Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Ion effects on the conformation and dynamics of repetitive domains of a spider silk protein: implications for solubility and β-sheet formation†
Nur Alia
Oktaviani
 a,
Akimasa
Matsugami
b,
Fumiaki
Hayashi
a,
Akimasa
Matsugami
b,
Fumiaki
Hayashi
 b and
Keiji
Numata
b and
Keiji
Numata
 *a
*a
aBiomacromolecules Research Team, RIKEN Center for Sustainable Resource Scieences, 2-1 Hirosawa, Wako, Saitama 351-0198, Japan. E-mail: keiji.numata@riken.jp
bAdvanced NMR Application and Platform Team, NMR Research and Collaboration Group, NMR Science and Development Division, RIKEN SPring-8 Center, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama, Kanagawa 230-0045, Japan
First published on 29th July 2019
Abstract
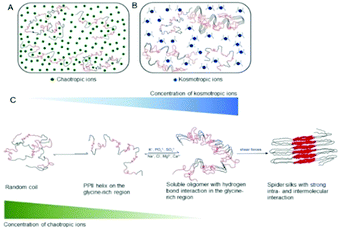
The effect of ions on the structure and dynamics of a spider silk protein is elucidated. Chaotropic ions prevent intra- and inter-molecular interactions on the repetitive domain, which are required to maintain the solubility, while kosmotropic ions promote hydrogen bond interactions in the glycine-rich region, which are a prerequisite for β-sheet formation.
Spider dragline silk is considered one of nature's super materials due to its superior mechanical properties. A dragline silk protein (spidroin) consists of an N-terminal domain (NTD) and a C-terminal domain (CTD) flanked by a long repetitive domain (approximately 100 repeat units).1–4 Based on solid state NMR studies, the repetitive domains of dragline silk fibers are mainly polyalanine domains which contain a β-sheet structure separated by glycine-rich regions (GGX), which forms a disordered 31 helix (31 helix is often recognized as polyproline type II, PPII, helix).5–7 Furthermore, other studies showed that LGXQ (X = S, G, N) motifs of dragline silk fibers from Nephila clavipes spider form a compact-turn like structure.8 In comparison, recently, it was reported that the glycine-rich region of the 47-mer peptide of dragline silk fibers, except LGSQG motifs, contains predominantly random coil populations.9 Interestingly, solution-state NMR data showed that the glycine-rich region of the soluble recombinant repetitive domain consists predominantly of a random coil (∼65%) and in addition, a PPII helix (∼25%) at temperature 10 °C.10 This PPII helix is considered a soluble prefibrillar form that might form intramolecular interactions by forming a reverse turn.10,15
Soluble spidroin is stored at high concentrations (20–50% w/v) in the spinning dope and is converted to an insoluble fiber after the CTD and NTD of spidroin exhibit pH- and ion-dependent conformations, which are crucial for controlling the pH-dependent assembly of spidroin.11–13 Another study also indicated that upon biomimetic spinning with pH gradient, recombinant minispidroins form fibers consisting of polyalanine domains with a β-sheet structure.14 However, the conformation of the recombinant repetitive spidroin domain is pH independent.10
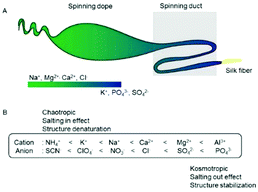
The spinning dope in the major ampullate gland contains high concentrations of Na+, Cl−, Mg2+ and Ca2+. The pH and chaotropic ion concentrations decrease and the PO43− and SO42− ion concentrations increase when approaching the spinning duct (Fig. 1A).16,17 In general, the Hofmeister series (Fig. 1B) is a classification of ions based on their ability to salt out and salt in proteins. Although, Mg2+ and Ca2+ ions are often categorized as kosmotropic ions, we classify Mg2+ and Ca2+ as chaotropic ions since they show the capability to salt-in repetitive domains of a spider silk protein.
In nature, the real concentration of chaotropic ions in spider glands has not been fully clarified. However, changes in the ionic composition in spider glands indicate that ions might be crucial for fiber formation.16 Currently, the effects of ions on the conformation and dynamics of the repetitive domain of spidroin for fiber formation still remain elusive. Herein, we present the effect of chaotropic and kosmotropic ions on the conformation and dynamics of the repetitive domain using NMR spectroscopy.
We prepared a recombinant silk protein consisting of 15 repeating domain units, namely, a 15-mer. A repetitive domain monomer, consisting of 1 repeat unit, was also prepared as a control to evaluate the effects of inter- and intramolecular interactions. Both of these repetitive domain constructs were designed based on the amino acid sequences in major spidroin I of N. clavipes (Fig. S1A, ESI†). All the backbone resonances of this domain were assigned at a pH of 7 and a temperature of 15 °C using 2D and 3D triple-resonance NMR spectra (Fig. S1B, ESI†).
In the presence of Na+ and Cl− ions, the signal intensities of the glycine-rich region of the repetitive domain increased drastically (Fig. 2), particularly in the SQGTSG region (Fig. 2B, blue shadow). In contrast, those ions did not affect the signal intensity ratios of the polyalanine region (Fig. 2B, red shadow). Similar phenomena were observed in the signal intensities of the 15-mer in the presence of other chaotropic ions (K+, Mg2+, and Ca2+) (Fig. S2, ESI†). The weak signal intensities of atoms in the glycine-rich region in the absence of chaotropic ions suggest the presence of weak interactions in this region, which normally cause intermediate exchange processes, resulting in line broadening in the NMR spectra.18 These interactions may have originated from weak hydrogen bonds in the glycine-rich region, which contains a PPII helix population according to previous studies.10,19 Upon addition of chaotropic ions, the signal intensities of the glycine-rich region increase, which indicates that these ions prevent interactions in this region, particularly the SQGTSG region (Fig. 2B). This motif is located in the turn region of spider silks.8 On the other hand, the signal intensities of polyalanine were not affected by the presence of chaotropic ions at different concentrations (intensity ratio ≈ 1) (Fig. 2B). This result suggests that the polyalanine region, which contains a predominantly random coil population based on previous studies,10,20 is not affected by the concentration of chaotropic ions.
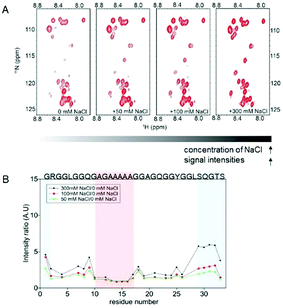 | ||
| Fig. 2 Effect of the chaotropic ions (NaCl) on the conformation of the repetitive domain (15-mer) of spidroin. (A) 2D 1H–15N HSQC spectra of the 15-mer in solutions at different NaCl concentrations (from 0 to 300 mM NaCl). As the concentration of NaCl increases, the signal intensities of the 15-mer also increase. (B) Plot of the intensity ratio of the 15-mer as a function of the residue number. The red shadow indicates the polyalanine region, which is not affected by the ion concentration. The blue shadow indicates the intensity ratios on the SQGTSG region, which is highly affected by NaCl. Similar results were observed for the effects of K+, Mg2+ and Ca2+ on the 15-mer (Fig. S2, ESI†). | ||
In comparison, an increase in the signal intensity ratios in the monomer spectra was observed for the glycine-rich region (Fig. S3, ESI†), but such a drastic increase in the intensity ratios was not observed for the SQGTSG region at the C-terminal end of the monomer. These results indicate that chaotropic ions also prevent intermolecular interactions, but the significant increase in the intensity ratio in the SQGTSG region of the 15-mer was mainly due to the fact that chaotropic ions prevent intramolecular interactions involving this SQGTSG region.
The addition of chaotropic ions (Na+ and Cl−) at different concentrations did not change the 1H–15N HSQC spectra (Fig. 2A) or the conformation of the repetitive domain (Fig. S4, ESI†). The 3JHNHA coupling constants of the 15-mer in the absence and presence of 300 mM NaCl at 15 °C showed similar patterns (Fig. S4E, ESI†). These NMR results were also in agreement with the vibrational circular dichroism (VCD) data of the 15-mer, which showed that in the presence of 300 mM NaCl (Fig S5, ESI†), the PPII helix population does not change significantly. The {1H}–15N heteronuclear NOE data of the 15-mer also demonstrated that the chaotropic ions do not affect the local dynamics of the 15-mer (Fig. S6A, ESI†). Furthermore, the 15NT2 relaxation times of the 15-mer increased as the ionic strength of the chaotropic ions (Mg2+) increased (Fig. S6B, ESI†), even though the trend in the 15NT2 relaxation times did not change as a function of the residue number. These results imply that the higher ionic strengths of the chaotropic ions prevent inter and intramolecular interactions of the repetitive domains without changing the conformation of the soluble repetitive domains.
The relaxation data of the 15-mer in the absence and presence of 300 mM NaCl at 15 °C were recorded at 700 and 800 MHz (Fig. S7, ESI†). Spectral densities, which reflect the probability of the molecular motion of this protein being at the angular frequency of the magnetic field (Fig. S8, ESI†), were calculated. The spectral densities (J(0.87ωH) and J(ωN)) of the 15-mer in both the absence and presence of 300 mM NaCl were quite similar. However, the J(0) of the 15-mer in the absence of salt was higher than the J(0) of the 15-mer in the presence of 300 mM NaCl. The smaller J(0) value of the 15-mer in the presence of 300 mM NaCl suggests that the chaotropic ions have two functions, namely, to suppress conformational exchanges of the 15-mer and to prevent inter and intramolecular interactions. These findings are also in agreement with a previous hydrodynamic study on spidroin, which revealed that NaCl maintains the compact form of spidroin and prevents oligomerization.21
Contrary to the chaotropic ions, higher concentrations of a kosmotropic ion (PO43−) caused a change in the 1H–15N HSQC spectrum of the 15-mer (Fig. 3A). The dispersion of the glycine signals of the 15-mer in the presence of potassium phosphate at concentrations above 300 mM indicates that the 15-mer becomes more rigid due to the presence of hydrogen bonds, especially around residue G3 (Fig. 3B) or the SQGTSGRG motif (Fig. S10C, ESI†). In natural dragline silk, G3 is located close to the turn region.8 In addition to the proton signal of G3, the chemical shifts of the amide protons of serine, threonine and other glycine signals were also shifted downfield (Fig. 3A), indicating an increase in the hydrogen bonding interactions in the glycine-rich region.22 These findings also agree with {1H}–15N heteronuclear NOE data of the 15-mer at higher phosphate concentrations (300 mM and 600 mM), which demonstrated a decrease in the NOEs as the potassium phosphate concentration increased. These dynamics data indicated that the local flexibility of the 15-mer decreases as the potassium phosphate concentration increases (Fig. 3C).
A similar effect was also observed on the 15-mer in the presence of sulfate (SO42−) (Fig. S9, ESI†). At a higher concentration of sulfate, the proton chemical shifts of glycine were shifted downfield, suggesting that the hydrogen bonding interaction increased in this region (Fig. S9A, ESI†) and the G3 signal did not overlap with other glycine signals (Fig. S9B, ESI†). Accordingly, the NOE values of the 15-mer slightly increased, particularly in the glycine-rich region, indicating the reduced local flexibility (Fig. S9C, ESI†) in the presence of 300 mM potassium sulfate. However, the differences in the amide proton chemical shifts in the presence and absence of SO42− are smaller than the changes in the presence and absence of PO43− because SO42− has a lower ionic strength than PO43−; thus, SO42− has less ability to promote hydrogen bonding interactions in this region (Fig. S10, ESI†). A higher concentration of potassium phosphate (≥600 mM) leads to precipitation of the 15-mer (Fig. S11, ESI†). The increase in the NOE values of the monomer in the presence of 300 mM potassium phosphate suggests that the local flexibility of the monomer decreases due to intermolecular interactions (Fig. S12, ESI†). These findings explain the phenomena reported in a previous study, in which potassium phosphate can promote nucleation as well as fibril growth in a recombinant spider silk protein.23
The dynamics data of the 15-mer in the presence of 600 mM potassium phosphate were recorded at 700 and 800 MHz (Fig S13, ESI†). The spectral density J(0.87ωH) of the 15-mer in the presence of 600 mM potassium phosphate (Fig. S14A, ESI†) and 10 mM potassium phosphate remained quite similar. However, the spectral densities J(ωN) of the 15-mer in the presence of 600 mM potassium phosphate were slightly larger than that in 10 mM potassium phosphate (Fig. S14B, ESI†), suggesting a decrease in the dynamics of the 15-mer in the presence of 600 mM potassium phosphate on the sub-nanosecond time scale.24 Interestingly, the spectral density J(0) of the 15-mer in the presence of 600 mM potassium phosphate (Fig. S14C, ESI†) was still much smaller than that in the presence of 10 mM potassium phosphate (Fig. S14C, ESI†) because the high concentrations of potassium phosphate could suppress the conformational exchange of the 15-mer. Although based on amide proton signals, hydrogen-bonding interaction increases on the SQGTSGRG motif in the presence of high concentration of the kosmotropic ions, which will enhance rigidity on the local dynamics of repetitive domains, the significant change in the secondary structure population on soluble repetitive domains cannot be observed clearly under this condition.
Low charge density ions (chaotropes) could bind directly to the amide group of the repetitive domain, particularly in the glycine-rich region.25 These ions might preferentially interact with the SQGTSG region, which is more polar, and thus prevent weak interactions in this region, which is required for maintaining the solubility of spidroin (Fig. 4A). Similar phenomena were also observed with an elastomeric protein in the presence of 300 mM NaCl.26 On the other hand, high charge densities of the kosmotropic ions polarize water molecules that are hydrogen bonded with amide groups, decreasing the hydration of the amide protons, which will promote hydrogen bonding in this region (Fig. 4B). The smaller J(0) value of the SQGTSGR region of the 15-mer in the presence of 600 mM potassium phosphate relative to that in the presence of 300 mM NaCl (Fig. S15, ESI†) implies that this region is more likely to have a turn. This finding agrees with a previous study which demonstrated that this motif in the repetitive domain forms turns in N. clavipes dragline silk fibers.8 Furthermore, the increase in hydrogen bonding interactions in the SQGTSGR region together with the shear forces cause polyalanine interactions and finally promote β-sheet formation of spidroin.
Previously, the effect of ions on recombinant repetitive domains of spidroin using other analytical methods such as DLS, Raman spectroscopy, and FT-IR spectroscopy had been reported21,23,27 and those results agree with this study. However, the novelty of this study over previous reports is that by using solution-state NMR spectroscopy, the effects of ions on the conformation, dynamics and self-assembly of repetitive domains of spidroin were elucidated in molecular details. In conclusion, we showed that the effect of ions on repetitive domains is crucial for regulating the solubility and β-sheet formation of the dragline silk protein. The chaotropic ions prevent intra and intermolecular interactions on the repetitive domains, which are necessary to maintain the solubility of spidroin, while the kosmotropic ions promote hydrogen bond interactions in the glycine-rich region, especially in the SQGTSGRG region. The formation of hydrogen bonds in this region is a preliminary step that leads to polyalanine interactions to form β-sheets. Our in vitro study demonstrated the functional role of ions in the self-assembly of spidroin, which will provide a better understanding of this process to facilitate the preparation of artificial spider silks with outstanding mechanical properties.
Conflicts of interest
The authors have no conflicts to declare.Notes and references
- A. Sponner, W. Vater, W. Rommerskirch, F. Vollrath, E. Unger, F. Grosse and K. Weisshart, Biochem. Biophys. Res. Commun., 2005, 338, 897–902 CrossRef CAS PubMed.
- D. Motriuk-Smith, A. Smith, C. Y. Hayashi and R. V. Lewis, Biomacromolecules, 2005, 6, 3152–3159 CrossRef CAS PubMed.
- G. Candelas, T. Candelas, A. Ortiz and O. Rodríguez, Biochem. Biophys. Res. Commun., 1983, 116, 1033–1038 CrossRef CAS PubMed.
- M. Xu and R. V. Lewis, Proc. Natl. Acad. Sci. U. S. A., 1990, 87, 7120–7124 CrossRef CAS PubMed.
- J. M. Gosline, P. A. Guerette, C. S. Ortlepp and K. N. Savage, J. Exp. Biol., 1999, 202, 3295–3303 CAS.
- J. Kümmerlen, J. D. van Beek, F. Vollrath and B. H. Meier, Macromolecules, 1996, 29, 2920–2928 CrossRef.
- G. P. Holland, M. S. Creager, J. E. Jenkins, R. V. Lewis and J. L. Yarger, J. Am. Chem. Soc., 2008, 130, 9871–9877 CrossRef CAS PubMed.
- C. A. Michal and L. W. Jelinski, J. Biomol. NMR, 1998, 12, 231–241 CrossRef CAS PubMed.
- T. Asakura, A. Nishimura and Y. Tasei, Macromolecules, 2018, 51, 3608–3619 CrossRef CAS.
- N. A. Oktaviani, A. Matsugami, A. D. Malay, F. Hayashi, D. L. Kaplan and K. Numata, Nat. Commun., 2018, 9, 2121 CrossRef PubMed.
- G. Askarieh, M. Hedhammar, K. Nordling, A. Saenz, C. Casals, A. Rising, J. Johansson and S. D. Knight, Nature, 2010, 465, 236–238 CrossRef CAS PubMed.
- F. Hagn, L. Eisoldt, J. G. Hardy, C. Vendrely, M. Coles, T. Scheibel and H. Kessler, Nature, 2010, 465, 239–242 CrossRef CAS PubMed.
- F. Hagn, C. Thamm, T. Scheibel and H. Kessler, Angew. Chem., Int. Ed., 2011, 50, 310–313 CrossRef CAS PubMed.
- M. Otikovs, M. Andersson, Q. Jia, K. Nordling, Q. Meng, L. B. Andreas, G. Pintacuda, J. Johansson, A. Rising and K. Jaudzems, Angew. Chem., Int. Ed., 2017, 56, 12571–12575 CrossRef CAS PubMed.
- A. A. Adzhubei and M. J. E. Sternberg, J. Mol. Biol., 1993, 229, 472–493 CrossRef CAS PubMed.
- D. P. Knight and F. Vollrath, Naturwissenschaften, 2001, 88, 179–182 CrossRef CAS PubMed.
- M. Andersson, G. Chen, M. Otikovs, M. Landreh, K. Nordling, N. Kronqvist, P. Westermark, H. Jörnvall, S. Knight, Y. Ridderstråle, L. Holm, Q. Meng, K. Jaudzems, M. Chesler, J. Johansson and A. Rising, PLoS Biol., 2014, 12, e1001921 CrossRef PubMed.
- A. K. Mittermaier and L. E. Kay, Trends Biochem. Sci., 2009, 34, 601–611 CrossRef CAS PubMed.
- T. Lefèvre, J. Leclerc, J.-F. Rioux-Dubé, T. Buffeteau, M.-C. Paquin, M.-E. Rousseau, I. Cloutier, M. Auger, S. M. Gagné, S. Boudreault, C. Cloutier and M. Pézolet, Biomacromolecules, 2007, 8, 2342–2344 CrossRef PubMed.
- D. Xu, C. Guo and G. P. Holland, Biomacromolecules, 2015, 16, 2072–2079 CrossRef CAS PubMed.
- J. Leclerc, T. Lefèvre, M. Gauthier, S. M. Gagné and M. Auger, Biopolymers, 2013, 99, 582–593 CrossRef CAS PubMed.
- B. Berglund and R. W. Vaughan, J. Chem. Phys., 1980, 73, 2037–2043 CrossRef CAS.
- M. Humenik, A. M. Smith, S. Arndt and T. Scheibel, J. Struct. Biol., 2015, 191, 130–138 CrossRef CAS PubMed.
- N. A. Farrow, O. Zhang, A. Szabo, D. A. Torchia and L. E. Kay, J. Biomol. NMR, 1995, 6, 153–162 CrossRef CAS PubMed.
- M. C. Gurau, S.-M. Lim, E. T. Castellana, F. Albertorio, S. Kataoka and P. S. Cremer, J. Am. Chem. Soc., 2004, 126, 10522–10523 CrossRef CAS PubMed.
- S. E. Reichheld, L. D. Muiznieks, F. W. Keeley and S. Sharpe, Proc. Natl. Acad. Sci. U. S. A., 2017, 201701877 Search PubMed.
- K. Jastrzebska, E. Felcyn, M. Kozak, M. Szybowicz, T. Buchwald, Z. Pietralik, T. Jesionowski, A. Mackiewicz and H. Dams-Kozlowska, Sci. Rep., 2016, 6, 28106 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c9cc03538a |
| This journal is © The Royal Society of Chemistry 2019 |