Bacterial flagella as an osteogenic differentiation nano-promoter†
Dong
Li
a,
Ye
Zhu
a,
Tao
Yang
b,
Mingying
Yang
c and
Chuanbin
Mao
 *a
*a
aDepartment of Chemistry and Biochemistry, Stephenson Life Sciences Research Center, Institute for Biomedical Engineering, Science and Technology, University of Oklahoma, Norman, Oklahoma 73072, USA. E-mail: cbmao@ou.edu
bSchool of Materials Science and Engineering, Zhejiang University, Hangzhou, Zhejiang 310027, P. R. China
cInstitute of Applied Bioresource Research College of Animal Science, Zhejiang University, Yuhangtang Road 866, Hangzhou 310058, China
First published on 12th June 2019
Abstract
Flagella as protein nanofibers (∼14 nm wide) on the surface of swimming bacteria are molecular machines for assisting bacteria to swim in the liquid. They are mainly assembled from protein subunits (FliC) that can be genetically engineered to display peptides. However, so far, no study has been made to show whether flagella with or without displaying peptides could direct stem cell fate. Here we show that flagella detached from bacteria could promote the osteogenic differentiation of bone marrow derived mesenchymal stem cells (BMSCs), and the display of a functional peptide and mineralization of bone mineral (hydroxylapatite, HAP) on the flagella further collectively enhance the promotion effect. The functional peptide is made of two fused amino acid sequences, RGD and E8, which are responsible for promoting cell adhesion onto flagella-bearing substrates and inducing HAP mineralization on flagella from an HAP-supersaturated solution, respectively. Our work shows that the unique nanotopography and surface chemistry of both mineralized and non-mineralized flagella enable them to present physical and chemical cues favoring the osteogenic differentiation of stem cells. Thus flagella are nanofibrous osteogenic differentiation promoters that can be used to build extracellular matrix-like materials.
New conceptsStem cells need to be specifically differentiated into bone forming cells (i.e., osteoblasts) for successful bone regeneration. However, stem cells naturally differentiate into multiple cells randomly. Thus a nanostructured material is needed to promote their differentiation into osteoblasts. Currently reported nanostructured materials usually do not bear the possibility of genetic modification that is desired in precisely displaying a signaling peptide. Here for the first time, we discover that protein nanofibers used as a molecular machine to enable bacteria to swim, called flagella, could promote the differentiation of stem cells into osteoblasts. Flagella are helically assembled from genetically modifiable protein subunits. When the flagella are genetically modified to bear cell-signaling molecules or chemically modified with bone minerals, the flagella become more capable of promoting the differentiation. Since bacteria naturally grow flagella on their surface, which can be purified, the discovery of flagella being a differentiation nano-promoter provides a cost-effective new nanomaterial that can either fill the bone defects as a scaffold or modify the implant surface as a film to enhance bone regeneration to repair bone defects. This work also suggests that the flagella can be used to control the stem cell differentiation due to their unique morphology and surface chemistry. |
Introduction
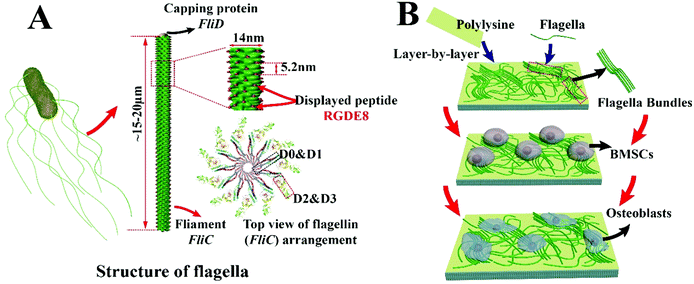
Due to their versatile chemistry, molecular recognition properties and biocompatibility, peptide and protein based nanomaterials gained much attention in directing cell fates.1–6 Nanostructured biomaterials with cell-favorable surface properties promote cell fates since cells directly interact with the nanostructured extra-cellular matrix (ECM) in the development of tissues such as bone.2 Many biomolecular materials, such as collagen, chitosan, silk proteins, peptide amphiphiles and bacteriophages,7–12 have been employed for the synthesis of bone-like ECM. Bone ECM is not a simple mixture of organic (mainly collagen) and inorganic (mainly hydroxylapatite, HAP) phases but has a unique spatial relation with respect to each other. At nanometer level, collagen molecules self-assemble into parallel collagen fibrils and HAP nanocrystals are nucleated along the collagen fibrils.13 Thereafter, use of a protein nanofiber to nucleate HAP is a feasible approach to the formation of an HAP-mineralized collagen fibril-like nanofiber.14 The synthesis should take place under mild conditions such as at room temperature and in aqueous solution.In this study, a naturally occurring protein nanofiber, bacterial flagellum, which is orderly assembled from several thousand copies of monomers called flagellin (FliC), is employed for the display of functional peptide and nucleation of HAP nanocrystals on the surface. FliC is synthesized inside bacteria and then assembled on the surface of bacteria into flagella. The resultant flagella can also be detached from bacteria by simple vortexing and isolated by ultracentrifugation, allowing us to genetically engineer and physically purify flagella. The flagellar nanofiber has an outer and inner diameter of ∼14 and 2 nm, respectively, and a length up to several microns.15,16 The N- and C-terminal domains of FliC (D0 and D1) are highly conserved; on the contrary, the central regions (D2 and D3) are hyper variable and thus can be modified by the insertion or deletion of sequences without losing its self-assembly properties (Scheme 1A).17 Here a peptide, RGDEEEEEEEE (RGDE8), fused from two peptide motifs, the integrin-binding motif Arg-Gly-Asp (RGD) as a cell-adhesion peptide (commonly seen in collagen and other ECM proteins) and 8 contiguous Glu residues (E8, which is an HAP-nucleating domain in a non-collagenous bone protein called bone sialoprotein),18 was displayed on the hyper variable solvent-exposed exterior surface of flagella. Then we used the RGDE8-displayed flagella to induce HAP mineralization from an HAP-supersaturated solution and form a mineralized nanofibrous matrix mimicking some aspects of the bone ECM. We discovered that the flagella-bearing matrix promoted the adhesion, proliferation and osteogenic differentiation of bone marrow derived stem cells (BMSCs) for the first time (Scheme 1B).
Results and discussions
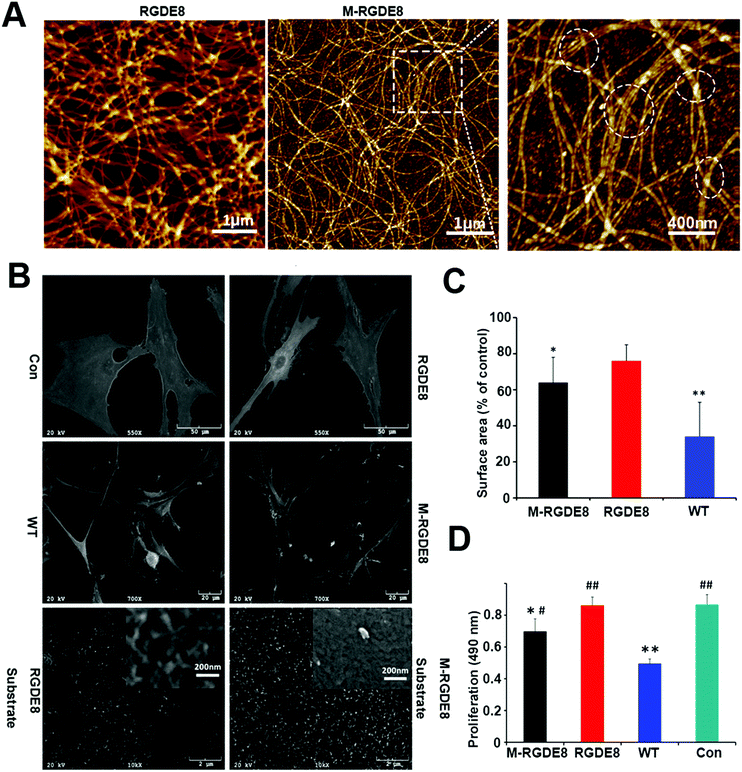
The distance between adjacent FliC subunits on the highly ordered surface of flagella is about 2.6 nm,15 resulting in the similar periodicity of the bioactive RGD motif on the flagellar surface introduced by flagellar display (Scheme 1A). A high density of about 3.43 × 104 epitopes per μm2 can thus be constructed on the flagellar surface. Surface topography on the material surface especially nanotopography is another important factor that can trigger signaling cascades to modulate cell behaviour.2,7 Thus we studied the synergistic effect of nanotopography (contributed by mineralization and assembly of flagellar filaments) and surface chemistry (due to the peptide display and mineral formation on the flagellar surface) on the proliferation and osteogenic differentiation of BMSCs.Peptides and proteins are important in controlling the biomineralization of hard tissue. The negatively charged amino acids, especially the E8, are known to be the main factor to attract calcium ions followed by HAP formation.18,19 Biomimetic mineralization has been considered one of the promising approaches for the fabrication of inorganic nanoparticles20 and bone-like biomaterials.21 During the flagella production process, RGDE8-displayed flagellins expressed in the bacteria were transported to the outer surface of the bacteria and in situ assembled into RGDE8-displayed flagella on the surface of bacteria. Then the bacteria were vortexed to detach the flagella from them and the flagella were further purified from the resultant mixture through centrifugation. The RGDE8-displayed flagella were then allowed to be mineralized in a 4 mM supersaturated HAP solution (prepared following our previous protocol22,23) for 6 days at room temperature. We found that the bioengineered flagella could mediate the nucleation of HAP nanocrystals (Fig. 1A and 2). The biomineralized RGDE8 flagella were deposited on the polylysine substrate by a layer-by-layer (LBL) assembly technique. Purified RGDE8 flagella (4.264 μg μl−1) exhibited a characteristic curly morphology when air-dried on the polylysine substrate (Fig. 1A, left). After HAP mineralization, RGDE8-displaying flagellar filaments were coated by a layer of mineral and became relatively “straight” (Fig. 1A, middle). Interestingly, at some areas, the mineralized flagella are assembled into parallel bundle-like structures (Fig. 1A and 2), mimicking some key structural features of mineralized collagen fibrils. In natural bone, the mineralized collagen fibrils are also parallel-aligned to generate organized nanostructures.
We then studied the stem cell behaviors on four types of substrates, including RGDE8-displaying flagella (RGDE8), biomineralized RGDE8 flagella (M-RGDE8), wild-type flagella (WT), and polylysine as the control (Con). After being seeded on the substrates for 24 h, BMSCs showed a characteristic fibroblastic morphology (bipolar to polygonal). They spread better on RGDE8 and M-RGDE8 flagella than on WT flagella (Fig. 1C and Fig. S1, ESI†). On WT flagella, BMSCs spread much less and some of the cells exhibited a round-like morphology, indicating less favorable adhesion due to the lack of RGD. However, BMSCs could still adhere and remain viable. As a well-known integrin-binding peptide, RGD is ubiquitous in many ECM proteins (e.g., collagen, fibronectin, and vitronectin) that can promote cell attachment and adhesion.24 Our data show that the display of RGD peptide on the flagella highly improved the cell spreading on their surface.
Scanning Electron Microscopy (SEM) also revealed a similar morphology of BMSCs on the different substrates (Fig. 1B). The cells were flat and stretched on the polylysine control substrates but spread less on the flagella coated substrates, especially on WT flagella. Consequently, the surface area of BMSCs decreased on the flagella-coated substrates (Fig. 1C). This is consistent with previous report of cellular morphologies on the nano-fibrous scaffolds.25 The cells were attached on the flagella-coated substrates via filopodia-like extensions and exhibited more and longer ones on the M-RGDE8 flagella-coated substrates (Fig. 1B and Fig. S2, ESI†). The increased surface roughness and structural complexity after mineralization on the flagella could induce more and longer filopodia-like extensions. More compact but rough surfaces were observed on the M-RGDE8 flagella-coated substrates due to the nucleation of HAP nanocrystals (Fig. 1B, inset), which is consistent with AFM observation (Fig. 1A).
The growth rate of BMSCs was significantly altered due to the peptide display and mineralization on the flagella, as well as the unique nanotopography of the flagella (Fig. 1D). Significant higher growth rates of the cells were observed on the RGDE8-displaying flagella but slightly decreased after biomineralization. The decreased proliferation rates of BMSCs were also observed on the biomimetic mineralized collagen membranes with nanostructures.26 As a highly osteoconductive material, HAP promotes cell differentiation and is often used in hard tissue regeneration.27–29 After biomineralization of flagella, a high proportion of cells were induced to undergo a differentiation pathway, and thus proliferation rates of the cells were limited.30 Another possibility is that some of the RGD epitopes on the flagellar surface may be embedded underneath mineral layers and become unavailable to the cells. The proliferation rates of cells were significantly limited on the WT flagella-coated substrates due to the lack of bioactive signals. In another word, the display of RGD ligand with a high density on the flagella highly improved the biocompatibility of flagella-coated substrates. Because polylysine can promote cell attachment and proliferation by altering the negatively charged surface to positive,31 as expected, we observed a higher growth rate of BMSCs on polylysine surface.
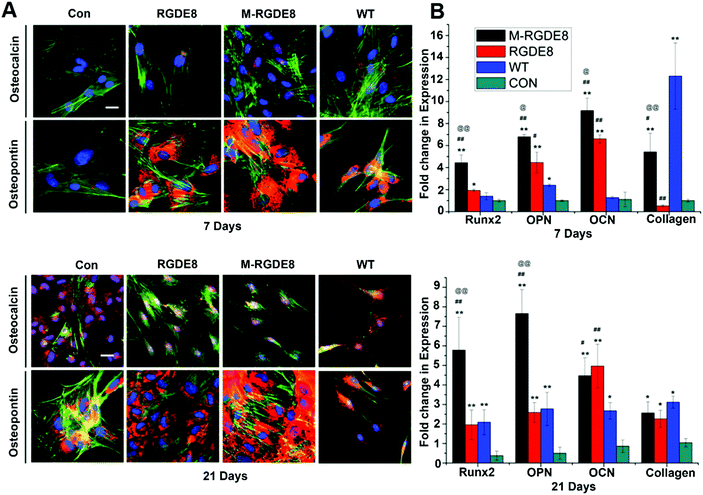
To determine BMSCs’ differentiation towards osteoblasts on the flagella based substrates in osteogenic medium, the production of two bone-specific marker proteins, including osteopontin (OPN) and osteocalcin (OCN), were monitored by immunofluorescence (Fig. 3A). It has been found that the surface chemistry and nanotopography can direct both the initial cell adhesion and the differentiation signaling pathways, depending on the organization and density of functional groups on the substrate.1,7 Peptide display enables precise control of flagella functionality through homogeneous surface modification with multivalent ligands and nanoscale distance.17 On both day 7 and day 21, the expression of OPN for different groups was found to be in the order of M-RGDE8 > RGDE8 > WT > Con. OCN showed a similar trend although it is not as obvious as OPN. Overall the immunofluorescence data showed that WT flagella could promote osteogenic differentiation and sequential addition of RGDE8 (by genetic display) and HAP (by mineralization) onto WT flagella further enhanced such promotion effect.
The effects of surface chemistry and nanotopography of flagella on the osteogenic differentiation of cells were also measured by real-time quantitative PCR (RT-qPCR) analysis. The gene expression levels of selected osteogenesis specific gene markers (OCN, OPN, and Runx2) and non-specific gene marker (type I collagen) were detected at different time points during osteogenic differentiation for all flagella-bearing substrates (Fig. 3B), suggesting that flagella could promote osteogenic differentiation. Because the differentiation markers almost had no expression on day 1, gene expression of the cells for each marker was normalized to that on polylysine coated substrate on day 7. On day 7, the expression of the four osteogenesis specific gene markers showed the order of M-RGDE8 > RGDE8 > WT > Con, consistent with the immunefluorescence characterization of the OCN and OPN markers at the protein level (Fig. 3A). These results indicated that the differentiation of BMSCs toward osteoblasts was accelerated on the RGDE8 flagella compared to the WT flagella. RGD ligands with nanoscale distribution and lower spacing were known to enhance osteoblast proliferation and differentiation.24,32 Apparently, the high density of RGD ligands with nanoscale spacing on the flagellar surface highly promoted early expression of osteogenic marker genes. Due to the biomeneralization of RGDE8 flagella, the nucleated HAP nanocrystals on the flagellar surface further promoted early expression of the osteogenic marker genes. After BMSCs were cultured for 21 days, the M-RGDE8 still kept a significantly higher level of osteogenesis-specific genes (OPN and Runx2) than the other groups. The level of the osteogenesis-specific genes (OPN and Runx2) on the RGDE8 and WT flagella substrates, though lower than that of the M-RGDE8 flagella substrate, is significantly higher than the control polylysine substrate. For OCN, M-RGDE8 and RGDE8 presented a significantly higher level than WT flagella, which exhibited a significantly higher level than the control polylysine substrate. As for the non-specific marker (type I collagen), the flagella-containing groups demonstrated a higher level than the control polylysine substrate. It is well known that the polylysine substrates could improve osteogenesis.31 Our flagella-bearing matrix showed even higher capability in promoting osteogenesis.
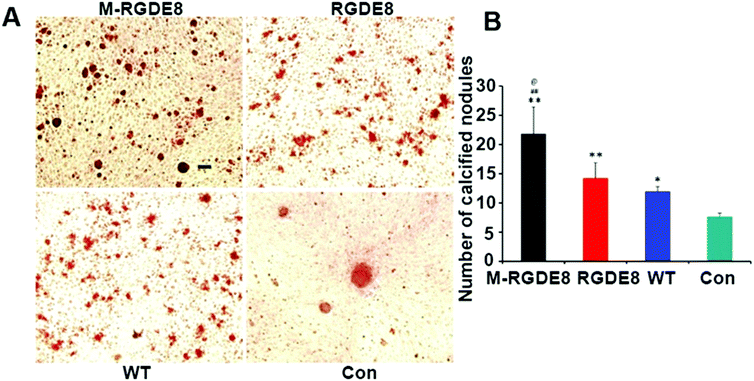
Calcium-containing mineral deposits are also important markers of osteogenic differentiation. The self-mineralized calcium deposits represent the final stages of osteogenic differentiation. Alizarin red stain was used to observe the presence of calcium-containing minerals. The highest density of calcified nodule-like structures in M-RGDE8 indicated that calcium deposition was significantly enhanced in M-RGDE8 (Fig. 4A). The calcium-containing mineral deposits were also detected on RGDE8 and WT with RGDE8 bearing more deposits than WT. On control substrate, however, more fibroblast-like cells were detected. Quantitative analysis also revealed extracellular calcium deposits (nodules of mineralization) were highly increased on flagella scaffolds with an order of M-RGDE8 > RGDE8 > WT > Con. After mineralization, a much higher number of calcified nodule-like structures were detected (Fig. 4B). The results from Alizarin red stain assay are generally consistent with those from the RT-qPCR and immunefluorescence assay. Collectively, they show WT flagella could promote osteogenic differentiation of BMSCs as compared to the substrates without flagella. Moreover, when RGDE8 was displayed on the flagella, the resultant substrates further enhanced the promotion effect. Eventually, when the HAP minerals are coated on the RGDE8 flagella by biomimetic mineralization, the resultant M-RGDE8 flagella more significantly enhanced the promotion effect than the RGDE8 flagella.
In addition to the surface chemistry, the nanotopography due to the filamentous nature and mineralization of flagella should also get involved in regulating cell behavior such as the cell morphology and anchorage, proliferation and differentiation. The less spreading of BMSCs on the flagella-bearing substrates than on the polylysine substrates could also be attributed to the nanotopography on the nanofibrous surfaces.25 Moreover, the expression of osteogenic marker genes is sensitive to surface nanotopography. The topography of nanostructures had significant effects on osteogenesis of human mesenchymal stem cell (hMSCs) by triggering speedy expression of osteo-specific markers and enhancing osteogenic differentiation.2,33 The nanostructured scaffolds could absorb more proteins such as fibronectin from cell culture media and, in turn, promoted up-regulated gene expression and increased mineralization.21 Compared to the control non-flagella substrates, up-regulated expression of osteogenic specific markers was seen on the flagella substrates during the entire period of study. This indicated that the nanotopography of flagellar substrate should also accelerate BMSCs differentiation toward osteoblastic lineage.
The flagella are chemically stable at various conditions (e.g., under the pH values ranging from 2 to 10) and quite rigid because they have a Young's modulus of ∼109 Pa.17,34FliC has been used as radioprotective drugs in vivo.35 In addition, as long as 302 amino acids can be inserted into the hyper variable region of FliC, which allows us the flexibility in the selection of functional motifs or their combination to be presented on the flagella.17 These characteristics of flagella make them suitable for use as a building block in generating biomaterials.
Conclusions
In conclusion, we employed bioengineered bacterial flagella to fabricate an ECM-like matrix by a layer-by-layer assembly approach. This matrix mimics some key features of bone architecture. The HAP nanocrystals nucleated on flagella resemble minerals in bone and the RGD-displaying flagella mimic collagen fibrils. The novel genetically engineered flagella can support the adhesion and proliferation of BMSCs, indicating surface peptide or protein motifs of the bioengineered flagella can be recognized by BMSCs. The uniformly displayed functional motifs on the surface of flagella with multivalent ligands and nanoscale distance could highly up-regulate osteogenic differentiation and early maturation of the cells to the osteoblast-like cells. After biomineralization, the expression of osteogenic markers was further increased and kept at a high level during the entire cell culture time. The biomimetic mineralized matrix derived from genetically engineered flagella incorporated with bioactive motifs and bone minerals could be a promising osteogenic biomaterial. In essence, the flagella represent a new type of nanoscale osteogenic differentiation promoters.Conflicts of interest
There are no conflicts to declare.Acknowledgements
We would like to thank the partial financial support from the Office of Basic Energy Sciences within the Department of Energy (DOE) Office of Science (DE-SC0016567). We would also like to thank Salete Newton and Philip Klebba for assisting the peptide display on flagella.Notes and references
- A. M. Kushner and Z. Guan, Angew. Chem., Int. Ed., 2011, 50, 9026–9057 CrossRef CAS PubMed.
- L. Zhang and T. J. Webster, Nano Today, 2009, 4, 66–80 CrossRef CAS.
- S. Gomes, I. B. Leonor, J. F. Mano, R. L. Reis and D. L. Kaplan, Prog. Polym. Sci., 2012, 37, 1–17 CrossRef CAS PubMed.
- R. E. Abouzeid, R. Khiari, D. Beneventi and A. Dufresne, Biomacromolecules, 2018, 19, 4442–4452 CrossRef CAS PubMed.
- P. Gupta, M. Adhikary, M. J. Christakiran, M. Kumar, N. Bhardwaj and B. B. Mandal, ACS Appl. Mater. Interfaces, 2016, 8, 30797–30810 CrossRef CAS PubMed.
- G. Z. Yu, D. T. Chou, D. Hong, A. Roy and P. N. Kumta, ACS Biomater. Sci. Eng., 2017, 3, 648–657 CrossRef CAS PubMed.
- A. George and S. Ravindran, Nano Today, 2010, 5, 254–266 CrossRef CAS PubMed.
- M. Heim, D. Keerl and T. Scheibel, Angew. Chem., Int. Ed., 2009, 48, 3584–3596 CrossRef CAS PubMed.
- J. D. Hartgerink, E. Beniash and S. I. Stupp, Science, 2001, 294, 1684–1688 CrossRef CAS PubMed.
- Y. J. Shuai, C. B. Mao and M. Y. Yang, ACS Appl. Mater. Interfaces, 2018, 10, 31988–31997 CrossRef CAS PubMed.
- K. S. Sunderland, M. Y. Yang and C. B. Mao, Angew. Chem., Int. Ed., 2017, 56, 1964–1992 CrossRef CAS PubMed.
- B. R. Cao, M. Y. Yang and C. B. Mao, Acc. Chem. Res., 2016, 49, 1111–1120 CrossRef CAS PubMed.
- T. Elsdale and J. Bard, J. Cell Biol., 1972, 54, 626–637 CrossRef CAS PubMed.
- K. J. C. van Bommel, A. Friggeri and S. Shinkai, Angew. Chem., Int. Ed., 2003, 42, 980–999 CrossRef CAS PubMed.
- K. Yonekura, S. Maki-Yonekura and K. Namba, Nature, 2003, 424, 643–650 CrossRef CAS PubMed.
- R. M. Macnab, Annu. Rev. Microbiol., 2003, 57, 77–100 CrossRef CAS PubMed.
- B. Westerlund-Wikstrom, Int. J. Med. Microbiol., 2000, 290, 223–230 CrossRef CAS PubMed.
- J. D. Termine, Non-Collagen Proteins in Bone, John Wiley & Sons, Ltd., 2007 Search PubMed.
- D. Li, S. M. Newton, P. E. Klebba and C. Mao, Langmuir, 2012, 28, 16338–16346 CrossRef CAS PubMed.
- M. I. Setyawati, J. Xie and D. T. Leong, ACS Appl. Mater. Interfaces, 2014, 6, 910–917 CrossRef CAS PubMed.
- A.-W. Xu, Y. Ma and H. Calfen, J. Mater. Chem., 2007, 17, 415–449 RSC.
- C. B. Mao, H. D. Li, F. Z. Cui, Q. L. Feng and C. L. Ma, J. Mater. Chem., 1999, 9, 2573–2582 RSC.
- C. B. Mao, H. D. Li, F. H. Cui, C. L. Ma and Q. L. Feng, J. Cryst. Growth, 1999, 206, 308–321 CrossRef CAS.
- R. G. LeBaron and K. A. Athanasiou, Tissue Eng., 2000, 6, 85–103 CrossRef CAS PubMed.
- L. A. Smith, X. H. Liu, J. A. Hu and P. X. Ma, Biomaterials, 2010, 31, 5526–5535 CrossRef CAS PubMed.
- A. Bernhardt, A. Lode, S. Boxberger, W. Pompe and M. Gelinsky, J. Mater. Sci.: Mater. Med., 2008, 19, 269–275 CrossRef CAS PubMed.
- A. Merzlyak, S. Indrakanti and S. W. Lee, Nano Lett., 2009, 9, 846–852 CrossRef CAS PubMed.
- D. A. Wahl and J. T. Czernuszka, Eur. Cells Mater., 2006, 11, 43–56 CrossRef CAS PubMed.
- N. M. Alves, I. B. Leonor, H. S. Azevedo, R. L. Reis and J. F. Mano, J. Mater. Chem., 2010, 20, 2911–2921 RSC.
- A. W. Ettinger, M. Wilsch-Brauninger, A.-M. Marzesco, M. Bickle, A. Lohmann, Z. Maliga, J. Karbanova, D. Corbeil, A. A. Hyman and W. B. Huttner, Nat. Commun., 2011, 2, 503 CrossRef PubMed.
- D. Galli, L. Benedetti, M. Bongio, V. Maliardi, G. Silvani, G. Ceccarelli, F. Ronzoni, S. Conte, F. Benazzo, A. Graziano, G. Papaccio, M. Sampaolesi and M. G. Cusella De Angelis, J. Biomed. Mater. Res., Part A, 2011, 97, 118–126 CrossRef PubMed.
- K. Y. Lee, E. Alsberg, S. Hsiong, W. Comisar, J. Linderman, R. Ziff and D. Mooney, Nano Lett., 2004, 4, 1501–1506 CrossRef CAS PubMed.
- M. J. Dalby, N. Gadegaard, R. Tare, A. Andar, M. O. Riehle, P. Herzyk, C. D. W. Wilkinson and R. O. C. Oreffo, Nat. Mater., 2007, 6, 997–1003 CrossRef CAS PubMed.
- H. Hoshikawa and R. Kamiya, Biophys. Chem., 1985, 22, 159–166 CrossRef CAS PubMed.
- L. G. Burdelya, V. I. Krivokrysenko, T. C. Tallant, E. Strom, A. S. Gleiberman, D. Gupta, O. V. Kurnasov, F. L. Fort, A. L. Osterman, J. A. DiDonato, E. Feinstein and A. V. Gudkov, Science, 2008, 320, 226–230 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c9nh00124g |
| This journal is © The Royal Society of Chemistry 2019 |

![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif)