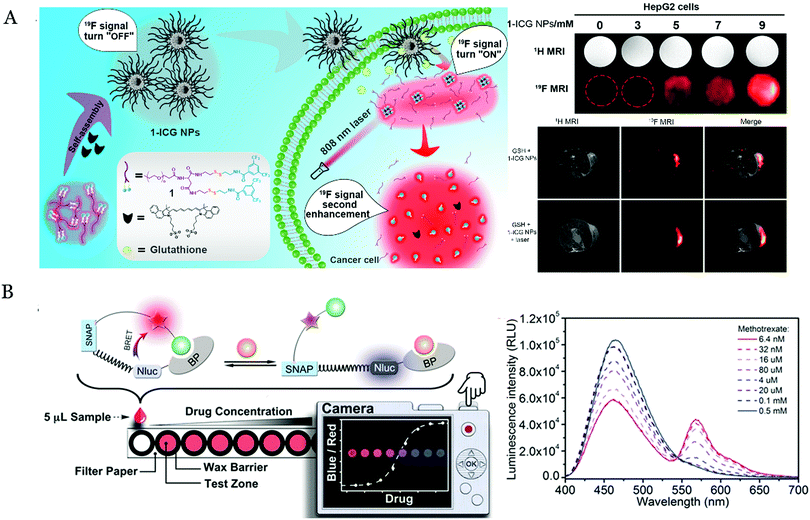 | ||
| Fig. 5 (A) Schematic of the detection of GSH by the novel 1-ICG probes with the cleavage of the s–s bonds and the photothermal treatment. Reproduced with permission.110 Copyright 2019, Nano Lett. (B) Detection of the therapeutic drug molecules based on their corresponding antibodies and the BRET mechanism. Reproduced with permission.112 Copyright 2017, Angew. Chem., Int. Ed. | ||
Recently, the BRET has been adopted to detect small molecules with a highly sensitive and simple manner. For example, a visual biosensor based on BRET was constructed to assess glucose by detecting NADPH involved in the process of glucose oxidation.111 The result was represented by the color of the paper-based biosensor in the digital camera. This method achieved the detection of a variety of related metabolites in the context of limited resources, such as phenylalanine, glucose, and glutamic acid. Xue et al.112 reported a visual sensor of bioluminescence based on antibodies and BRET to detect the drugs of methotrexate, theophylline and quinine. This sensor consisted of the drug-targeted antibodies, nanoLuc luciferases and SNAP-tag-containing fluorophores. In the presence of the competing antigens (drug molecules), the binding of the antigens might destroy the tethered fluorescent competitor (Fig. 5B), thereby disrupting the BRET between the luciferase and the fluorophore. The change in the fluorescence caused by the BRET could be observed using a digital camera. This system might be extended to detect other molecules by changing the specific antibodies. In summary, the abovementioned methods had the possibility to construct the POC testing devices for the small molecule detection in the complex biological samples.113 The simple classification and characteristics of these nanomaterials are summarized in Table 2, and a comparison of different target detections by the visual detection is listed in Table 3.
| The type of nanomaterials | Materials | Colorimetric method | Advantage | Disadvantage | Ref. |
|---|---|---|---|---|---|
| Natural enzyme | HRP; Gox ALP, AchE, lactate dehydrogenase; xanthine oxidoreductase | Catalytic color reaction | High catalytic property; simple operation; mild reaction conditions; naked-eye observation | Expensive; low stability; difficulty to massive produce | 55, 56, 114 and 115 |
| Nanozyme | Iron based; vanadium based; noble metal based; carbon based; MOF based; gold based; copper based | Catalytic color reaction | High stability; low cost; naked-eye detection | Low catalytically efficiency; poor biocompatibility difficult modification or labelling | 58 and 116 |
| Fluorescent materials | Quantum dots; UCNP; metal cluster (Au/Ag/Cu); carbon dots; silicon dots | Fluorescence | Simple operation; fluorescence stability; high sensitivity | UV light irradiation; autofluorescence and high background | 25, 107 and 116 |
| Target | Detection method | Visual mode | Device | Reaction time | LOD | Linear range | Medium | Reference |
|---|---|---|---|---|---|---|---|---|
| a Commercial kit from Shanghai Haling Biotechnology Co., Ltd. | ||||||||
| Tryptophan hydroxylase | Colorimetric | UV | Microcentrifuge tube | 1.5 h | ∼200 μg mL−1 | — | Buffer | Commercial kita |
| L-Ascorbic acid | Colorimetric | UV | Microcentrifuge tube | 1.5 h | — | 0.5–250 μg mL−1 | Buffer | Commercial kita |
| Urinary iodine | Colorimetric | UV | Microcentrifuge tube | 2 h | — | 0–300 μg L−1 | Urine | Commercial kita |
| she | Colorimetric reaction | UV | Microtiter plate | 2 h | 0.05 ng mL−1 | — | PBS | 59 |
| PSA | Colorimetric reaction | UV/Naked eye | Lateral flow assay | 5 min | 3.1 pg mL−1 | 10–200 pg mL−1 | Plasma | 66 |
| PSA | Colorimetric reaction | UV | Microtiter plate | 2 h | 48 pg mL−1 | 0.2–100 ng mL−1 | Plasma | 67 |
| mIgG | Colorimetric reaction | UV | Microtiter plate | 3 h | 0.34 ng mL−1 | 1–100 ng mL−1 | Serum | 68 |
| Heparinase | Colorimetric reaction | UV/Naked eye | Microcentrifuge tube | 20 min | 15 ng mL−1 | 20–1000 ng mL−1 | Serum | 69 |
| Hb | Fluorescence | Fluorescence/Naked eye | Cellphone-based acoustofluidic platform | 2 h | 31 fg mL−1 | 0.2–200 pg mL−1 | Blood | 70 |
| BoNTA/SEB | Fluorescence | Fluorescence/Naked eye | Lateral flow assay | 30 min | 2.52/2.86 pg mL−1 | 1 pg mL−1–100 ng mL−1 | Milk/Juice | 71 |
| Anti-HIV1/anti-HA/anti-DEN1 | BRET | Fluorescence | Paper-based platform | 30 min | 2.8 nM/7.1 nM/19.3 nM | — | Plasma | 72 |
| α-Thrombin/PSA | BRET | Fluorescence | Smartphone | 1.5 h | 12.8 pM/6.4 pM | 14.4 pM–9 nM/11.2 pM–7 nM | Blood | 73 |
| TfR | Colorimetric reaction | UV | Microtiter plate | 1 h | 4 cell | 50–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cell 000 cell |
Buffer | 35 |
| AFP | Colorimetric reaction | UV | — | 1 h | 0.46 ng mL−1 | 0.1–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ng mL−1 000 ng mL−1 |
Serum | 117 |
| HIV p24 | Colorimetric reaction | UV/Naked eye | Lateral flow assay | 20 min | 0.8 pg mL−1 | 100–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 pg mL−1 000 pg mL−1 |
Serum | 118 |
| ACP | Colorimetric reaction (UV) | UV | — | 35 min | 8.3 μU/mL | 0.01–6.0 mU mL−1 | Serum | 119 |
| miRNA-21 | Fluorescence | Fluorescence/Naked eye | Microcentrifuge tube | 4 h | 30 × 10−12 M | 0.09–6.25 × 10−9 M | PBS | 74 |
| Hepatitis C virus | Fluorescence | Fluorescence | — | 2 h | 0.008 μM | 0.008–1 μM | Blood | 75 |
| miRNA-122 | FRET | Fluorescence | — | 8 h | 10−13 M | 0–10−12 M | PBS | 76 |
| BRAFV600E | AuNPs color change | UV | Microcentrifuge tube | 1 h | 500 fM | — | Plasma | 77 |
| Lambda DNA | Fluorescence | Fluorescence/Naked eye | Smartphone | 1 h | 2.8 × 103 copies μL−1 | — | Tris HCl buffer | 80 |
| V. parahaemolyticus. | Colorimetric reaction | UV/Naked eye | Microcentrifuge tube | 1 h | 1 CFU mL−1 | — | PBS | 84 |
| Salmonella | Colorimetric reaction | UV/Naked eye | — | 1 h | 4 CFU mL−1 | 101–105 CFU mL−1 | Buffer | 85 |
| miRNA-21 | Colorimetric reaction | UV | — | 6.5 h | 33 aM | 100 aM–100 nM | Serum | 86 |
| TDNA | Enzyme induced metallization (Au/AgNPs) | UV/Naked eye | — | 2.5 h | 10 pM | 10 pM–1 nM | Blood | 39 |
| miRNA | BRET | Fluorescence | Paper-based device | 2.5 h | 1.7 fM | 2 fM–50 pM | Serum | 87 |
| Salmonella | Colorimetric reaction | UV/Naked eye | — | 1.5 h | 6 CFU mL−1 | 102–107 CFU mL−1 | Yoghurt starter | 120 |
| Listeria monocytogenes | ||||||||
| African swine fever virus | AuNPs aggregation | UV/Naked eye | Lateral flow assay | 1 h | 150 copies μL−1; 200 copies μL−1 | — | Serum | 90 |
| Pseudomonas aeruginosa | ||||||||
| Listeria monocytogenes | Colorimetric reaction | UV/Naked eye | Lateral flow assay | 1 h | 8 ng total RNA; 150 copies μL−1 | — | Serum | 91 |
| SARS-CoV-2 | Fluorescence | Fluorescence | Lateral flow assay | 2 h | 10 copies μL−1 | — | Swab | 94 |
| EBOV | Colorimetric/fluorescence | UV/Fluorescence/Naked eye | Lateral flow assay | 20 min | 0.18 ng mL−1 | 2–1000 ng mL−1 | Urine/Plasma | 95 |
| HPV | Fluorescence | UV/Naked eye | Microfluidic platform | 2 h | 7.205 amol | — | Endocervical brush | 96 |
| Exosome | Colorimetric reaction | UV/Naked eye | — | 1.5 h | 3.94 × 105 particles mL−1 | 8.3 × 105–5.3 × 107 particles mL−1 | Serum | 98 |
| Exosome | Colorimetric reaction | UV/Naked eye | Microcentrifuge tube | 40 min | 5.2 × 105 particles μL−1 | 1.84 × 106–2.21 × 107 particles μL−1 | Serum | 99 |
| Exosome | Enzyme induced metallization (Au/AgNPs) | UV/Naked eye | Microcentrifuge tube | 5 h | 1.6 × 102 particles μL−1 | 1.4 × 103–2.8 × 105 particles μL−1 | Serum | 100 |
| Exosome | Colorimetric reaction | UV/Naked eye | Microcentrifuge tube | 3 h | 2.2 × 103 particles μL−1 | 2.3 × 103–2.3 × 105 particles μL−1 | Serum | 101 |
| Exosome | Colorimetric reaction | UV/Naked eye | ZnO-chip device | 2.5 h | 2.2 × 104 particles μL−1 | 2.2 × 105–2.4 × 107 particles μL−1 | Blood | 28 |
| Exosome | Colorimetric reaction | UV/Naked eye | Microcentrifuge tube | 2 h | 103 particles μL−1 | 103 –107 particles μL−1 | Serum | 102 |
| ROS (˙OH) | Fluorescence | Fluorescence | six-well plates | 1 h | 0.5 μM | 0–20 μM | Living cell | 105 |
| ROS (˙OH) | Fluorescence | Fluorescence | — | 3 h | 0.1 μM | 0.2–100 μM | Living cell | 106 |
| ClO− | Fluorescence | Fluorescence | 96-well plates | 24 h | 0.17 μM | 0–18 μM | Living cell | 107 |
| VB1 | Colorimetric reaction | UV | — | 10 min | 0.46 μg mL−1 | 0.67–202.38 μg mL−1 | Water | 108 |
| Cysteine | Fluorescence | Fluorescence/Naked eye | Paper-based platform | 10 min | 16 nM | 0.04–70.04 μM | Plasma | 109 |
| Methotrexate/theophylline/quinine | BRET | Fluorescence | Camera | 15 min | — | 53 nM–0.13 mM/0.84 mM–0.93 mM/0.49 mM–0.59 mM | Serum/Blood | 112 |
| Glucose/nitrite/protein/phenylpyruvate | Colorimetric reaction | UV | Smartphone | 8/5/8/12 min | 0.33 mM/18.96 μM/3.50 μM/0.32 mM | — | Urine | 113 |
4. Conclusions and perspective
In this review, we summarized the latest advances in biomarker detection technologies based on the visual colorimetric reaction and fluorescence, including protein, nucleic acid, virus, exosome and small molecule detection. The general principle of the visual biosensors is based on the reaction to transfer the direct or indirect event to the visualized signals. Meanwhile, the biosensors based on visual detection could provide excellent access to sensitive signal transduction choices to detect the biomarkers without the need for large detection equipment. In addition, the visual detection can be integrated with new technology that enables the creation of the “next generation biosensors” that might meet the need of clinical applications, such as POC devices for convenient detection, microfluidic chip for multiple infectious pathogen and biomarker detection with limited sample, and CRISPR/Cas-based amplification for enhancing sensitivity.Indeed, there are still many challenges that should be taken into accounts, e.g. how to improve the sensitivity of the visual biosensors. For instance, it is a challenge to detect the cancer at the early stage due to low abundance of the biomarkers in the whole blood. Although “liquid biopsy” obtained a large amount of the financing investment, there are still few of the commercial products. Thus, the fast, low-cost, highly sensitive visual biosensors should be constructed and put forward to the commercially available products. The reproducibility of detection methods and stability of reagents during the store and transportation should be carefully evaluated. Some new reagent storage approaches might be adopted.121 The second challenge is how to improve the visual detection throughput for the multiplexed detection since the single target detection was commonly with a poor specificity. It is much more difficult for the high-throughput and high-resolution detection, for example, to detect >100 kinds of targets simultaneously.122 This challenge might be overcome by combining the special and color barcoding technology as well as multiplexed imaging process.120 Many new technologies such as machine leaning123 may be induced into the visual detection system for the high-throughput and high-resolution detection of the infectious pathogens and the cancer biomarkers. Indeed, the high-throughput and high-resolution visual biosensor might require great efforts and cooperation of the experts from different professional fields. Some targets such as exosomes may have the heterogeneous property,124 therefore, how to selectively capture the targets by new affinity reagents but bypassing the heterogeneous characterization should be taken into account. An additional challenge is the capability of the visual biosensors to accurately and selectively probe the biomarkers in the complex clinical samples, ranging from urine to serum and whole blood. Moreover, in the resource-limited areas, an emerging demand is to develop more simple biosensors that do not rely on sophisticated equipment facilities and provide accurate results in a simple, fast and inexpensive manner.
These problems might be resolved by developing “smart” systems based on visual detection with integrated new technologies, such as some new materials (e.g. nanozymes), new mechanism (e.g. BRET, enzyme-induced metallization and CRISPR/Cas-based signal amplification), and new systems by integrating molecular amplification technologies (e.g. PCR, LAMP) and visual methods (smart phone as the detector) into new flexible devices, such as microfluidic chips, paper-based devices and/or lateral flow-based test strips. These POC testing may provide excellent access to achieve the criteria for the disease-specific biomarker detection. With the continuous efforts, we believe that novel techniques, and various novel materials might be integrated into the visual system and generate next generation sensing systems that solve the existing challenges with state-of-the-art ways.
Abbreviations
| ACP | Acid phosphatase |
| AFP | α-Fetoprotein |
| ALP | Alkaline phosphatase |
| ASFV | African swine fever virus |
| AuNPs | Gold nanoparticles |
| BoNT/A | Botulinum neurotoxin type A |
| BRET | Bioluminescence resonance energy transfer |
| CEA | Carcinoembryonic antigen |
| CRISPR | Clustered regularly interspaced short palindromic repeat |
| EBOV | Ebola virus |
| ELISA | Enzyme-linked immunosorbent assay |
| FRET | Fluorescence resonance energy transfer |
| GO | Graphene oxide |
| GSH | Glutathione |
| Hb | Hemoglobin |
| HCP | Hairpin capture probes |
| HCR | Hybridization chain reaction |
| HDA | Helicase-dependent amplification |
| HRP | Horseradish peroxidase |
| ICG | Indocyanine green |
| LAMP | Loop-mediated isothermal amplification |
| LFA | Lateral flow assay |
| LOD | Limit of detection |
| mNG | mNeonGreen |
| miRNA | microRNA |
| MOF | Metal–organic framework |
| MUC1 | Mucin 1 |
| NADH | Nicotinamide adenine dinucleotide |
| NLuc | Luciferase |
| OsNPs | Osmium nanoparticles |
| p-AP | p-Aminophenol |
| p-APP | p-Aminophenyl phosphate monohydrate |
| PCR | Polymerase chain reaction assay |
| PEG | Polyethylene glycol |
| POC | Point-of-care |
| PolyHRP | Polymeric horseradish peroxidase |
| PSA | Prostate-specific antigen |
| RCA | Rolling circle amplification |
| RPA | Recombination polymerase amplification |
| SEB | Staphylococcal enterotoxin B |
| sEH | Soluble epoxide hydrolase |
| SERS | Surface-enhanced Raman scattering |
| SPR | Surface plasmon resonance |
| TfR | Transferrin receptors |
| TMB | 3,3′,5,5′-Tetramethylbenzidine |
| UCNP | Up-conversion nanoparticle |
| VB1 | Vitamin B1 |
| ZFP | Zinc finger protein |
Conflicts of interest
The authors declare no conflict of interest.Acknowledgements
This work is supported by the National Natural Science Foundation of China (Grant No. 21804105) and by the Fundamental Research Funds for the Central Universities of China (Grant No. 2172019KFYRCPY112, 2020kfyXJJS082).References
- X. Yu, Y. Xia, Y. Tang, W. L. Zhang, Y. T. Yeh, H. Lu and S. Y. Zheng, Small, 2017, 13, 1700425 CrossRef.
- N. I. Lindeman, P. T. Cagle, D. L. Aisner, M. E. Arcila, M. B. Beasley, E. H. Bernicker, C. Colasacco, S. Dacic, F. R. Hirsch, K. Kerr, D. J. Kwiatkowski, M. Ladanyi, J. A. Nowak, L. Sholl, R. Temple-Smolkin, B. Solomon, L. H. Souter, E. Thunnissen, M. S. Tsao, C. B. Ventura, M. W. Wynes and Y. Yatabe, J. Thorac. Oncol., 2018, 13, 323–358 CrossRef CAS.
- X. Liu, Y. Gao, R. Chandrawati and L. Hosta-Rigau, Nanoscale, 2019, 11, 21046–21060 RSC.
- C. A. Klein, Nat. Rev. Cancer, 2009, 9, 302–312 CrossRef CAS.
- X. Yu, N. Wu, F. Chen, J. Wei and Y. Zhao, TrAC, Trends Anal. Chem., 2019, 117, 27–38 CrossRef CAS.
- L. A. Parker, E. Chilet-Rosell, I. Hernandez-Aguado, M. Pastor-Valero, S. Gea and B. Lumbreras, Clin. Chem., 2018, 64, 1657–1667 CrossRef CAS.
- R. Singh, M. D. Mukherjee, G. Sumana, R. K. Gupta, S. Sood and B. D. Malhotra, Sens. Actuators, B, 2014, 197, 385–404 CrossRef CAS.
- Y. Zhang, A. Hu, N. Andini and S. Yang, Biotechnol. Adv., 2019, 37, 476–490 CrossRef CAS.
- Y. Choi, J. H. Hwang and S. Y. Lee, Small Methods, 2018, 2, 1700351 CrossRef.
- Y. Tang, X. Yu, H. Chen and Y. Diao, Biosens. Bioelectron., 2016, 86, 255–261 CrossRef CAS.
- S. M. Yoo and S. Y. Lee, Trends Biotechnol., 2016, 34, 7–25 CrossRef CAS.
- A. Pandey, N. Dhas, P. Deshmukh, C. Caro, P. Patil, M. Luisa García-Martín, B. Padya, A. Nikam, T. Mehta and S. Mutalik, Coord. Chem. Rev., 2020, 409, 213212 CrossRef CAS.
- H. Xu, A. Xia, J. Luo, M. Gao, R. Liao, F. Li, Q. Zhong, W. Zhang, Y. Wang, J. Cui, W. Fu, K. Chang, M. Gan, W. Jiang and M. Chen, Sens. Actuators, B, 2020, 308, 127750 CrossRef CAS.
- L. Gloag, M. Mehdipour, D. Chen, R. D. Tilley and J. J. Gooding, Adv. Mater., 2019, 31, e1904385 CrossRef.
- Y. Y. Lee, S. G. Parker, A. Barfidokht, M. T. Alam, D. B. Walker, B. A. Messerle and J. J. Gooding, Electroanalysis, 2015, 27, 1078–1085 CrossRef CAS.
- M. Sriram, K. Zong, S. R. Vivekchand and J. J. Gooding, Sensors, 2015, 15, 25774–25792 CrossRef CAS.
- H. Aoki, R. M. Corn and B. Matthews, Biosens. Bioelectron., 2019, 142, 111565 CrossRef CAS.
- X. Liu, M. Chen, T. Hou, X. Wang, S. Liu and F. Li, Biosens. Bioelectron., 2014, 54, 598–602 CrossRef CAS.
- S. Slomovic, K. Pardee and J. J. Collins, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 14429–14435 CrossRef CAS.
- J. Liu, Z. Geng, Z. Fan, J. Liu and H. Chen, Biosens. Bioelectron., 2019, 132, 17–37 CrossRef CAS.
- J. Wang, Biosens. Bioelectron., 2006, 21, 1887–1892 CrossRef CAS.
- A. Mohamad, H. Teo, N. A. Keasberry and M. U. Ahmed, Crit. Rev. Biotechnol., 2018, 1–17, DOI:10.1080/07388551.2018.1496063,.
- M. Zhang, J. Ye, J. S. He, F. Zhang, J. Ping, C. Qian and J. Wu, Anal. Chim. Acta, 2020, 1099, 1–15 CrossRef CAS.
- X. Wang and L. Hu, J. Electrochem. Soc., 2020, 167, 037535 CrossRef CAS.
- A. den Hamer, P. Dierickx, R. Arts, J. de Vries, L. Brunsveld and M. Merkx, ACS Sens., 2017, 2, 729–734 CrossRef CAS.
- T. Kong, N. Backes, U. Kalwa, C. Legner, G. J. Phillips and S. Pandey, ACS Sens., 2019, 4, 2638–2645 CrossRef CAS.
- F. Wang, Y. Lu, J. Yang, Y. Chen, W. Jing, L. He and Y. Liu, Analyst, 2017, 142, 3177–3182 RSC.
- Z. Chen, S. B. Cheng, P. Cao, Q. F. Qiu, Y. Chen, M. Xie, Y. Xu and W. H. Huang, Biosens. Bioelectron., 2018, 122, 211–216 CrossRef CAS.
- K. Mao, H. Zhang, Z. Wang, H. Cao, K. Zhang, X. Li and Z. Yang, Biosens. Bioelectron., 2020, 148, 111785 CrossRef CAS.
- Q. Yang, X. Wang, H. Peng, M. Arabi, J. Li, H. Xiong, J. Choo and L. Chen, Sens. Actuators, B, 2020, 302, 127176 CrossRef CAS.
- K. Quan, C. Yi, X. Yang, X. He, J. Huang and K. Wang, TrAC, Trends Anal. Chem., 2020, 124, 115784 CrossRef CAS.
- X. Wang, L. Cohen, J. Wang and D. R. Walt, J. Am. Chem. Soc., 2018, 140, 18132–18139 CrossRef CAS.
- X. Ren, J. Liu, J. Ren, F. Tang and X. Meng, Nanoscale, 2015, 7, 19641–19646 RSC.
- G. Fu, S. T. Sanjay, W. Zhou, R. A. Brekken, R. A. Kirken and X. Li, Anal. Chem., 2018, 90, 5930–5937 CrossRef CAS.
- P. Weerathunge, D. Pooja, M. Singh, H. Kulhari, E. L. H. Mayes, V. Bansal and R. Ramanathan, Sens. Actuators, B, 2019, 297, 126737 CrossRef CAS.
- C. H. Zhou, J. Y. Zhao, D. W. Pang and Z. L. Zhang, Anal. Chem., 2014, 86, 2752–2759 CrossRef CAS.
- S. Xu, W. Ouyang, P. Xie, Y. Lin, B. Qiu, Z. Lin, G. Chen and L. Guo, Anal. Chem., 2017, 89, 1617–1623 CrossRef CAS.
- J. Chen, A. A. Jackson, V. M. Rotello and S. R. Nugen, Small, 2016, 12, 2469–2475 CrossRef CAS.
- X. Yu, Z. L. Zhang and S. Y. Zheng, Biosens. Bioelectron., 2015, 66, 520–526 CrossRef CAS.
- Y. Zhou, J. F. Zhang and J. Yoon, Chem. Rev., 2014, 114, 5511–5571 CrossRef CAS.
- A. B. Dippel, W. A. Anderson, J. H. Park, F. H. Yildiz and M. C. Hammond, ACS Chem. Biol., 2020, 15, 904–914 CrossRef CAS.
- F. Weihs and H. Dacres, TrAC, Trends Anal. Chem., 2019, 116, 61–73 CrossRef CAS.
- M. N. Islam, I. Ahmed, M. I. Anik, M. S. Ferdous and M. S. Khan, Front. Chem., 2018, 6, 496 CrossRef CAS.
- Y. Liu, Z. Zhang, J. Yu, J. Xie and C. M. Li, Anal. Chim. Acta, 2017, 963, 129–135 CrossRef CAS.
- H. Zhang, L. Xue, F. Huang, S. Wang, L. Wang, N. Liu and J. Lin, Biosens. Bioelectron., 2019, 127, 142–149 CrossRef CAS.
- N. Yang, Y. Huang, G. Ding and A. Fan, Anal. Chem., 2019, 91, 4906–4912 CrossRef CAS.
- S. Mariani and M. Minunni, Anal. Bioanal. Chem., 2014, 406, 2303–2323 CrossRef CAS.
- J. F. Masson, ACS Sens., 2017, 2, 16–30 CrossRef CAS.
- H. Yuan, W. Ji, S. Chu, Q. Liu, S. Qian, J. Guang, J. Wang, X. Han, J. F. Masson and W. Peng, ACS Sens., 2019, 4, 704–710 CrossRef CAS.
- C. C. Chang, C. P. Chen, T. H. Wu, C. H. Yang, C. W. Lin and C. Y. Chen, Nanomaterials, 2019, 9, 861 CrossRef CAS.
- M. Sharifi, S. H. Hosseinali, P. Yousefvand, A. Salihi, M. S. Shekha, F. M. Aziz, A. JouyaTalaei, A. Hasan and M. Falahati, Mater. Sci. Eng., C, 2020, 108, 110422 CrossRef CAS.
- M. Wuithschick, A. Birnbaum, S. Witte, M. Sztucki, U. Vainio, N. Pinna, K. Rademann, F. Emmerling, R. Kraehnert and J. Polte, ACS Nano, 2015, 9, 7052–7071 CrossRef CAS.
- M. Endo and T. Ozawa, Int. J. Mol. Sci., 2020, 21, 6538 CrossRef CAS.
- V. Jayanthi, A. B. Das and U. Saxena, Biosens. Bioelectron., 2017, 91, 15–23 CrossRef CAS.
- J. Qin, D. G. Jo, M. Cho and Y. Lee, Biosens. Bioelectron., 2018, 113, 82–87 CrossRef CAS.
- M. Cheng, F. Zhang, A. Zhu, X. Zhang, Y. Wang, X. Zhao, L. Chen, Z. Hua, Y. Zhang and X. Zhang, Sens. Actuators, B, 2020, 309, 127759 CrossRef CAS.
- Y. Jia, Y. Li, S. Zhang, P. Wang, Q. Liu and Y. Dong, Biosens. Bioelectron., 2020, 149, 111842 CrossRef CAS.
- S. Liang, X.-L. Wu, J. Xiong, M.-H. Zong and W.-Y. Lou, Coord. Chem. Rev., 2020, 406, 213149 CrossRef CAS.
- D. Li, Y. Cui, C. Morisseau, S. J. Gee, C. S. Bever, X. Liu, J. Wu, B. D. Hammock and Y. Ying, Anal. Chem., 2017, 89, 6248–6256 CrossRef CAS.
- J. Wu, S. Li and H. Wei, Nanoscale Horiz., 2018, 3, 367–382 RSC.
- J. Wu, X. Wang, Q. Wang, Z. Lou, S. Li, Y. Zhu, L. Qin and H. Wei, Chem. Soc. Rev., 2019, 48, 1004–1076 RSC.
- D. Jiang, D. Ni, Z. T. Rosenkrans, P. Huang, X. Yan and W. Cai, Chem. Soc. Rev., 2019, 48, 3683–3704 RSC.
- H. Wang, K. Wan and X. Shi, Adv. Mater., 2019, 31, e1805368 CrossRef.
- X. Zhang, D. Wu, X. Zhou, Y. Yu, J. Liu, N. Hu, H. Wang, G. Li and Y. Wu, TrAC, Trends Anal. Chem., 2019, 121, 115668 CrossRef CAS.
- L. Gao, J. Zhuang, L. Nie, J. Zhang, Y. Zhang, N. Gu, T. Wang, J. Feng, D. Yang, S. Perrett and X. Yan, Nat. Nanotechnol., 2007, 2, 577–583 CrossRef CAS.
- Z. Gao, H. Ye, D. Tang, J. Tao, S. Habibi, A. Minerick, D. Tang and X. Xia, Nano Lett., 2017, 17, 5572–5579 CrossRef CAS.
- H. Ye, K. Yang, J. Tao, Y. Liu, Q. Zhang, S. Habibi, Z. Nie and X. Xia, ACS Nano, 2017, 11, 2052–2059 CrossRef CAS.
- C. Wang, J. Gao and H. Tan, ACS Appl. Mater. Interfaces, 2018, 10, 25113–25120 CrossRef CAS.
- S. B. He, Q. Q. Zhuang, L. Yang, M. Y. Lin, Y. Kuang, H. P. Peng, H. H. Deng, X. H. Xia and W. Chen, Anal. Chem., 2020, 92, 1635–1642 CrossRef CAS.
- L. Zhang, Z. Tian, H. Bachman, P. Zhang and T. J. Huang, ACS Nano, 2020, 14, 3159–3169 CrossRef CAS.
- C. Wang, R. Xiao, S. Wang, X. Yang, Z. Bai, X. Li, Z. Rong, B. Shen and S. Wang, Biosens. Bioelectron., 2019, 146, 111754 CrossRef CAS.
- K. Tenda, B. van Gerven, R. Arts, Y. Hiruta, M. Merkx and D. Citterio, Angew. Chem., Int. Ed., 2018, 57, 15369–15373 CrossRef CAS.
- Y. Li, P. Yang, N. Lei, Y. Ma, Y. Ji, C. Zhu and Y. Wu, Anal. Chem., 2018, 90, 11495–11502 CrossRef CAS.
- G. Udayan, A. Marsella and P. Valentini, Nanoscale, 2020, 12, 2973–2979 RSC.
- N. Yan, L. Lin, C. Xu, H. Tian and X. Chen, Small, 2019, 15, e1903016 CrossRef.
- X. Luo, B. Xue, G. Feng, J. Zhang, B. Lin, P. Zeng, H. Li, H. Yi, X. L. Zhang, H. Zhu and Z. Nie, J. Am. Chem. Soc., 2019, 141, 5182–5191 CrossRef CAS.
- H. Ren, Z. Long, X. Shen, Y. Zhang, J. Sun, J. Ouyang and N. Na, ACS Appl. Mater. Interfaces, 2018, 10, 25621–25628 CrossRef CAS.
- F. Tian, C. Liu, J. Deng, Z. Han, L. Zhang, Q. Chen and J. Sun, Sci. China: Chem., 2020, 1–9, DOI:10.1007/s11426-020-9800-6,.
- L. Zhang, F. Tian, C. Liu, Q. Feng, T. Ma, Z. Zhao, T. Li, X. Jiang and J. Sun, Lab Chip, 2018, 18, 610–619 RSC.
- V. K. Rajendran, P. Bakthavathsalam, P. L. Bergquist and A. Sunna, Biosens. Bioelectron., 2019, 134, 68–75 CrossRef CAS.
- Y. Chen, C. Qian, C. Liu, H. Shen, Z. Wang, J. Ping, J. Wu and H. Chen, Biosens. Bioelectron., 2020, 153, 112049 CrossRef CAS.
- T. Notomi, H. Okayama, H. Masubuchi, T. Yonekawa, K. Watanabe, N. A. Mino and T. Hase, Chem. Res., 2000, 28, e63 CAS.
- Y. Zhao, F. Chen, Q. Li, L. Wang and C. Fan, Chem. Rev., 2015, 115, 12491–12545 CrossRef CAS.
- J. E. Lee, H. Mun, S. R. Kim, M. G. Kim, J. Y. Chang and W. B. Shim, Biosens. Bioelectron., 2020, 151, 111968 CrossRef CAS.
- J. Tian, H. Chu, Y. Zhang, K. Li, H. Tian, X. Zhang and W. Xu, ACS Appl. Mater. Interfaces, 2019, 11, 46504–46514 CrossRef CAS.
- L. Tian, J. Qi, O. Oderinde, C. Yao, W. Song and Y. Wang, Biosens. Bioelectron., 2018, 110, 110–117 CrossRef CAS.
- Y. Li, L. Zhou, W. Ni, Q. Luo, C. Zhu and Y. Wu, Anal. Chem., 2019, 91, 14838–14841 CrossRef CAS.
- J. S. Chen, E. Ma, L. B. Harrington, M. D. Costa, X. Tian, J. M. Palefsky and J. A. Doudna, Science, 2018, 360, 436–439 CrossRef CAS.
- J. S. Gootenberg, O. O. Abudayyeh, M. J. Kellner, J. Joung, J. J. Collins and F. Zhang, Science, 2018, 360, 439–444 CrossRef CAS.
- X. Wang, E. Xiong, T. Tian, M. Cheng, W. Lin, H. Wang, G. Zhang, J. Sun and X. Zhou, ACS Nano, 2020, 14, 2497–2508 CrossRef CAS.
- M. Hu, C. Yuan, T. Tian, X. Wang, J. Sun, E. Xiong and X. Zhou, J. Am. Chem. Soc., 2020, 142, 7506–7513 CrossRef CAS.
- Y. Xia, Y. Chen, Y. Tang, G. Cheng, X. Yu, H. He, G. Cao, H. Lu, Z. Liu and S. Y. Zheng, ACS Sens., 2019, 4, 3298–3307 CrossRef CAS.
- G. Eom, A. Hwang, H. Kim, S. Yang, D. K. Lee, S. Song, K. Ha, J. Jeong, J. Jung, E. K. Lim and T. Kang, ACS Sens., 2019, 4, 2282–2287 CrossRef CAS.
- J. P. Broughton, X. Deng, G. Yu, C. L. Fasching, V. Servellita, J. Singh, X. Miao, J. A. Streithorst, A. Granados, A. Sotomayor-Gonzalez, K. Zorn, A. Gopez, E. Hsu, W. Gu, S. Miller, C. Y. Pan, H. Guevara, D. A. Wadford, J. S. Chen and C. Y. Chiu, Nat. Biotechnol., 2020, 38, 870–874 CrossRef CAS.
- J. Hu, Y. Z. Jiang, L. L. Wu, Z. Wu, Y. Bi, G. Wong, X. Qiu, J. Chen, D. W. Pang and Z. L. Zhang, Anal. Chem., 2017, 89, 13105–13111 CrossRef CAS.
- N. R. Y. Ho, G. S. Lim, N. R. Sundah, D. Lim, T. P. Loh and H. Shao, Nat. Commun., 2018, 9, 3238 CrossRef.
- S. Wang, A. Khan, R. Huang, S. Ye, K. Di, T. Xiong and Z. Li, Biosens. Bioelectron., 2020, 154, 112056 CrossRef CAS.
- Y. Zhou, H. Xu, H. Wang and B. C. Ye, Analyst, 2019, 145, 107–114 RSC.
- Y. Xia, M. Liu, L. Wang, A. Yan, W. He, M. Chen, J. Lan, J. Xu, L. Guan and J. Chen, Biosens. Bioelectron., 2017, 92, 8–15 CrossRef CAS.
- Y. Zhang, D. Wang, S. Yue, Y. Lu, C. Yang, J. Fang and Z. Xu, ACS Sens., 2019, 4, 3210–3218 CrossRef CAS.
- F. He, H. Liu, X. Guo, B. C. Yin and B. C. Ye, Anal. Chem., 2017, 89, 12968–12975 CrossRef CAS.
- K. Boriachek, M. K. Masud, C. Palma, H. P. Phan, Y. Yamauchi, M. S. A. Hossain, N. T. Nguyen, C. Salomon and M. J. A. Shiddiky, Anal. Chem., 2019, 91, 3827–3834 CrossRef CAS.
- Y. Yang, E. Kannisto, G. Yu, M. E. Reid, S. K. Patnaik and Y. Wu, ACS Appl. Mater. Interfaces, 2018, 10, 43375–43386 CrossRef CAS.
- H. J. Oh, J. Kim, H. Park, S. Chung, D. W. Hwang and D. S. Lee, Biosens. Bioelectron., 2019, 126, 647–656 CrossRef CAS.
- Y. Xie, Y. Xianyu, N. Wang, Z. Yan, Y. Liu, K. Zhu, N. S. Hatzakis and X. Jiang, Adv. Funct. Mater., 2018, 28, 1702026 CrossRef.
- X. Ran, Z. Wang, F. Pu, Z. Liu, J. Ren and X. Qu, Chem. Commun., 2019, 55, 15097–15100 RSC.
- C. Yin, H. Zhu, C. Xie, L. Zhang, P. Chen, Q. Fan, W. Huang and K. Pu, Adv. Funct. Mater., 2017, 27, 1700493 CrossRef.
- Y. Chen, T. Chen, X. Wu and G. Yang, Small, 2019, 15, e1903153 CrossRef.
- J. Zhang, Y. Miao, Z. Cheng, L. Liang, X. Ma and C. Liu, Analyst, 2020, 145, 1878–1884 RSC.
- X. Tang, X. Gong, A. Li, H. Lin, C. Peng, X. Zhang, X. Chen and J. Gao, Nano Lett., 2020, 20, 363–371 CrossRef CAS.
- Q. Yu, L. Xue, J. Hiblot, R. Griss, S. Fabritz, C. Roux, P. A. Binz, D. Haas, J. G. Okun and K. Johnsson, Science, 2018, 361, 1122–1126 CrossRef CAS.
- L. Xue, Q. Yu, R. Griss, A. Schena and K. Johnsson, Angew. Chem., Int. Ed., 2017, 56, 7112–7116 CrossRef CAS.
- X. He, Q. Pei, T. Xu and X. Zhang, Sens. Actuators, B, 2020, 304, 127415 CrossRef CAS.
- M. G. Battelli, M. Bortolotti, L. Polito and A. Bolognesi, Redox Biol., 2019, 21, 101070 CrossRef CAS.
- H. Yin, H. Li, A. Grofe and J. Gao, ACS Catal., 2019, 9, 4236–4246 CrossRef CAS.
- M. Vazquez-Gonzalez, W. C. Liao, R. Cazelles, S. Wang, X. Yu, V. Gutkin and I. Willner, ACS Nano, 2017, 11, 3247–3253 CrossRef CAS.
- F. Ma, C. W. Yuan, J. N. Liu, J. H. Cao and D. Y. Wu, ACS Appl. Mater. Interfaces, 2019, 11, 19902–19912 CrossRef CAS.
- C. N. Loynachan, M. R. Thomas, E. R. Gray, D. A. Richards, J. Kim, B. S. Miller, J. C. Brookes, S. Agarwal, V. Chudasama, R. A. McKendry and M. M. Stevens, ACS Nano, 2018, 12, 279–288 CrossRef CAS.
- C. Chen, W. Liu, P. Ni, Y. Jiang, C. Zhang, B. Wang, J. Li, B. Cao, Y. Lu and W. Chen, ACS Appl. Mater. Interfaces, 2019, 11, 47564–47570 CrossRef CAS.
- Y. Zhang, J. Tian, K. Li, H. Tian and W. Xu, Anal. Chim. Acta, 2019, 1075, 144–151 CrossRef CAS.
- C. Wang, S. Tadepalli, J. Luan, K. K. Liu, J. J. Morrissey, E. D. Kharasch, R. R. Naik and S. Singamaneni, Adv. Mater., 2017, 29, 1604433 CrossRef.
- C. M. Ackerman, C. Myhrvold, S. G. Thakku, C. A. Freije, H. C. Metsky, D. K. Yang, S. H. Ye, C. K. Boehm, T. F. Kosoko-Thoroddsen, J. Kehe, T. G. Nguyen, A. Carter, A. Kulesa, J. R. Barnes, V. G. Dugan, D. T. Hung, P. C. Blainey and P. C. Sabeti, Nature, 2020, 582, 277–282 CrossRef CAS.
- Y.-T. Yeh, K. Gulino, Y. Zhang, A. Sabestien, T.-W. Chou, B. Zhou, Z. Lin, I. Albert, H. Lu, V. Swaminathan, E. Ghedin and M. Terrones, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 895–901 CrossRef CAS.
- K. Lee, K. Fraser, B. Ghaddar, K. Yang, E. Kim, L. Balaj, E. A. Chiocca, X. O. Breakefield, H. Lee and R. Weissleder, ACS Nano, 2018, 12, 494–503 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2021 |