Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Stereoselective synthesis of allele-specific BET inhibitors†
Adam G.
Bond
 ,
Andrea
Testa‡
,
Andrea
Testa‡
 and
Alessio
Ciulli
and
Alessio
Ciulli
 *
*
Division of Biological Chemistry and Drug Discovery, School of Life Sciences, University of Dundee, James Black Centre, Dow Street, Dundee DD1 5EH, UK. E-mail: a.ciulli@dundee.ac.uk
First published on 5th August 2020
Abstract
Developing stereoselective synthetic routes that are efficient and cost-effective allows easy access to biologically active molecules. Our previous syntheses of allele-selective bumped inhibitors of the Bromo and Extra-Terminal (BET) domain proteins, Brd2, Brd3, Brd4 and BrdT, required a wasteful, late-stage alkylation step and expensive chiral separation. To circumvent these limitations, we developed a route based on stereocontrolled alkylation of an N-Pf protected aspartic acid derivative that was used in a divergent, racemisation-free protocol to yield structurally diverse and enantiopure triazolodiazepines. With this approach, we synthesized bumped thienodiazepine-based BET inhibitor, ET-JQ1-OMe, in five steps and 99% ee without the need for chiral chromatography. Exquisite selectivity of ET-JQ1-OMe for Leu-Ala and Leu-Val mutants over wild-type bromodomain was established by isothermal titration calorimetry and X-ray crystallography. Our new approach provides unambiguous chemical evidence for the absolute stereochemistry of the active, allele-specific BET inhibitors and a viable route that will open wider access to this compound class.
Chemical biology and therapeutic development rely on the design or discovery of biologically active compounds that typically contain one or more stereocenters. Amongst the different stereoisomers for a given compound, it is often the case that only one (so-called eutomer) exhibits the desired biological activity, while the other(s) (distomers) may be inactive or have toxic and off-target effects.1 Studying and testing diastereomeric or racemic mixtures has limitations and could lead to unwanted or artefactual results, it is therefore important to develop stereoselective routes and processes which yield solely the desired biologically active molecule.2–5
The four Bromo and Extra-Terminal (BET) proteins, Brd2, Brd3, Brd4 and BrdT, play a crucial role in transcriptional regulation and other processes such as cell proliferation and cell cycle progression.6–8 BET proteins have become attractive therapeutic targets as their misregulation has been linked to diseases such as cancer, neurological disorders and inflammation.9,10 Association to disease has fuelled great interest in the field to develop small molecule BET inhibitors, many of which are in the clinic.11–14 Many BET inhibitors include a triazolodiazepine scaffold, including JQ1 (ref. 15) and I-BET762 (ref. 16) (Fig. 1). Due to the high conservation of BET bromodomains at the acetyl-lysine binding pocket, these inhibitors are pan-selective so do not significantly discriminate between the bromodomains within and across the BET family.17,18 More recently, compounds have been reported to show selective binding to the first bromodomains (BD1), or to the second bromodomains (BD2), of the BET family.19–22 Although these compounds can discriminate between BD1 and BD2 within a given BET protein, they still cannot discriminate across the four BET proteins.
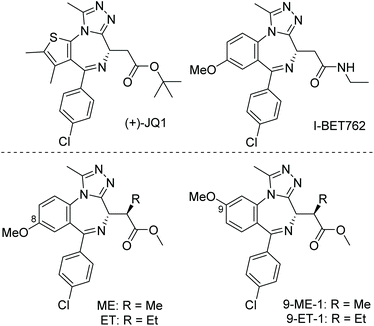 | ||
| Fig. 1 Pan and allele-selective BET inhibitors. Pan-selective inhibitors (+)-JQ1 and I-BET762 (top) and allele selective probes ME, ET, 9-ME-1 and 9-ET-1 (bottom). | ||
To aid individual intra-BET selectivity, we previously developed a chemical genetics approach to engineer orthogonal protein/ligand pairs between the BET proteins and selective inhibitors.23 Our “bump & hole” approach involved the introduction of a single point mutation to the target BET bromodomain by replacing a leucine residue that is conserved across all BET bromodomains, with a smaller residue (e.g. alanine) to generate a “hole”.24 Simultaneously, an alkyl “bump” is incorporated at a diastereotopic, methylene group on the parent scaffold based on the I-BET762 inhibitor, aimed to both complement the size of the “hole” and provide a steric clash to the wild-type protein. This approach led to the generation of allele specific chemical probes ET (ref. 24) and 9-ME-1 (ref. 25) (Fig. 1) targeting the leucine to alanine (L/A) or the less disruptive, leucine to valine (L/V) mutation, respectively. We used this new system to dissect individual roles of BD1 vs. BD2 in Brd4 and the other BET proteins and showed that while the BD1 is necessary and sufficient for chromatin binding, the BD2 plays a role in transcriptional regulation.24,25
Our previous approaches for incorporation of an alkyl “bump” into the I-BET scaffold involved alkylation of a potassium enolate to afford bumped I-BET derivatives with undesired diastereoselectivity and in low yields (Fig. 2A). Epimerisation was required to enhance the amount of desired diastereomer, and further separation by high performance liquid chromatography (HPLC) to isolate the desired isomer. Due to the use of potassium hexamethyldisilazide (KHMDS) for the enolization, and the need of an epimerisation step at such a late stage of the synthesis, the products are racemic and require chiral separation to isolate the eutomers.24–26 Separation of the enantiomers can be costly (up to £1000 for <150 mg of racemate) and leads to loss of material during the separation.
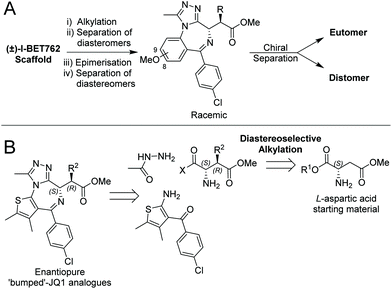 | ||
| Fig. 2 Synthetic routes to bumped BET inhibitors. (A) Previous synthetic strategy for bumped I-BET762 derivatives. (B) Retrosynthetic analysis for enantiopure bumped JQ1 derivatives. | ||
We sought to address these problems by developing a new stereoselective synthetic strategy. We hypothesised that incorporating the “bump” earlier in the synthesis via a diastereoselective alkylation of an aspartic acid derivative would circumvent the limitations of the original route (Fig. 2B). Here, we describe a new synthetic route that allows the preparation of both novel and previously described bumped BET inhibitors stereoselectively in 99% ee. We also provide unambiguous chemical evidence to the absolute stereochemistry of the active allele-specific ligand, previously only assumed based on co-crystal structures.25
Our first efforts to alkylate (+)-JQ1 directly proved unsuccessful, likely due to the steric hindrance caused by the tert-butyl ester. Conversion from the tert-butyl to a methyl ester was required to allow for the introduction of the bump. Alkylation with KHMDS and alkyl iodides proceeded with undesired diastereoselectivity towards the (S,S) diastereomer over the (S,R) diastereomer in overall alkylation yields of approx. 20%. Epimerisation of the major (S,S) isomer with sodium methoxide allows access to the desired (S,R) isomer in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio with the starting (S,S) isomer. However, the use of strong bases during alkylation and additional epimerisation steps may lead to complete racemisation of both stereocentres as previously reported,24–26 (see Scheme S1 in ESI†). With enantiopure (+)-JQ1 costing >$750 per gram, and its derivatives e.g. (+)-JQ1 carboxylic acid being even more expensive, this wasteful approach is not a viable strategy for the preparation of enantiopure bumped JQ1.
1 ratio with the starting (S,S) isomer. However, the use of strong bases during alkylation and additional epimerisation steps may lead to complete racemisation of both stereocentres as previously reported,24–26 (see Scheme S1 in ESI†). With enantiopure (+)-JQ1 costing >$750 per gram, and its derivatives e.g. (+)-JQ1 carboxylic acid being even more expensive, this wasteful approach is not a viable strategy for the preparation of enantiopure bumped JQ1.
To efficiently prepare the desired bumped JQ1 derivatives as single enantiomers, we sought to stereoselectively introduce the “bump” earlier in the synthesis. Our synthetic methodology was based on previously reported stereoselective alkylation of L-aspartic acid diester derivatives.27–30 We so hypothesized that protection of the amino group with both 9-phenyl-9-fluorenyl (Pf) and benzyl (Bn) groups could drive diastereoselective alkylation, with the Pf group providing strong steric hindrance of the β-carbon to the nitrogen while also protecting the α-proton from epimerisation.
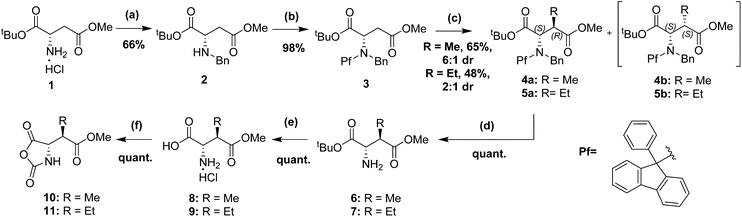
L-Aspartic acid derived diester 1 was first treated with benzaldehyde in dichloromethane (DCM) and the formation of the intermediate imine was monitored by 1H-NMR. Reduction of the imine with sodium borohydride yielded the mono-benzyl protected amine 2. Amine 2 was treated with 9-phenyl-9-fluorenyl bromide, lead(II) nitrate and tribasic potassium phosphate in acetonitrile to form the N-diprotected diester 3. Deprotonation of diester 3 with lithium hexamethyldisilazide (LHMDS) at −78 °C in tetrahydrofuran (THF) afforded the desired E-lithium enolate, which was reacted with methyl iodide at −40 °C over 16 h. This yielded methylated diastereomers 4a (S,R) and 4b (S,S) in a 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio respectively. Ethylated compounds 5a and 5b were prepared in a similar way by deprotonation of diester 3 with LHMDS at −78 °C in THF followed by addition of ethyl iodide and stirring at −78 °C for 16 h. This was left for a further 24 h at −23 °C to yield diastereomers 5a (S,R) and 5b (S,S) in a 2
1 ratio respectively. Ethylated compounds 5a and 5b were prepared in a similar way by deprotonation of diester 3 with LHMDS at −78 °C in THF followed by addition of ethyl iodide and stirring at −78 °C for 16 h. This was left for a further 24 h at −23 °C to yield diastereomers 5a (S,R) and 5b (S,S) in a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio respectively. The choice of LHMDS over the respective potassium base, KHMDS, was motivated by prior findings that switching between these bases can reverse the diastereoselectivity on a similar aspartate derived diester to compound 3.27 Using the potassium base leads to a chelate controlled enolate-ester intermediate which has the opposite geometry to the non-chelated, ‘open’ lithium enolate intermediate and influences facial selectivity to attack by the electrophile (alkyl iodide).
1 ratio respectively. The choice of LHMDS over the respective potassium base, KHMDS, was motivated by prior findings that switching between these bases can reverse the diastereoselectivity on a similar aspartate derived diester to compound 3.27 Using the potassium base leads to a chelate controlled enolate-ester intermediate which has the opposite geometry to the non-chelated, ‘open’ lithium enolate intermediate and influences facial selectivity to attack by the electrophile (alkyl iodide).
Removal of both Pf and Bn groups were performed by hydrogenation of alkylated diesters 4a and 5a with a suspension of 10% palladium on carbon (Pd/C) in acetic acid to give free amines 6 and 7 in high yields. The resulting free amines were dissolved in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of trifluoroacetic acid (TFA) and dichloromethane to achieve the tert-butyl ester deprotection and leave the free amino acid as a TFA salt. The TFA salts were then dissolved in 2 M HCl and freeze-dried to obtain the amino acids 8 and 9 as HCl salts. Conversion of salts proved crucial for the next step as any TFA present led to the formation of trifluoroacetamide by-products. Amino acids 8 and 9 (HCl salts) were treated with triphosgene in THF over 16 h to yield the key N-carboxyanhydrides (NCAs) 10 and 11, as precursors of the alkylated sidechain fragment. These were used in the next steps without the need for further purification (Scheme 1).
1 mixture of trifluoroacetic acid (TFA) and dichloromethane to achieve the tert-butyl ester deprotection and leave the free amino acid as a TFA salt. The TFA salts were then dissolved in 2 M HCl and freeze-dried to obtain the amino acids 8 and 9 as HCl salts. Conversion of salts proved crucial for the next step as any TFA present led to the formation of trifluoroacetamide by-products. Amino acids 8 and 9 (HCl salts) were treated with triphosgene in THF over 16 h to yield the key N-carboxyanhydrides (NCAs) 10 and 11, as precursors of the alkylated sidechain fragment. These were used in the next steps without the need for further purification (Scheme 1).
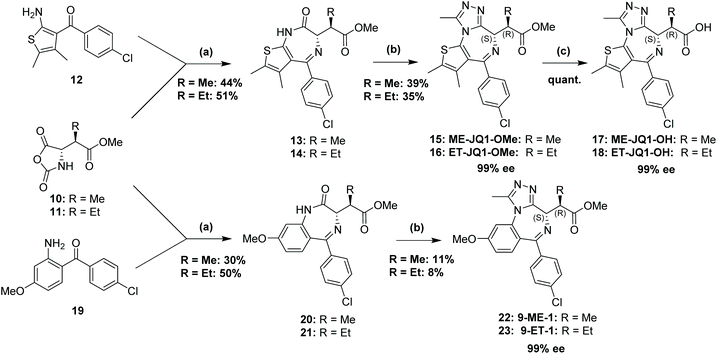
Next, thienodiazepines 13 and 14 were formed in a condensation reaction between NCAs 10 and 11 and amino ketone 12 by heating in the presence of TFA and subsequently triethylamine (TEA) in toluene as reported by Fier et al.31 The use of this methodology for the benzodiazepine ring formation was crucial in our synthetic strategy as it was found to retain the stereochemistry of the amino acid derived NCA. Deprotonation of the lactam in both 13 and 14 with potassium tert-butoxide and addition of diethyl chlorophosphate, as described,32 gave the activated phosphorylimidate intermediate which was not isolated. This was subsequently reacted with acetylhydrazine which led to the formation of the triazole ring in the final compounds 15 (ME-JQ1-OMe) and 16 (ET-JQ1-OMe) with 99% ee determined with supercritical fluid chiral chromatography (Scheme 2, see ESI† for analytical details). Overall, we were able to achieve ∼40 mg of enantiopure product in just five, yield limiting, steps from <£100 worth of starting materials. In comparison, our previous approach required six steps, including expensive chiral purification (∼£1000) and 1 g of JQ1 ($750) to achieve the same amount of pure product.
Quantitative conversion to carboxylic acids 17 (ME-JQ1-OH) and 18 (ET-JQ1-OH) was achieved by treating esters 15 and 16 in a 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of THF to either a 0.54 M or 0.65 M lithium hydroxide (LiOH) solution respectfully. Heating to 45 °C was required for ethyl bumped compound 16 due to the conversion being much slower in comparison to the methyl bumped compound 15. These very mild conditions were essential to avoid epimerisation of the alkylated stereocenter adjacent to the carbonyl group. Using higher concentrations of LiOH and/or higher temperatures resulted in an increased rate of hydrolysis but led to substantial epimerisation to the undesired (S,S) diastereomer. Access to these free acids allows the possibility for further functionalisation (e.g. via amide or ester bond formation).
1 mixture of THF to either a 0.54 M or 0.65 M lithium hydroxide (LiOH) solution respectfully. Heating to 45 °C was required for ethyl bumped compound 16 due to the conversion being much slower in comparison to the methyl bumped compound 15. These very mild conditions were essential to avoid epimerisation of the alkylated stereocenter adjacent to the carbonyl group. Using higher concentrations of LiOH and/or higher temperatures resulted in an increased rate of hydrolysis but led to substantial epimerisation to the undesired (S,S) diastereomer. Access to these free acids allows the possibility for further functionalisation (e.g. via amide or ester bond formation).
Having achieved the novel bumped JQ1 derivatives, we next sought to demonstrate the versatility and scope of our new route by attempting to synthesise the I-BET-based bumped probes, 9-ME-1 and 9-ET-1.25 By using NCA precursors 10 and 11, and treating them with aminobenzophenone 19 in the same condensation reaction as described previously, yielded benzodiazepines 20 and 21. Subsequent triazole ring formation via a similar phosphorylimidate intermediate as described above yielded the final ligands, 22 (9-ME-1) and 23 (9-ET-1), with 99% ee determined with supercritical fluid chiral chromatography (see ESI† for analytical details). We have also demonstrated the accessibility for the (S,S) diastereomer (reported as 16* in the ESI†) of 16 using the minor ethylated diastereomer 5b as the starting point. Stereoselective access to ‘inactive’ stereoisomers provides important negative controls to increase validity and robustness of findings in both biophysical and biological assays.
To further characterise our novel bumped compound, we studied the binding of 16 to Brd4(2) L387A, L387V and wild-type using isothermal titration calorimetry (ITC). We found undetectable/no binding of 16 to the wild-type protein (Fig. 3A, see also ESI Fig. S5†), consistent with ethyl bumped compounds reported previously.24,25 Crucially, 16 demonstrated very high binding affinity to both L/A and L/V mutants with equipotent Kd values of 65 nM (Fig. 3A, see also ESI Fig. S3 and S4†). To validate the binding mode, we solved a high resolution (1.56 Å) X-ray structure of Brd2(2)L383V co-crystallised with 16 (Fig. 3B, see also ESI Fig. S2 and Table S1†). We found that 16 adopts a similar binding mode to 9-ME-1 and 9-ET-1 (see ESI Fig. S1†), positioning the ethyl “bump” towards the “hole” formed by the L/V mutation.
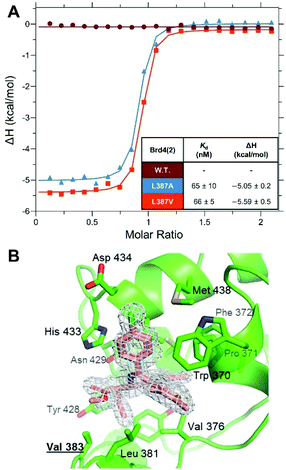 | ||
| Fig. 3 Biophysical and structural characterization of novel JQ1-based bumped inhibitor. (A) ITC titrations of ET-JQ1-OMe (16) against wild-type (W.T.) (maroon), L387A (blue) and L387V (red) constructs of Brd4(2). Table shows values taken as a mean and standard deviation from three replicates. (B) Co-crystal structure of 16 (pink carbons) with Fo-Fc omit map (grey mesh, contour: 3σ) bound to Brd2(2)L383V mutant (green, cartoon representation; green carbons, binding site side chains) (PDB code: 6YTM). Bump and hole residue Val 383 is highlighted. | ||
In summary, we describe a versatile stereoselective approach to successfully synthesise bumped BET inhibitors. We demonstrate scope by synthesizing both novel JQ1 derivatives and previously described I-BET762 derivatives, all in 99% ee. Compared to the previous method, our new route achieves enantiopure products in one less step, from widely available and relatively inexpensive starting materials, while avoiding wasteful, late-stage alkylation steps and chiral separation. We qualified the remarkable allele-selectivity of ET-JQ1-OMe over wild-type and provided unambiguous chemical evidence to the absolute stereochemistry of the eutomer. Access to carboxylic acid derivatives retaining enantiomeric purity enables functionalisation into conjugates such as biotinylated and fluorescent probes and PROTACs,23 which will further expand the scope and utility of this new synthetic strategy for chemical biology investigation.
Abbreviations
| AcNHNH2 | Acetyl hydrazine |
| AcOH | Acetic acid |
| BD1 | First bromodomain |
| BD2 | Second bromodomain |
| BET | Bromo and extra-terminal |
| Brd | Bromodomain |
| DCM | Dichloromethane |
| ITC | Isothermal titration calorimetry |
| KHMDS | Potassium hexamethyldisilazide |
| LHMDS | Lithium hexamethyldisilazide |
| NCA | N-Carboxyanhydride |
| PfBr | Phenylfluorenyl bromide |
| PROTACs | Proteolysis targeting chimeras |
| TFA | Trifluoroacetic acid |
| THF | Tetrahydrofuran |
Author contributions
The manuscript was written through contributions of all authors, and all authors have given approval to the final version of the manuscript.Funding sources
Research reported in this publication was supported by the Medical Research Scotland (PhD studentship 1170-2017 to A.G.B.). The A.C. lab was funded by awards from the UK Biotechnology and Biological Sciences Research Council (BBSRC, grant BB/J001201/2) and the European Research Council (ERC, Starting Grant ERC-2012-StG-311460 DrugE3CRLs). Biophysics and drug discovery activities were supported by Wellcome Trust strategic awards to Dundee (100476/Z/12/Z and 094090/Z/10/Z, respectively).Conflicts of interest
The authors declare the following competing financial interest(s): The Ciulli laboratory receives or has received sponsored research support from Amphista Therapeutics, Boehringer Ingelheim, Eisai, Nurix, and Ono Pharmaceuticals.A.C. is a scientific founder, shareholder, non-executive director and consultant of Amphista Therapeutics, a company that is developing targeted protein degradation therapeutic platforms. The remaining authors report no competing interests.
Acknowledgements
We thank Kwok-Ho Chan for the gift of purified recombinant proteins; Angus Cowan for help with solving the crystal structure; Sarath Ramachandran and Ryan Casement for assisting with ITC and crystallography; Darryl McConnell (Boehringer Ingelheim) for discussions; Reach Separations for chiral analysis; and staff at Diamond Light Source for synchrotron access.References
- L. A. Nguyen, H. He and C. Pham-Huy, Chiral drugs: an overview, Int. J. Biomed. Sci., 2006, 2(2), 85–100 CAS.
- K. C. Nicolaou, T. K. Chakraborty, A. D. Piscopio, N. Minowa and P. Bertinato, Total synthesis of rapamycin, J. Am. Chem. Soc., 1993, 115(10), 4419–4420 CrossRef CAS.
- X. Wu, J. L. Stockdill, P. Wang and S. J. Danishefsky, Total Synthesis of Cyclosporine: Access to N-Methylated Peptides via Isonitrile Coupling Reactions, J. Am. Chem. Soc., 2010, 132(12), 4098–4100 CrossRef CAS PubMed.
- J. Skotnitzki, L. Spessert and P. Knochel, Regio- and Stereoselective Allylic Substitutions of Chiral Secondary Alkylcopper Reagents: Total Synthesis of (+)-Lasiol, (+)-13-Norfaranal, and (+)-Faranal, Angew. Chem., Int. Ed., 2019, 58(5), 1509–1514 CrossRef CAS PubMed.
- K. Mitachi, B. A. Aleiwi, C. M. Schneider, S. Siricilla and M. Kurosu, Stereocontrolled Total Synthesis of Muraymycin D1 Having a Dual Mode of Action against Mycobacterium tuberculosis, J. Am. Chem. Soc., 2016, 138(39), 12975–12980 CrossRef CAS PubMed.
- M. G. Baratta, A. C. Schinzel, Y. Zwang, P. Bandopadhayay, C. Bowman-Colin, J. Kutt, J. Curtis, H. Piao, L. C. Wong, A. L. Kung, R. Beroukhim, J. E. Bradner, R. Drapkin, W. C. Hahn, J. F. Liu and D. M. Livingston, An in-tumor genetic screen reveals that the BET bromodomain protein, BRD4, is a potential therapeutic target in ovarian carcinoma, Proc. Natl. Acad. Sci. U. S. A., 2015, 112(1), 232–237 CrossRef CAS PubMed.
- T. Fujisawa and P. Filippakopoulos, Functions of bromodomain-containing proteins and their roles in homeostasis and cancer, Nat. Rev. Mol. Cell Biol., 2017, 18(4), 246–262 CrossRef CAS PubMed.
- Y. Taniguchi, The Bromodomain and Extra-Terminal Domain (BET) Family: Functional Anatomy of BET Paralogous Proteins, Int. J. Mol. Sci., 2016, 17(11), 1849 CrossRef PubMed.
- A. C. Belkina and G. V. Denis, BET domain co-regulators in obesity, inflammation and cancer, Nat. Rev. Cancer, 2012, 12(7), 465–477 CrossRef CAS PubMed.
- A. G. Cochran, A. R. Conery and R. J. Sims, Bromodomains: a new target class for drug development, Nat. Rev. Drug Discovery, 2019, 18(8), 609–628 CrossRef CAS PubMed.
- C. Massard, J. Soria, A. Stathis, J. Delord, A. Awada, S. Peters, J. Lewin, M. Bekradda, K. Rezai and Z. Zeng, A phase Ib trial with MK-8628/OTX015, a small molecule inhibitor of bromodomain (BRD) and extra-terminal (BET) proteins, in patients with selected advanced solid tumors, Eur. J. Cancer, 2016, 69, S2–S3 CrossRef.
- S. A. Piha-Paul, J. C. Sachdev, M. Barve, P. LoRusso, R. Szmulewitz, S. P. Patel, P. N. Lara, X. Chen, B. Hu, K. J. Freise, D. Modi, A. Sood, J. E. Hutti, J. Wolff and B. H. Neil, First- in-Human Study of Mivebresib (ABBV-075), an Oral Pan-Inhibitor of Bromodomain and Extra Terminal Proteins, in Patients with Relapsed/Refractory Solid Tumors, Clin. Cancer Res., 2019, 25(21), 6309–6319 CrossRef CAS PubMed.
- A. Alqahtani, K. Choucair, M. Ashraf, D. M. Hammouda, A. Alloghbi, T. Khan, N. Senzer and J. Nemunaitis, Bromodomain and extra-terminal motif inhibitors: a review of preclinical and clinical advances in cancer therapy, Future Sci. OA, 2019, 5(3), FSO372 CrossRef PubMed.
- P. Filippakopoulos and S. Knapp, Targeting bromodomains: epigenetic readers of lysine acetylation, Nat. Rev. Drug Discovery, 2014, 13(5), 337–356 CrossRef CAS PubMed.
- P. Filippakopoulos, J. Qi, S. Picaud, Y. Shen, W. B. Smith, O. Fedorov, E. M. Morse, T. Keates, T. T. Hickman, I. Felletar, M. Philpott, S. Munro, M. R. McKeown, Y. Wang, A. L. Christie, N. West, M. J. Cameron, B. Schwartz, T. D. Heightman, N. La Thangue, C. A. French, O. Wiest, A. L. Kung, S. Knapp and J. E. Bradner, Selective inhibition of BET bromodomains, Nature, 2010, 468, 1067–1073 CrossRef CAS PubMed.
- C.-w. Chung, H. Coste, J. H. White, O. Mirguet, J. Wilde, R. L. Gosmini, C. Delves, S. M. Magny, R. Woodward, S. A. Hughes, E. V. Boursier, H. Flynn, A. M. Bouillot, P. Bamborough, J.-M. G. Brusq, F. J. Gellibert, E. J. Jones, A. M. Riou, P. Homes, S. L. Martin, I. J. Uings, J. Toum, C. A. Clément, A.-B. Boullay, R. L. Grimley, F. M. Blandel, R. K. Prinjha, K. Lee, J. Kirilovsky and E. Nicodeme, Discovery and Characterization of Small Molecule Inhibitors of the BET Family Bromodomains, J. Med. Chem., 2011, 54(11), 3827–3838 CrossRef CAS PubMed.
- C. Galdeano and A. Ciulli, Selectivity on-target of bromodomain chemical probes by structure- guided medicinal chemistry and chemical biology, Future Med. Chem., 2016, 8(13), 1655–1680 CrossRef CAS PubMed.
- X. Li, Y. Wu, G. Tian, Y. Jiang, Z. Liu, X. Meng, X. Bao, L. Feng, H. Sun, H. Deng and X. D. Li, Chemical Proteomic Profiling of Bromodomains Enables the Wide-Spectrum Evaluation of Bromodomain Inhibitors in Living Cells, J. Am. Chem. Soc., 2019, 141(29), 11497–11505 CrossRef CAS PubMed.
- K. Cheung, G. Lu, R. Sharma, A. Vincek, R. Zhang, A. N. Plotnikov, F. Zhang, Q. Zhang, Y. Ju, Y. Hu, L. Zhao, X. Han, J. Meslamani, F. Xu, A. Jaganathan, T. Shen, H. Zhu, E. Rusinova, L. Zeng, J. Zhou, J. Yang, L. Peng, M. Ohlmeyer, M. J. Walsh, D. Y. Zhang, H. Xiong and M.-M. Zhou, BET N-terminal bromodomain inhibition selectively blocks Th17 cell differentiation and ameliorates colitis in mice, Proc. Natl. Acad. Sci. U. S. A., 2017, 2952–2957 CrossRef CAS PubMed.
- E. J. Faivre, K. F. McDaniel, D. H. Albert, S. R. Mantena, J. P. Plotnik, D. Wilcox, L. Zhang, M. H. Bui, G. S. Sheppard, L. Wang, V. Sehgal, X. Lin, X. Huang, X. Lu, T. Uziel, P. Hessler, L. T. Lam, R. J. Bellin, G. Mehta, S. Fidanze, J. K. Pratt, D. Liu, L. A. Hasvold, C. Sun, S. C. Panchal, J. J. Nicolette, S. L. Fossey, C. H. Park, K. Longenecker, L. Bigelow, M. Torrent, S. H. Rosenberg, W. M. Kati and Y. Shen, Selective inhibition of the BD2 bromodomain of BET proteins in prostate cancer, Nature, 2020, 578(7794), 306–310 CrossRef CAS PubMed.
- S. Picaud, C. Wells, I. Felletar, D. Brotherton, S. Martin, P. Savitsky, B. Diez-Dacal, M. Philpott, C. Bountra, H. Lingard, O. Fedorov, S. Müller, P. E. Brennan, S. Knapp and P. Filippakopoulos, RVX-208, an inhibitor of BET transcriptional regulators with selectivity for the second bromodomain, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(49), 19754 CrossRef CAS PubMed.
- O. Gilan, I. Rioja, K. Knezevic, M. J. Bell, M. M. Yeung, N. R. Harker, E. Y. N. Lam, C.-w. Chung, P. Bamborough, M. Petretich, M. Urh, S. J. Atkinson, A. K. Bassil, E. J. Roberts, D. Vassiliadis, M. L. Burr, A. G. S. Preston, C. Wellaway, T. Werner, J. R. Gray, A.-M. Michon, T. Gobbetti, V. Kumar, P. E. Soden, A. Haynes, J. Vappiani, D. F. Tough, S. Taylor, S.-J. Dawson, M. Bantscheff, M. Lindon, G. Drewes, E. H. Demont, D. L. Daniels, P. Grandi, R. K. Prinjha and M. A. Dawson, Selective targeting of BD1 and BD2 of the BET proteins in cancer and immunoinflammation, Science, 2020, 368(6489), 387–394 CAS.
- A. C. Runcie, K.-H. Chan, M. Zengerle and A. Ciulli, Chemical genetics approaches for selective intervention in epigenetics, Curr. Opin. Chem. Biol., 2016, 33, 186–194 CrossRef CAS PubMed.
- M. G. J. Baud, E. Lin-Shiao, T. Cardote, C. Tallant, A. Pschibul, K.-H. Chan, M. Zengerle, J. R. Garcia, T. T.-L. Kwan, F. M. Ferguson and A. Ciulli, A bump-and-hole approach to engineer controlled selectivity of BET bromodomain chemical probes, Science, 2014, 346(6209), 638–641 CrossRef CAS PubMed.
- A. C. Runcie, M. Zengerle, K. H. Chan, A. Testa, L. van Beurden, M. G. J. Baud, O. Epemolu, L. C. J. Ellis, K. D. Read, V. Coulthard, A. Brien and A. Ciulli, Optimization of a “bump-and-hole” approach to allele-selective BET bromodomain inhibition, Chem. Sci., 2018, 9(9), 2452–2468 RSC.
- M. G. J. Baud, E. Lin-Shiao, M. Zengerle, C. Tallant and A. Ciulli, New Synthetic Routes to Triazolo-benzodiazepine Analogues: Expanding the Scope of the Bump-and-Hole Approach for Selective Bromo and Extra-Terminal (BET) Bromodomain Inhibition, J. Med. Chem., 2016, 59(4), 1492–1500 CrossRef CAS PubMed.
- J. M. Humphrey, R. J. Bridges, J. A. Hart and A. R. Chamberlin, 2, 3- Pyrrolidinedicarboxylates as neurotransmitter conformer mimics: Enantioselective synthesis via chelation-controlled enolate alkylation, J. Org. Chem., 1994, 59(9), 2467–2472 CrossRef CAS.
- P. J. Dunn, R. Haener and H. Rapoport, Stereoselective synthesis of 2, 3-diamino acids. 2, 3-Diamino-4-phenylbutanoic acid, J. Org. Chem., 1990, 55(17), 5017–5025 CrossRef CAS.
- H. Nishida, T. Eguchi and K. Kakinuma, Amino acid starter unit in the biosynthesis of macrolactam polyketide antitumor antibiotic vicenistatin, Tetrahedron, 2001, 57(39), 8237–8242 CrossRef CAS.
- T. Yoshida, M. Takeshita, H. Orita, N. Kado, S. Yasuda, H. Kato and Y. Itoh, A Large-Scale Preparation of (3S, 4S)-3-(tert-Butoxycarbonyl)amino-4-methylpyrrolidine and Its Analogs from L-Aspartic Acid, Chem. Pharm. Bull., 1996, 44(5), 1128–1131 CrossRef CAS.
- P. S. Fier and A. M. Whittaker, An Atom-Economical Method To Prepare Enantiopure Benzodiazepines with N-Carboxyanhydrides, Org. Lett., 2017, 19(6), 1454–1457 CrossRef CAS PubMed.
- A. Walser, T. Flynn, C. Mason, H. Crowley, C. Maresca, B. Yaremko and M. O'Donnell, Triazolobenzo- and triazolothienodiazepines as potent antagonists of platelet activating factor, J. Med. Chem., 1991, 34(3), 1209–1221 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available: Supplementary results (Fig. S1–S5 and Table S1); Materials and methods section and Scheme S1; supplementary references and NMR spectra. Atomic coordinates and structure factors for the crystal structure of Brd2(BD2)-L383V mutant in complex with ET-JQ1-OMe has been deposited in the protein databank (PDB) under accession number 6YTM. See DOI: 10.1039/d0ob01165g |
| ‡ Present address: Andrea Testa: Amphista Therapeutics Ltd, Bo'Ness Road, Newhouse, ML1 5UH, United Kingdom. |
| This journal is © The Royal Society of Chemistry 2020 |