Open Access Article
Open Access ArticleSilver nanoparticle modified surfaces induce differentiation of mouse kidney-derived stem cells†
Neelika
Roy Chowdhury
a,
Isabel
Hopp
b,
Peter
Zilm
c,
Patricia
Murray
*b and
Krasimir
Vasilev
 *a
*a
aSchool of Engineering, University of South Australia, Mawson Lakes, SA 5095, Australia. E-mail: Krasimir.vasilev@unisa.edu.au
bInstitute of Translational Medicine, University of Liverpool, Liverpool, UK. E-mail: P.A.Murray@liverpool.ac.uk
cMicrobiology Laboratory, Adelaide Dental School, The University of Adelaide, Adelaide, SA 5005, Australia
First published on 4th June 2018
Abstract
In this paper, we interrogate the influence of silver nanoparticle (AgNPs)-based model surfaces on mouse kidney-derived stem cells (mKSCs) differentiation. The widespread use of silver in biomedical and consumer products requires understanding of this element's effect on kidney cells. Moreover, the potential for using stem cells in drug discovery require methods to direct their differentiation to specialized cells. Hence, we generated coated model substrates containing different concentrations of surface immobilized AgNPs, and used them to evaluate properties and functions of mKSCs. Initially, mKSCs exhibited reduced viability on higher silver containing surfaces. However, longer culture periods assisted mKSCs to recover. Greater degree of cell spreading and arborization led by AgNPs, suggest podocyte differentiation. Proximal tubule cell marker's expression revealed differentiation to the specific lineage. Although the exact mechanism underpinning these findings require significant future efforts, this study demonstrate silver's capacity to stimulate mKSC differentiation, which may provide opportunities for drug screenings.
Introduction
Silver and silver nanoparticles (AgNPs) have been utilized in a number of consumer products because of their capacity to kill bacteria.1–7 In medicine, silver coated catheters, wound dressings and a range of other devices are currently used on a daily basis.3,8 With the potential benefits brought by the antibacterial properties of silver come concerns regarding toxicity to mammalian cells, tissues and overall human health. Although mammalian cells can generally tolerate greater amounts of silver than bacteria, the element can also be toxic to tissues if used in high concentrations.9–11 The inflammatory response to silver is still an open question with the area being rather controversial. Many studies claim reduced inflammation or no alteration of inflammatory response while others report adverse responses. Another area that needs further attention is related to the effect of silver on stem cell fate. We have recently looked at the effect of silver on the differentiation of mesenchymal stem/stromal cell (MSC) and found that oxidative stress caused by the silver led to preferential differentiation to adipocytes rather than osteoblasts.12 This report demonstrated the need for greater attention to how silver may affect stem cells.Chronic kidney disease (CKD) is recognized as a serious global health problem.13 End stage renal disease (ESRD) is the final stage of CKD and is associated with high rates of morbidity and mortality, as well being financially costly, with the United States alone spending >US$ 50,000 for each patient suffering from ESRD. Although current therapeutic interventions slow down the progression of CKD to ESRD, the current state of treatment does not generate satisfactory outcomes in all patients.14 Therefore, it is extremely important to develop new treatment strategies where stem cells can be directed to differentiate to specialized renal cell types that could be useful to treat CKD in order to prevent ESRD progression.
The kidney is a highly complex organ with several distinct cell types,15 but two cell types in particular, podocytes and proximal tubule cells (PTCs), play crucial roles in maintaining normal kidney function. For instance, podocytes are required to maintain the glomerular filtration barrier, and PTCs are necessary for secreting metabolites into the tubular lumen, and for the absorption of various molecules from the glomerular filtrate. PTCs also play a role in drug metabolism, and consequently, drug-induced nephrotoxicity is a major cause of kidney disease.16–18 Podocytes and PTCs are difficult cells to culture because the former are post-mitotic and the latter rapidly lose their phenotype following isolation from the kidney. There has thus been great interest in generating these cell types from stem cells in order to understand their physiology, investigate how this becomes altered in kidney disease, and to use them in drug screening programs.
We have previously isolated stem cells from neonatal mouse kidneys that are able to generate podocytes and proximal tubule cells (PTCs) in vitro, and ex vivo when transplanted into mouse embryo kidneys.19,20 Recent reports have also demonstrated that the chemistry and nanotopography of cell culture surfaces regulate the proliferation and differentiation of various stem cell lines.14,21,22 MacGregor-Ramiasa et al. have investigated the effect of concentration gradient of –NH2 functional groups on the differentiation of a clonal kidney stem cell (mKSCs) line derived from neonatal mouse kidney. Differentiation of proximal-tubule-like cells was observed on the surfaces with low amine content by the expression of specific markers.23
With the increased promise of cell therapies and the wide application of silver containing products in medicine and everyday life, it becomes important to evaluate the effect of silver based antibacterial coatings on mKSC fate. In this work, we used established platforms for generation of antibacterial surfaces combining plasma polymer coatings and silver nanoparticles.6 These types of surfaces were selected since they were demonstrated in our previous work to have excellent antibacterial properties but no toxicity to primary human fibroblasts or MSCs. The method of preparation also allowed us to generate two different silver nanoparticle surface concentrations which facilitated examination of dose dependent effects.6,12 The effect of the silver nanoparticle modified surface on the attachment, proliferation, viability, spreading and differentiation of mKSCs was evaluated via microscopic and immunostaining techniques.
Experimental
Materials
2-Methyl-2-oxazoline, NaOH pellets, phosphate buffer saline (PBS) tablets, foetal bovine serum (FBS), 2-mercaptosuccinic acid (MSA) (97%), sodium borohydride (NaBH4), nitric acid (70%) were purchased from Sigma-Aldrich Australia and used as received. Hydrochloric acid (36%, Ajax Finechem Pty. Ltd. Australia) was used as received. Dulbecco's Modified Eagle Medium (DMEM) was purchased from Gibco. Penicillin, streptomycin, L-glutamine, MEM non-essential amino acids, cell counting kit-8 (CCK-8), Naphthol AS-MX phosphate and Fast Red TR were obtained from Sigma. Silver nitrate (AgNO3) and glass coverslips were bought from ProSciTech, Australia. Ultra-pure MilliQ water (resistivity 18.2 Ω) was used for all experiments and cleaning procedures.Plasma deposition of 2-methyl-2-oxazoline
A custom built capacitively coupled bell-chamber reactor with a 13.56 MHz plasma generator and a matching network (Advanced Energy, USA) described elsewhere24 was used for plasma deposition. 13 mm glass coverslips and silicon wafers were cleaned with ethanol, dried with nitrogen stream and stored in airtight containers until further use. The substrates were cleaned using air plasma (50 W, 5 minutes, 2 × 10−1 mbar) prior to any plasma treatment. Deposition of 2-methyl-2-oxazoline was carried out at 50 W for 2 minutes at a precursor pressure of 2 × 10−1 mbar. To stabilize the plasma coating, the substrates were kept overnight at room temperature in sealed containers prior to any further modification.Synthesis of AgNPs@MSA
Silver nanoparticles were synthesized by sodium borohydride reduction of silver nitrate.6 Briefly, 12 ml of 2 mM AgNO3 was mixed with 5 ml of 2 mM MSA under ice cold condition. 0.5 ml of 0.5 M of NaBH4 was added to the solution under vigorous stirring. The color of the solution mixture turned to reddish brown after the addition of NaBH4 indicated the formation of AgNPs. The AgNPs were stored in light proof container until further use. The solutions remain stable for several months without showing any visible aggregation of particles.Immobilization of AgNPs@MSA on pPOX substrates
The plasma modified substrates were incubated at room temperature in AgNPs solution for time intervals of 6 and 24 h. Depending on the AgNPs@MSA immobilization time the samples will be represented as SL and SH (6 and 24 h AgNPs@MSA immobilization time respectively) in this paper. After incubation the AgNPs@MSA solution was discarded and the samples were rinsed 3 times with MilliQ water to remove any loosely bound AgNPs. Then the samples were dried with nitrogen stream and stored in airtight container until further use.Glass coverslips were used for atomic force microscope (AFM) and cell culture studies, while silicon wafers were used for ellipsometry and X-ray photoelectron spectroscopy (XPS).
Surface characterization
Cell culture
All solutions and buffers used during cell culture were pre-warmed to 37 °C prior to use. The stem cell line was derived from neonatal mouse kidney by Fuente Mora et al.19 Cells were cultured in DMEM in standard tissue culture plates (TCPs). The medium was changed every 2–3 days and cells were incubated with 5% CO2 at 37 °C. 13 mm coated and uncoated glass coverslips were placed in 24 well TCPs and mKSCs were seeded at a density of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well and incubated for a period of 48 and 96 h. Care was taken to ensure that the cell suspension was evenly distributed during plating.
000 cells per well and incubated for a period of 48 and 96 h. Care was taken to ensure that the cell suspension was evenly distributed during plating.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10) was added to each well and cells were incubated for 1 h. Then 100 μL of the reaction solution was transferred into a 96 well plate. The absorbance of the solution was measured by an Anthos Labtec LP400 spectrophotometer at wavelength of 450 nm for detection (620 nm reference filter). The background of the culture medium was measured and subtracted from all values.
10) was added to each well and cells were incubated for 1 h. Then 100 μL of the reaction solution was transferred into a 96 well plate. The absorbance of the solution was measured by an Anthos Labtec LP400 spectrophotometer at wavelength of 450 nm for detection (620 nm reference filter). The background of the culture medium was measured and subtracted from all values.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200) labelled with Alexa Fluor® 488 (Invitrogen) for 30 minutes at room temperature in the dark and then nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). Samples were imaged under epifluorescence illumination using a Leica DM2500 microscope fitted with a Leica DFC420C camera. An average of 10 images per coverslip was taken and cell number quantified by counting stained nuclei from the images obtained. All images are representative for the entire surface from the corresponding sample. The extent of cell spreading (phalloidin-stained cells) was measured using ImageJ.
200) labelled with Alexa Fluor® 488 (Invitrogen) for 30 minutes at room temperature in the dark and then nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI). Samples were imaged under epifluorescence illumination using a Leica DM2500 microscope fitted with a Leica DFC420C camera. An average of 10 images per coverslip was taken and cell number quantified by counting stained nuclei from the images obtained. All images are representative for the entire surface from the corresponding sample. The extent of cell spreading (phalloidin-stained cells) was measured using ImageJ.
From the total cell numbers at different time points, the mKSC population doubling times (PDTs) were calculated with the help of non-linear regression analysis using the following equation:
| PDT = ln(2)/K |
| Name | Type | Concentration | Manufacturer | |
|---|---|---|---|---|
| Primary antibody | LRP2 megalin | Mouse monoclonal IgG1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
Acris, DM3613P |
| Podocalyxin | Mouse monoclonal IgG2 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
R & D systems, MAB1556 | |
| Secondary antibody | Alexa Fluor 594 goat α mouse | IgG1-594 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 1000 |
Invitrogen, A-21125 |
For quantification of images, an average of 10 images were taken from each sample and number of positive cells was quantified. Images shown represent the entire sample surface as well as an average of all experiments.
All experiments were performed with three biological and technical replicates. Methyl-oxazoline plasma polymerized surfaces and uncoated glass coverslips were used as positive (PC) and negative controls (NC) respectively for all cell culture experiments.
Results and discussion
Substrate preparation and characterization
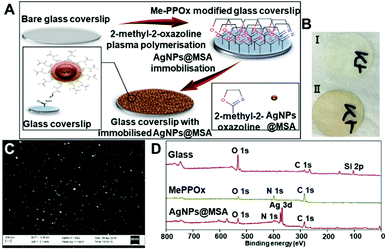
To generate model substrates which facilitate understanding of the effect of silver-based coating on the fate of mKSCs, we used silver nanoparticles (AgNPs@MSA) that we reported earlier.6We knew that coatings generated using these nanoparticles have excellent antibacterial properties and are non-cytotoxic to primary human fibroblasts and mesenchymal stem cells.6,12 Our early approach to immobilize these nanoparticles to the surface was via electrostatic coupling to allylamine plasma polymer films which are known to retain significant number of amine groups.27,28 In this work, we used covalent attachment of the nanoparticle to the substrate since it presents a more stable platform compared to electrostatic binding.29 To achieve this, we used relatively recent plasma polymer coatings developed by our group which are prepared from oxazoline (pPOX) based precursors. The resultant films have been demonstrated to retain a population of intact oxazoline functionalities which can be used to covalently bind nanoparticles and ligands containing carboxylic acid groups.30–32 The conditions selected for the plasma deposition resulted in a film thickness of 25.01 ± 0.03 nm as measured by ellipsometry. AgNPs@MSA were immobilized by immersion of the plasma polymer modified surface in a solution of silver nanoparticles for predetermined time (Fig. 1A).
After AgNPs@MSA immobilization, the samples showed a homogenous light brownish colour which is due to the plasmon resonance adsorption of the silver nanoparticles (Fig. 1B, II). The AgNPs@MSA were spherical with an average size of 12.27 nm similar to those in ref. 6. The SEM analysis showed uniform particle distribution on the plasma modified surfaces and absence of aggregations (Fig. 1C). AFM imaging before the immobilization of the AgNPs@MSA (Fig. SI 1A†) demonstrated that the coatings are continuous, pin hole free with a surface roughness of 0.569 nm (Fig. SI 1C†). The surface roughness increased to 5.511 nm after the deposition of AgNPs@MSA (Fig. SI 1D†). XPS survey spectra of the unmodified glass coverslips (NC), pPOX coated substrates (PC) and pPOX surfaces with the immobilized AgNPs@MSA is shown in Fig. 1D. As expected, only three elements are present on the bare glass surface – carbon, oxygen and silicone. Substrates with pPOX plasma polymer film showed the presence of nitrogen, in addition to carbon and oxygen, which is consistent with the chemical composition of the precursor. Along with the carbon, nitrogen and oxygen a distinct Ag3d peak was detected when AgNPs@MSA were immobilized on the plasma polymer modified samples. The atomic concentration of chemical elements on the surface of the samples is presented in Table 2. SH and SL denote different time of nanoparticle immobilization for 24 h and 6 h, respectively. As anticipated the atomic concentration of silver increased with the time of immobilization from 8.61 ± 0.3% (6 h, samples SL) to 13.95 ± 0.1% (24 h, samples SH). Longer immobilization time resulted in greater silver concentration corresponding to greater number of nanoparticles binding to the surface. This time dependent immobilization process allowed us to generate surfaces with different and controlled concentration of silver nanoparticles. Both silver surface concentrations are sufficient to completely eliminate bacterial growth as we demonstrated in ref. 6, however, this allows us to test the influence of different silver surface concentration on the fate of mKSCs.
| C1s | N1s | O1s | Si2p | Ag3d | |
|---|---|---|---|---|---|
| NC | 15.62 | 54.00 | 30.36 | ||
| PC | 72.45 | 18.06 | 9.48 | ||
| SL | 48.58 | 30.90 | 11.98 | 8.61 | |
| SH | 46.74 | 22.20 | 17.10 | 13.95 |
Effect of silver nanoparticle concentration on mKSC viability, proliferation and morphology
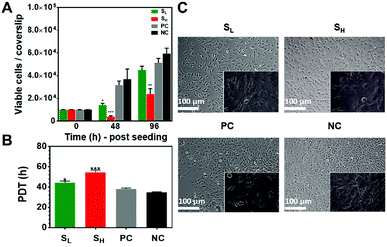
The viability of mKSCs cultured on silver nanoparticle modified surfaces and controls was determined using the CCK-8 kit assay (Fig. 2A). The results indicate strong reduction in cell viability after 48 h in culture, particularly for the higher silver nanoparticle concentration. However, the cells recovered after 96 h, especially at the lower nanoparticle concentration where cell viability was only 15–20% lower compared to than on control substrates without silver. The reduced initial viability and attachment of the cell on the silver containing samples could be due to the substantial amount of released silver ions from the surfaces.6,12 The silver ions are the active species that make silver such a potent antibacterial agent but the element could be toxic to mammalian cells too. In a pioneering study we demonstrated the mammalian cells can tolerate greater amount of silver than bacteria. Our finding was also confirmed by others and our follow up studies3,6,12 which showed that amounts of silver lethal to bacteria did not significantly affect primary human fibroblasts and mesenchymal stem cells (MSCs). However, mammalian cells may have different tolerance to silver and our current study demonstrate that, although capable of recovery, the viability of mKSCs was reduced in the early stages of growth. Overall, these data suggest both time- and concentration-dependent effects of silver nanoparticles on mKSC viability.We also assessed the mKSC population doubling time (Fig. 2B). It was greater on the silver nanoparticle modified surface i.e. 45 h and >52 h for SL and SH, respectively. In comparison, the PDT was <40 h on control surfaces.
Podocyte differentiation
mKSCs have been shown to differentiate on surfaces to podocytes and PTCs.19,20,23 To investigate if the propensity of cells to undergo podocyte differentiation varied between different substrates, we first measured the extent of cell spreading, because podocyte-like cells tend to have a large cytoplasm![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
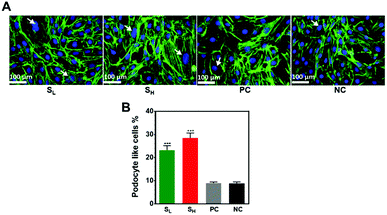
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) nuclear ratio. F-Actin staining was used to facilitate evaluation of cell spreading area (Fig. 3A). The average spreading areas of cells seeded on silver nanoparticle modified surfaces was nearly twice larger compared to glass coverslip control with (PC) or without (NC) a methyl oxazoline plasma polymer coating. The significant increase of cell spreading area on samples containing nanoparticles strongly suggests that the silver nanoparticles play a role in driving the observed phenomenon.
nuclear ratio. F-Actin staining was used to facilitate evaluation of cell spreading area (Fig. 3A). The average spreading areas of cells seeded on silver nanoparticle modified surfaces was nearly twice larger compared to glass coverslip control with (PC) or without (NC) a methyl oxazoline plasma polymer coating. The significant increase of cell spreading area on samples containing nanoparticles strongly suggests that the silver nanoparticles play a role in driving the observed phenomenon.
In some cells, F-actin staining showed a pattern typical of that in primary podocytes. These cells tended to be highly arborized and were often binucleate, all of which are typical characteristics of podocytes. The percentage of podocyte like cells on silver treated samples particularly on the SH samples was significantly higher than the controls (Fig. 3B). Upon differentiation, on both samples containing AgNPs@MSA, mKSCs increased in cell size, developed binucleate cells with arborized morphology (Fig. 3A) after 96 h. Immunostaining analysis showed the expression of podocyte specific marker podocalyxin by mKSCs cultured on the silver containing substrates. Representative images are provided in the ESI (Fig. SI 2†). The data also suggests that the greater silver concentration facilitate differentiation of mKSCs into podocyte-like cells.
Actin cytoskeleton have key roles in physiological function of podocytes and maintenance of glomerular filtration barrier.33–35 Several studies have reported that mutation or changes in the actin cytoskeleton lead to dysfunctional podocytes and subsequent proteinuria.33,36–38 No disruption in the actin cytoskeleton organization was observed on the silver treated substrates (Fig. 3A). Our results therefore confirm that the silver did not induce any derangement of the actin cytoskeleton.
Proximal tubule cell differentiation
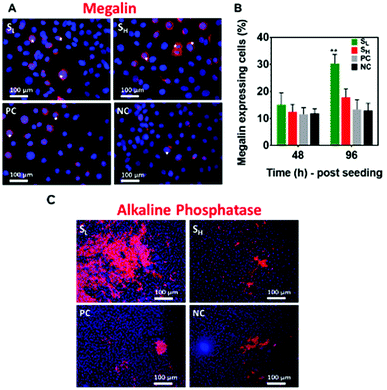
To examine the differentiation of mKSCs to PTCs we have carried out staining for megalin. The data showed a greater number of megalin expressing cells on silver nanoparticles modified surfaces and particularly on those with lower silver concentration (SL). The plasma polymer only coating did not show any notable difference to that of glass coverslip control (Fig. 4A), and these cells are significantly greater in number on the SL surfaces (Fig. 4B).To further confirm the differentiation to PTCs we carried out staining for alkaline phosphatase, an enzyme that is expressed by PTCs.39 The images presented in Fig. 4C show significantly stronger alkaline phosphatase activity of mKSCs grown on SL. This is in good agreement with the data obtained from the megalin assays and further strengthen the finding that silver may be stimulating mKSCs to differentiate into PTCs.
The silver nanoparticle modified surfaces appeared to trigger differentiation of mKSCs to podocyte and PTCs lineage. There are at least two possible explanations for the observed phenomena. Silver nanoparticles have been shown to cause oxidative stress in MSCs and in this way drive the differentiation of these cells.12 It may be possible that this is the reason behind the current findings pointing to increased differentiation of the mKSCs when silver was present. On another hand, the silver nanoparticles create surface roughness (Fig. S1†). We have demonstrated recently that tailored surface roughness, including roughness comparable with this inherent for the model samples used in this study, was capable of guiding the differentiation of mKSCs to podocytes.23 This finding was not surprising since it is now well accepted that surface nanotopography has significant influence on the fate of various types of cells, including immune and stem cells.27,40,41
Conclusions
Novel clinical approaches that involve the use of nanoengineered biomaterials to support and/or direct the differentiation and/or renewal of podocytes and PTCs, are urgently required to combat ESRD. Furthermore, the increasing use of silver in biomedical and consumer products require better understanding of the effect of this element on the fate of kidney stem cells. The results of this study showed that the AgNPs@MSA containing substrates (SL and SH) affect the viability, spreading, proliferation, population doubling time and differentiation of mKSCs to podocytes and PTCs as evaluated from the expression of markers. Bioreactors, cell culture substrates, cell therapies and tissue engineering constructs may be able to benefit from the antibacterial effect of silver nanoparticles and their capacity to stimulate mKSCs differentiation. However, before the later becomes a real opportunity much further work is required to fully and mechanistically understand how and in what direction silver nanoparticles affect the fate of kidney cells. Whether, the observed increased cell differentiation is due to surface nanoroughness or due to generation of reactive oxygen species, the combined effect of both, or due to other factors coming into play will be the subject of future studies.Conflicts of interest
There are no conflicts to declare.Acknowledgements
N. R. C gratefully acknowledges the Australian Government Research Training Program Scholarship from the Australian Federal Government and UniSA for International Travel Award. K. V. thanks ARC for DP15104212, NHMRC for Fellowship APP1122825 and Project grant APP1032738, and the Alexander von Humboldt Foundation for Fellowship for Experienced Researchers.References
- K. Chaloupka, Y. Malam and A. M. Seifalian, Trends Biotechnol., 2010, 28, 580–588 CrossRef PubMed.
- X. Chen and H. J. Schluesener, Toxicol. Lett., 2008, 176, 1–12 CrossRef PubMed.
- S. Chernousova and M. Epple, Angew. Chem., Int. Ed., 2013, 52, 1636–1653 CrossRef PubMed.
- B. Schäfer, J. Tentschert and A. Luch, Environ. Sci. Technol., 2011, 45, 7589–7590 CrossRef PubMed.
- B. Schäfer, J. Vom Brocke, A. Epp, M. Götz, F. Herzberg, C. Kneuer, Y. Sommer, J. Tentschert, M. Noll and I. Günther, Arch. Toxicol., 2013, 87, 2249–2262 CrossRef PubMed.
- S. Taheri, A. Cavallaro, S. N. Christo, L. E. Smith, P. Majewski, M. Barton, J. D. Hayball and K. Vasilev, Biomaterials, 2014, 35, 4601–4609 CrossRef PubMed.
- S. Taheri, K. Vasilev and P. Majewski, Recent Pat. Mater. Sci., 2015, 8, 166–175 CrossRef.
- J. W. Alexander, Surg. Infect., 2009, 10, 289–292 CrossRef PubMed.
- P. V. AshaRani, G. Low Kah Mun, M. P. Hande and S. Valiyaveettil, ACS Nano, 2009, 3, 279–290 CrossRef PubMed.
- R. Foldbjerg, D. A. Dang and H. Autrup, Arch. Toxicol., 2011, 85, 743–750 CrossRef PubMed.
- C. Greulich, D. Braun, A. Peetsch, J. Diendorf, B. Siebers, M. Epple and M. Koller, RSC Adv., 2012, 2, 6981–6987 RSC.
- W. He, T. A. Elkhooly, X. Liu, A. Cavallaro, S. Taheri, K. Vasilev and Q. Feng, J. Mater. Chem. B, 2016, 4, 1466–1479 RSC.
- C. Zoccali, A. Kramer and K. J. Jager, NDT Plus, 2009, 3, 213–224 Search PubMed.
- P. Murray, K. Vasilev, C. F. Mora, E. Ranghini, H. Tensaout, A. Rak-Raszewska, B. Wilm, D. Edgar, R. D. Short and S. E. Kenny, Biochem. Soc. Trans., 2010, 38, 1062–1066 CrossRef PubMed.
- M. H. Little, Cell Death Dis., 2016, 2, 16053 CrossRef PubMed.
- M. H. Little and A. P. McMahon, Cold Spring Harbor Perspect. Biol., 2012, 4, a008300 Search PubMed.
- N. Nakhoul and V. Batuman, in Experimental Models for Renal Diseases, Karger Publishers 2011, vol. 169, pp. 37–50 Search PubMed.
- H. Pavenstädt, W. Kriz and M. Kretzler, Physiol. Rev., 2003, 83, 253–307 CrossRef PubMed.
- C. Fuente Mora, E. Ranghini, S. Bruno, B. Bussolati, G. Camussi, B. Wilm, D. Edgar, S. E. Kenny and P. Murray, Stem Cells Dev., 2011, 21, 296–307 CrossRef PubMed.
- E. Ranghini, C. F. Mora, D. Edgar, S. E. Kenny, P. Murray and B. Wilm, PLoS One, 2013, 8, e62953 CrossRef PubMed.
- W. Chen, L. G. Villa-Diaz, Y. Sun, S. Weng, J. K. Kim, R. H. W. Lam, L. Han, R. Fan, P. H. Krebsbach and J. Fu, ACS Nano, 2012, 6, 4094–4103 CrossRef PubMed.
- L. C. Y. Lee, N. Gadegaard, M. C. de Andrés, L.-A. Turner, K. V. Burgess, S. J. Yarwood, J. Wells, M. Salmeron-Sanchez, D. Meek, R. O. C. Oreffo and M. J. Dalby, Biomaterials, 2017, 116, 10–20 CrossRef PubMed.
- M. MacGregor-Ramiasa, I. Hopp, A. Bachhuka, P. Murray and K. Vasilev, Acta Biomater., 2017, 56, 171–180 CrossRef PubMed.
- K. Vasilev, A. Michelmore, P. Martinek, J. Chan, V. Sah, H. J. Griesser and R. D. Short, Plasma Processes Polym., 2010, 7, 824–835 CrossRef.
- J. E. Sader, J. W. Chon and P. Mulvaney, Rev. Sci. Instrum., 1999, 70, 3967–3969 CrossRef.
- S. Shankland, J. Pippin, J. Reiser and P. Mundel, Kidney Int., 2007, 72, 26–36 CrossRef PubMed.
- R. V. Goreham, A. Mierczynska, L. E. Smith, R. Sedev and K. Vasilev, RSC Adv., 2013, 3, 10309–10317 RSC.
- J. C. Ruiz, S. Taheri, A. Michelmore, D. E. Robinson, R. D. Short, K. Vasilev and R. Förch, Plasma Processes Polym., 2014, 11, 888–896 CrossRef.
- J. Hernandez-Lopez, R. Bauer, W.-S. Chang, G. Glasser, D. Grebel-Koehler, M. Klapper, M. Kreiter, J. Leclaire, J.-P. Majoral and S. Mittler, Mater. Sci. Eng., C, 2003, 23, 267–274 CrossRef.
- M. Ramiasa, A. Cavallaro, A. Mierczynska, S. Christo, J. Gleadle, J. Hayball and K. Vasilev, Chem. Commun., 2015, 51, 4279–4282 RSC.
- D. L. Schmidt, C. E. Coburn, B. M. DeKoven, G. E. Potter, G. F. Meyers and D. A. Fischer, Nature, 1994, 368, 39–41 CrossRef.
- G. Tillet, B. Boutevin and B. Ameduri, Prog. Polym. Sci., 2011, 36, 191–217 CrossRef.
- C. Faul, M. Donnelly, S. Merscher-Gomez, Y. H. Chang, S. Franz, J. Delfgaauw, J.-M. Chang, H. Y. Choi, K. N. Campbell and K. Kim, Nat. Med., 2008, 14, 931–938 CrossRef PubMed.
- P. Mundel and J. Reiser, Kidney Int., 2010, 77, 571–580 CrossRef PubMed.
- J. Reiser, V. Gupta and A. D. Kistler, Kidney Int., 2010, 77, 662–668 CrossRef PubMed.
- L. Perico, S. Conti, A. Benigni and G. Remuzzi, Nat. Rev. Nephrol., 2016, 12, 692–710 CrossRef PubMed.
- X. Tian and S. Ishibe, Nephrol. Dial. Transplant., 2016, 31, 1577–1583 CrossRef PubMed.
- C. Wei, C. C. Möller, M. M. Altintas, J. Li, K. Schwarz, S. Zacchigna, L. Xie, A. Henger, H. Schmid and M. P. Rastaldi, Nat. Med., 2007, 14, nm1696 Search PubMed.
- E. D. Wachsmuth and J. P. Stoye, Histochemistry, 1976, 47, 315–337 Search PubMed.
- Z. Chen, A. Bachhuka, S. Han, F. Wei, S. Lu, R. M. Visalakshan, K. Vasilev and Y. Xiao, ACS Nano, 2017, 11, 4494–4506 CrossRef PubMed.
- S. N. Christo, A. Bachhuka, K. R. Diener, A. Mierczynska, J. D. Hayball and K. Vasilev, Adv. Healthcare Mater., 2016, 5, 956–965 CrossRef PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c8ra02145g |
| This journal is © The Royal Society of Chemistry 2018 |