Strigolactones: a plant phytohormone as novel anti-inflammatory agents†
Jun-Xia
Zheng‡
a,
Yu-Shui
Han‡
c,
Jin-Cai
Wang‡
 c,
Hui
Yang
c,
Hao
Kong
c,
Kang-Jia
Liu
c,
Si-Yu
Chen
c,
Yi-Rui
Chen
b,
Yi-Qun
Chang
c,
Wei-Min
Chen
c,
Hui
Yang
c,
Hao
Kong
c,
Kang-Jia
Liu
c,
Si-Yu
Chen
c,
Yi-Rui
Chen
b,
Yi-Qun
Chang
c,
Wei-Min
Chen
 c,
Jia-Liang
Guo
*b and
Ping-Hua
Sun
*c
c,
Jia-Liang
Guo
*b and
Ping-Hua
Sun
*c
aSchool of Chemical Engineering and Light Industry, Guangdong University of Technology, Guangzhou, PR China
bSchool of Stomatology and Medicine, Foshan University, Foshan, PR China. E-mail: janalguo@126.com; Tel: +86 2085224497
cCollege of Pharmacy, Jinan University, Guangzhou, PR China
First published on 5th December 2017
Abstract
Strigolactones (SLs) are a novel class of plant hormones with enormous potential for the prevention and treatment of inflammation. To further investigate the anti-inflammatory activities of SLs, a representative SL, GR24, and the reductive products of its D-ring were synthesized and their anti-inflammatory activities were fully evaluated on both in vitro and in vivo models. Among these compounds, the two most active optical isomers (2a and 6a) demonstrated strong inhibitory activity on the release of inflammatory cytokines, including nitric oxide (NO), tumor necrosis factor-alpha (TNF-α), and interleukin-6 (IL-6) by blocking the nuclear factor kappa B (NF-κB) and mitogen-activated protein kinase (MAPK) signaling pathways; they also greatly inhibited the migration of neutrophils and macrophages in fluorescent protein labeled zebrafish larvae. These results identified the promising anti-inflammatory effects of SLs, and suggested that both the absolute configuration of SL and the α,β-unsaturated D-ring structure are essential for the observed anti-inflammatory activity.
Introduction
Multifaceted biological functions of massive phytohormones such as abscisic acids (ABA),1 methyl jasmonates (MJ),2 and cytokinins (CTK)3,4 in mammals were recognized, which indicated high potential for application in the treatment of various clinical diseases. It appears these phytohormones could be an excellent source for the discovery of potential drugs from food and plants. Strigolactones (SLs) are a novel family of plant hormones produced in potential host plants such as wheat, rice, and corn, which control shoot branching architecture by inhibiting growth and self renewal of axillary meristem cells.5 Until now, most initial studies focused on the bioactivity of SLs as a germination stimulant of parasitic seed weeds.6–9 Interestingly, Yarden et al. recently reported that SL and its analogues are potent inhibitors of mammosphere formation and cancer stem-like cell survival, which opens up the possibility for the discovery of promising anticancer agents among SLs.10–12 It is well-known that inflammation is an immune response initiated by pathogen invasion or tissue injury. An increasing number of studies have realized that inflammation is invariably accompanied by various diseases, such as cancer,13,14 atherosclerosis,15,16 and diabetes.17,18 However, no study related to the promising anti-inflammatory effects of SLs have been reported.To date, more than 15 natural SLs have been isolated from plant root extracts or exudates.19 Based on the structure–activity relationships, a number of synthetic SL analogues, known as GR compounds, were designed and synthesized. Among them, GR24, a synthetic analogue with a simpler structure, is the most active and widely applicable as a reference compound for research into parasitic weeds, such as Striga and Orobanche, in bioassays.20–23 Since it exhibits inhibitory effects toward breast cancer cell lines growth and survival,10 GR24 has become an interesting leading compound and research hotspot in pharmaceutical studies.
Therefore, in the present study, (±)-GR24 and the reductive products of its D-ring were synthesized and their anti-inflammatory effects were evaluated on both in vitro and in vivo models, followed by an in-depth study of their mechanism by the western-blot analysis. The toxicity on cells was also evaluated using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. The results revealed the promising anti-inflammatory activity of GR24, particularly two most active optical isomers (2a and 6a), which greatly inhibited the release of TNF-α and NO in LPS-activated murine macrophages by affecting the MAPK and NF-κB pathways. These findings provide support for the potential pharmaceutical application of SLs as novel anti-inflammatory compounds.
Results and discussion
Preparation of optically active synthetic GR24 derivatives
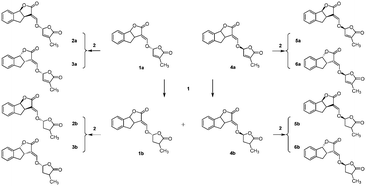
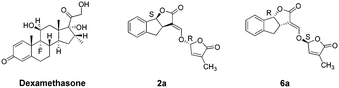
The synthetic routes to obtain GR24 followed previously reported methods with some improvements to increase the yield (Scheme S1†). The synthesis of the GR24 enantiomers (2a, 3a, 5a, and 6a) is shown in Scheme 1; in order to investigate the effect of the double bond in the D-ring, the reduction products, GR24 derivatives (1b and 4b), were obtained by a chemical reduction method from 1a and 4a, respectively. The optically active GR24 derivatives (2b, 3b, 5b, and 6b) were separated by chiral column chromatography. All the GR24 enantiomers were confirmed by 1H NMR, 13C NMR, and MS (shown in ESI†). The structures of 2a, 6a, and positive control dexamethasone are shown in Fig. 1.Inhibition of NO production in (LPS)-stimulated RAW264.7 cells
Nitric oxide (NO) acts as a significant pro-inflammatory mediator for acute or chronic inflammation. It is generally reported that NO inhibitors can offer potential opportunities to develop new therapeutic methods for inflammatory diseases.24,25 In this study, the inhibitory activity of the synthesized strigolactone compounds against NO release was first investigated in lipopolysaccharide (LPS)-stimulated RAW264.7 cells. As depicted in Table 1, a novel class of GR24 derivatives (1a–6a) displayed improved NO inhibitory activities when compared to dexamethasone. In particular, series a (1a–6a) exhibited better activities than series b (1b–6b). Among these, compounds 2a, 2b, 6a, and 6b showed better activities than the corresponding compounds 3a, 3b, 5a, and 5b.| Compounds | IC50a (μM) | Compounds | IC50a (μM) |
|---|---|---|---|
| a Results are shown as mean ± SD (n = 4) of at least three independent experiments. b Dex: dexamethasone. | |||
| Dexb | 2.66 ± 0.50 | 1b | 42.05 ± 5.77 |
| 1a | 4.06 ± 0.47 | 2b | 32.27 ± 4.23 |
| 2a | 3.14 ± 0.24 | 3b | 60.45 ± 6.54 |
| 3a | 6.34 ± 0.76 | 4b | 44.48 ± 4.72 |
| 4a | 4.32 ± 0.39 | 5b | 64.59 ± 4.32 |
| 5a | 6.62 ± 0.51 | 6b | 35.44 ± 2.83 |
| 6a | 3.51 ± 0.22 | ||
This study confirmed that the unreduced D-ring of strigolactones plays an essential role in anti-inflammation bioactivity. Compounds 2a and 2b and 3a and 3b have the (8bS, 2′R) and (8bR, 2′R)-configuration, respectively. In contrast, compounds 6a and 6b and 5a and 5b have the configuration of (8bR, 2′S) and (8bS, 2′S), respectively. Thus, we can speculate that the α,β-unsaturated furanones, including strigolactones with their D-ring, are important chemical pharmacophores. Moreover, compounds 3a, 3b, 5a, and 5b showed weak bioactivity in vitro because of the steric-hindrance.
Cytotoxicity in RAW264.7 cells
To determine whether the NO inhibitory activities of 2a and 6a were related to cell viabilities, further studies were carried out on the effect of these compounds upon RAW264.7 cell growth by adopting the MTT assay method.26 As observed from Table 2, the relative cell viabilities of RAW264.7 were all above 99%, which indicated that no evident cytotoxicity of the tested agents (dexamethasone, 2a, 6a, and LPS) were observed at the detected concentrations. This study was a further indication that NO inhibitory effects of 2a and 6a were probably attributed to the structure of these two compounds. These non-toxic concentrations were also used in the following experiments.| Compounds | Concentrations | Cell viabilitya (%) |
|---|---|---|
| a Results are expressed as mean ± SD (n = 5) of three independent experiments. b Blank: the cells cultured with fresh medium only. c Dex: dexamethasone. | ||
| Blankb | 100.0 ± 3.5 | |
| Dexc | 10 μM | 100.2 ± 2.1 |
| 2a | 10 μM | 100.2 ± 1.6 |
| 6a | 10 μM | 99.2 ± 2.1 |
| LPS | 500 ng mL−1 | 99.8 ± 2.0 |
| LPS | 100 ng mL−1 | 101.5 ± 4.2 |
Inhibition of TNF-α and IL-6 production in RAW264.7 cells
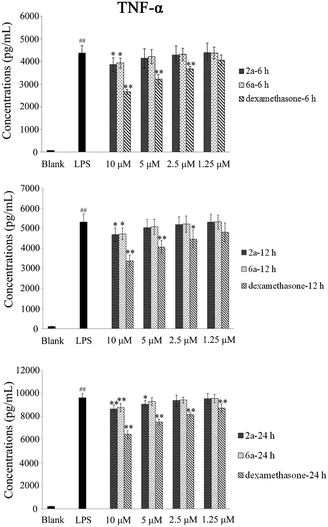
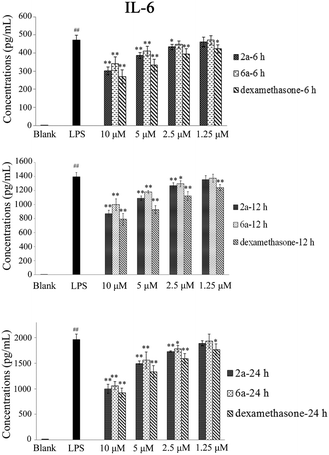
TNF-α and IL-6 play an important role in many biological processes and have been proposed as the main contributing factors to induce inflammatory diseases.27,28 The inhibitory effect of 2a and 6a on the production of pro-inflammatory cytokines is one of the major and effective methods for treatment of inflammation. Consequently, the expression of TNF-α and IL-6 cytokines in RAW264.7 cells was investigated and measured by ELISA.29 Dexamethasone was also used as a positive reference drug. The effects of these three compounds (2a, 6a, and dexamethasone) were evaluated at concentrations of 10 μM, 5 μM, 2.5 μM and 1.25 μM in a time-dependent manner.According to the results (Tables 3 and 4, Fig. 2, and 3), the levels of TNF-α and IL-6 in LPS-stimulated RAW264.7 cells were significantly decreased by 2a and 6a treatment in concentration- and time-dependent manners as compared with that in RAW264.7 control cells, and their inhibitory effects were slightly weaker than that of the positive control drug treatment on both TNF-α and IL-6 production at all three detection times (6 h, 12 h, and 24 h). For example, compounds 2a, 6a, and dexamethasone at 10 μM could significantly reduce the production of LPS-induced inflammation cytokine IL-6 with inhibition index of 1002.2 ± 87.5 pg mL−1, 1061.3 ± 82.1 pg mL−1, and 924.1 ± 84.1 pg mL−1, respectively. Hence, the above results obtained in our laboratory studies suggested that 6a, and particularly 2a, were excellent agents, which showed significant anti-inflammation effect. Next, in order to further examine the regulatory effect of 2a and 6a on inflammatory response, experiments were performed to explore the possible anti-inflammatory mechanism.
| TNF-αa (pg mL−1) | ||||
|---|---|---|---|---|
| Compounds | 6 h | 12 h | 24 h | |
| ## P < 0.01, #P < 0.05 versus the blank (cultured with fresh medium only) group; **P < 0.01, *P < 0.05 versus the LPS (treated with LPS only) group. a Results are shown as mean ± SD (n = 3) of three independent experiments. b Dex: dexamethasone. | ||||
| Blank | 75.9 ± 6.5 | 107.2 ± 10.4 | 204.1 ± 23.8 | |
| LPS | 4377.1 ± 318.6## | 5313.7 ± 420.9## | 9635.7 ± 377.3## | |
| LPS + Dexb | 10 μM | 2655.0 ± 106.2** | 3393.5 ± 280.9** | 6458.3 ± 362.9** |
| 5 μM | 3219.4 ± 208.8** | 4063.6 ± 326.8** | 7533.7 ± 260.2** | |
| 2.5 μM | 3671.8 ± 97.6** | 4455.7 ± 446.9* | 8187.9 ± 184.9** | |
| 1.25 μM | 4056.3 ± 231.3 | 4810.1 ± 462.8 | 8748.1 ± 370.5** | |
| LPS + 2a | 10 μM | 3862.3 ± 300.1* | 4685.5 ± 348.6* | 8660.7 ± 363.2** |
| 5 μM | 4140.7 ± 427.4 | 5036.1 ± 413.2 | 9085.5 ± 327.1* | |
| 2.5 μM | 4276.9 ± 415.4 | 5186.9 ± 386.2 | 9397.7 ± 483.9 | |
| 1.25 μM | 4388.3 ± 425.2 | 5303.7 ± 405.9 | 9565.4 ± 450.4 | |
| LPS + 6a | 10 μM | 3927.4 ± 222.8* | 4721.3 ± 286.7* | 8796.7 ± 342.8** |
| 5 μM | 4209.3 ± 316.3 | 5079.1 ± 376.0 | 9318.5 ± 348.4 | |
| 2.5 μM | 4307.9 ± 271.9 | 5212.4 ± 404.2 | 9444.5 ± 260.6 | |
| 1.25 μM | 4367.3 ± 264.7 | 5319.6 ± 354.3 | 9582.9 ± 362.0 | |
| IL-6a (pg mL−1) | ||||
|---|---|---|---|---|
| Compounds | 6 h | 12 h | 24 h | |
| ## P < 0.01, #P < 0.05 versus the blank (cultured with fresh medium only) group; **P < 0.01, *P < 0.05 versus the LPS (treated with LPS only) group. a Results are shown as mean ± SD (n = 3) of three independent experiments. b Dex: dexamethasone. | ||||
| Blank | 2.3 ± 0.5 | 6.8 ± 0.7 | 12.4 ± 1.9 | |
| LPS | 472.3 ± 25.8## | 1394.0 ± 62.6## | 1966.6 ± 102.3## | |
| LPS + Dexb | 10 μM | 269.5 ± 38.9** | 791.0 ± 82.9** | 924.1 ± 84.1** |
| 5 μM | 332.8 ± 32.8** | 927.7 ± 57.4** | 1336.5 ± 117.4** | |
| 2.5 μM | 393.4 ± 30.9** | 1120.3 ± 60.7** | 1594.9 ± 95.4** | |
| 1.25 μM | 422.7 ± 23.0* | 1240.5 ± 42.8** | 1770.4 ± 107.2* | |
| LPS + 2a | 10 μM | 302.3 ± 21.8** | 869.0 ± 47.2** | 1002.2 ± 87.5** |
| 5 μM | 387.1 ± 15.0** | 1090.7 ± 30.2** | 1499.2 ± 43.3** | |
| 2.5 μM | 433.6 ± 14.2* | 1267.7 ± 40.5** | 1729.3 ± 11.3** | |
| 1.25 μM | 461.2 ± 27.3 | 1351.9 ± 57.7 | 1892.3 ± 47.2 | |
| LPS + 6a | 10 μM | 340.6 ± 37.8** | 998.2 ± 84.1** | 1061.3 ± 82.1** |
| 5 μM | 410.3 ± 26.2** | 1177.0 ± 21.7** | 1561.1 ± 162.1** | |
| 2.5 μM | 447.9 ± 19.7 | 1296.0 ± 42.8* | 1787.6 ± 65.3* | |
| 1.25 μM | 470.1 ± 25.7 | 1374.3 ± 56.2 | 1936.5 ± 137.1 | |
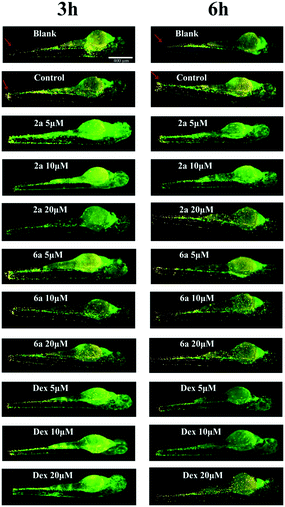
Effects of GR24 on migration of neutrophils and primitive macrophages in the injuries of transgenic zebrafish larvae
In order to further evaluate anti-inflammatory activity of GR24 derivatives, injured transgenic zebrafish larvae were used as in vivo models for the next step. To the best of our knowledge, neutrophils and macrophages are two most prominent types of inflammatory cells in the living body. Once inflammation occurs in vivo, higher quantities of these are produced and transported to the inflammation sites. In this study, the anti-inflammatory effects of 2a and 6a were investigated using a double-transgenic line (Coronilla-eGFP/Lyc-dsRed), in which neutrophils were double-labeled by eGFP and dsRed, presenting yellow colour, while the primitive macrophages were labeled by eGFP only, presenting green color.The generation of injury inflammation was induced by tail-cutting; dexamethasone was used as the control drug. As shown in Fig. 4, the blank group without tail cutting, which were the healthy larvae, always generated a clear image. However, after tail cutting, for 3 h and 6 h, the control group generated a fluorescence image, in which the inflammatory cells, neutrophils and primitive macrophages, clearly accumulated around the wound and could be visualized. Compared to the control group, the level of neutrophils and primitive macrophages around the wound slightly reduced after treatment with 5 μM of 2a and 6a, but significantly reduced after treatment with 10 and 20 μM dosages during the evaluation time, which is similar to the positive control group. This is also consistent with the behavior described in our early reports.30 In summary, all these results illustrate the in vivo anti-inflammation activity of the GR24 derivatives, 2a and 6a. Therefore, further efforts are essential to explore their possible anti-inflammatory mechanism.
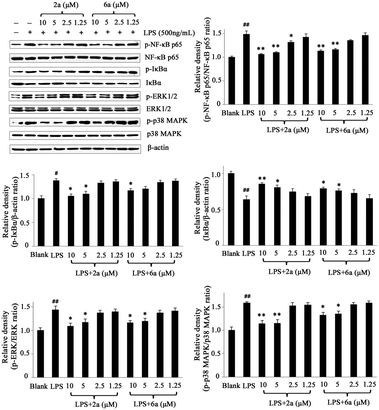
Western blot for interpreting possible mechanism
It has been reported that the NF-κB and MAPK pathways are quite paramount and significant in inflammation by interplaying with various inflammatory diseases because of their crucial roles in the regulation of the intracellular and extracellular production of inflammatory cytokines in activated macrophages, such as NO, TNF-α, and IL-6.31–33 Accumulating literature reports indicated that the inflammatory mediators induced by LPS could activate the NF-κB and MAPK signaling pathways.34,35Therefore, in order to investigate whether the anti-inflammatory effects of 2a and 6a were associated with the activation of NF-κB and MAPK, western blot analysis was carried out. Herein, we detected two NF-κB pathway proteins, NF-κB p65 and inhibitor kappa B alpha (IκBα) kinase36–38 as well as two MAPK cascades, extracellular regulated protein kinases (ERK1/2, or p42/44) and p38 MAPK.39–41 As shown in Fig. 5, the levels of phosphorylated NF-κB p65, IκBα, ERK1/2, and p38 MAPK significantly increased by treatment of LPS and decreased in varying degrees in RAW264.7 cells by 2a and 6a treatment. Furthermore, we observed that the phosphorylation of NF-κBp65, IκBα, ERK1/2, and p38 MAPK were significantly inhibited after treatment with 2a and 6a in a concentration dependent manner when compared with LPS-induced macrophages, while they had a slightly antagonistic effect on the phosphorylation of IκBα. Collectively, it can be concluded that the anti-inflammation effects of 2a and 6a were due to, in part, their ability to suppress the activation of NF-κB and MAPK cascades, resulting in decreased NO, TNF-α, and IL-6 levels.
To further investigate the anti-inflammatory mechanism of GR24, reverse docking of 2a in the NF-κB signaling pathway was performed. From the PDB website, 62 proteins of NF-κB were obtained, and its protein target structure database was built (Table S1†). The molecular docking software AutoDock Vina was used to dock 2a with the protein targets. According to the docking scores and the docking conformational interaction, the potential targets were selected. A total of 3 target proteins, PARP1, CK2, and AKT were found to interact with GR24 with high affinity. By comparison of their docking conformational interaction (Fig. S1–S3†), PARP1 is superior to CK2 and AKT. Anti-inflammation of GR24 relates to its function of competitive inhibition of the 3 target proteins, which provides an important reference for the study of the anti-inflammatory mechanism of strigolactone and its analogues. However, it also requires further confirmation.
Conclusions
In conclusion, a novel class of synthetic GR24 derivatives was synthesized and their bioactivities were evaluated. To the best of our knowledge, the anti-inflammatory activity of SLs and their derivatives was studied for the first time. Among these SLs, two promising compounds (2a and 6a) were identified for the treatment of inflammatory diseases. The in vitro anti-inflammatory experiments revealed a similar or even better inhibitory effect on the release of inflammatory mediators and cytokines (NO, TNF-α, and IL-6) as compared to dexamethasone. Moreover, the migration of neutrophils and primitive macrophages were also measured in an in vivo zebrafish model; the results also revealed strong inhibition by 2a and 6a. It was demonstrated that the anti-inflammatory activity of 2a and 6a was due to, at least in part, their ability to suppress the activation of NF-κB and MAPK cascades, resulting in decreased NO, TNF-α, and IL-6 levels. These results further confirm that compounds with the unreduced D-ring generally exhibit improved anti-inflammatory activity, which suggests that the reserved D-ring would be an effective strategy to develop anti-inflammatory agents. Further studies would be undertaken in our group in order to evaluate the possible anti-inflammatory action targets of these SLs.Experimental
Apparatus
1H and 13C NMR spectra were recorded in CDCl3 on a Bruker 300 spectrometer (Karlsruhe, Germany) using tetramethylsilane (TMS) as an internal standard. Chemical shifts were measured in δ (ppm). Mass spectra were recorded on a Thermo Finnigan LCQ Advantage MAX mass spectrometer (Applied Biosystems, 4000 Q TRAP). Optical rotation was obtained on a SEPA-300 digital polarimeter (Horiba, Tokyo, Japan). Melting points were determined with a Reichert Thermopan microscope and are uncorrected. Kieselgel 60 F254 (Merck, Germany) was used for column chromatography, and pre-coated silica gel 60 F254 plates were used to analyze thin layer chromatography (TLC). Spots were visualized using a UV lamp. The fluorescence photos of larvae were recorded using a fluorescence microscope (Olympus, Japan).Materials and reagents
RAW264.7 murine macrophages were purchased from the American Type Culture Collection (ATCC) and cultured in DMEM containing 10% FBS. Foetal bovine serum (FBS) and Dulbecco's modified Eagle's medium (DMEM) were purchased from Gibco (Invitrogen, California, USA). Lipopolysaccharide (LPS, Escherichia coli 0127:B8) and MTT were obtained from Sigma-Aldrich (St. Louis, Missouri, USA). Dexamethasone was purchased from the National institutes for food and drug control (Beijing, China). N-(1-Naphthyl) ethylenediamine dihydrochloride and sulfanilamide were purchased from Sinopharm chemical reagent Co. Ltd (Shanghai, China). TNF-α enzyme immunoassay kits (EK2822) and IL-6 enzyme immune assay kits (EK2062) were purchased from Multi Sciences Biotechnology (Hangzhou, China). Phorbol-12-myristate-13-acetate (PMA, S1819), IP cell lysis buffer (P0013), and phenylmethanesulfonyl fluoride (PMSF, ST506) were purchased from Beyotime Biotechnology (Shanghai, China). Anti-beta actin monoclonal antibody (E021020-01) was purchased from EarthOx, LLC, CA. Primary anti-rabbit inducible NO synthase (iNOS, #2982), NF-κB pathway sampler Kit (#9936S), p-MAPK family antibody sampler kit (#9910S), and MAPK family antibody sampler kit (#9926) were purchased from Cell Signaling Technology. BCA protein assay kit (KGPBCA) was purchased from KeyGEN Biotechnology (Nanjing, China). The enhanced chemiluminescent (ECL) Detection Kit (WBKLS0050) was purchased from Merck Millipore Corporation (Merck KGaA, Darmstadt, Germany).Chemicals
(±)-GR24 and its derivatives were synthesized from the commercially available material, indan-1-one, using previously reported methods42–44 as an epimer mixture of 1a and 4a (ca. 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) at C-2′, which were then separated by chromatography on silica gel with hexane/EtOAc (v/v, 7
1) at C-2′, which were then separated by chromatography on silica gel with hexane/EtOAc (v/v, 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3) as the eluent. Then, Pd/C (5 mg) was added to a solution of 1a/4a (50 mg) in ethyl acetate (5 mL). The mixture was stirred under hydrogen atmosphere for 0.5 h at room temperature and filtered; the filtrate was concentrated in vacuo to obtain the crude product, which was purified by flash chromatography on a silica gel column to yield the diastereomer 1b/4b. The optical isomers of 2a and 3a were achieved from compound 1a using an optically active column as reported by Ueno et al.45 Similarly, the isomers of 5a and 6a were also prepared from compound 4a. In addition, a series of optical isomers of 2b, 3b, 5b, and 6b were obtained in the same way (Scheme 1). All products in stock including dexamethasone were dissolved in dimethyl sulfoxide (DMSO) to a concentration of 50 mM and the amount of DMSO was fixed at 0.1% (v/v) when added to cells.
3) as the eluent. Then, Pd/C (5 mg) was added to a solution of 1a/4a (50 mg) in ethyl acetate (5 mL). The mixture was stirred under hydrogen atmosphere for 0.5 h at room temperature and filtered; the filtrate was concentrated in vacuo to obtain the crude product, which was purified by flash chromatography on a silica gel column to yield the diastereomer 1b/4b. The optical isomers of 2a and 3a were achieved from compound 1a using an optically active column as reported by Ueno et al.45 Similarly, the isomers of 5a and 6a were also prepared from compound 4a. In addition, a series of optical isomers of 2b, 3b, 5b, and 6b were obtained in the same way (Scheme 1). All products in stock including dexamethasone were dissolved in dimethyl sulfoxide (DMSO) to a concentration of 50 mM and the amount of DMSO was fixed at 0.1% (v/v) when added to cells.
Cell culture
RAW264.7 murine macrophages cells were routinely cultured in DMEM with 10% foetal calf serum (FCS), 100 units per mL penicillin, and 100 μg mL−1 streptomycin at 37 °C in a 5% CO2 humidified atmosphere.Assay for NO production
RAW264.7 cells, a murine macrophage cell line, were inoculated at 5 × 104 cells per well in a 96-well plate and left to culture for 18 h before any treatment. The concentrations of each compound including the positive drug (dexamethasone) were fixed at 100 μM, 50 μM, 20 μM, 10 μM, 5 μM, 2 μM, and 1 μM. The cells were then pre-treated with different concentrations of compounds for 2 h before stimulation with LPS (100 ng mL−1), while the cells in control group were treated with LPS only. After stimulation for 24 h by LPS, the NO produced in the culture medium was quantified by Greiss reagent. In brief, 100 μL cell culture supernatant was transferred to another 96-well plate; then, 100 μL of Griess reagent was added and incubated for 10 min at room temperature. Then, the absorbance of the samples was measured at 540 nm (OD540) in a microplate reader (Bio-Rad Laboratories, CA, USA). Dexamethasone was used as the positive control. IC50, the half maximal inhibitory concentration, represented the concentration of a compound that is required for inhibition of 50% NO production in RAW264.7 cells and was determined by constructing a dose–response curve in vitro.Cell cytotoxicity
The cell cytotoxicity of the RAW264.7 cells was measured by the MTT assay. Briefly, the cells were seeded at 4 × 103 cells per well in a 96-well plate. After being cultured overnight, the cells were treated with different compounds for 48 h. Then to each well, 20 μL of 0.5 mg mL−1 MTT solution was added. Cells were incubated for 4 h at 37 °C. After 4 h, the cell culture supernatant was removed and then solubilized with 150 μL dimethyl sulfoxide (DMSO) solution. The optical density was measured at 570 nm (OD570) using a Bio-Rad microplate reader. Each experiment was repeated thrice, and the data was collected from the three results. The density of formazan formed in the blank group was set as 100% viability. Cell viability (%) = [Compound (OD570) − Blank (OD570)]/[Control (OD570) − Blank (OD570)] × 100%.Blank: cultured with fresh medium only.
Control: the cells cultured with fresh medium only.
Compound: the cells treated with different compounds.
Measurement of TNF-α and IL-6
The levels of TNF-α and IL-6 were measured using the mouse ELISA kit (TNF-α: MultiSciences, EK2822; IL-6: MultiSciences, EK2062), which was operated according to the manufacturer's instructions. All samples were obtained as follows: RAW264.7 cells (5 × 105 cells per well) were cultured in a 24-well plate and pretreated with 10 μM, 5 μM, 2.5 μM, and 1.25 μM of compounds for 2 h; then, LPS (100 ng mL−1) was added. The production of TNF-α and IL-6 was incubated for 6 h, 12 h, and 24 h.Zebrafish larvae live image analysis
The neutrophil/macrophage double transgenic zebrafish (Coronilla-eGFP/Lyc-dsRed) experiments involving 2a and 6a were performed according to our previous studies.30,46–48 The preparation of four days post fertilization zebrafish larvae, tail-cutting surgery, compound treatment, and image analysis all followed the reported methods with a minor modifications. Dexamethasone was used as the control drug. All experiments were performed in compliance with the relevant laws and institutional guidelines, and were approved by the Zebrafish Research Centre of Southern Medical University in China.Western blot analysis
RAW264.7 cells were seeded into a 6-well culture plate at a density of 2 × 106 cells per well. Then, the culture medium was replaced by fresh medium containing different concentrations of compounds (10 μM, 5 μM, 2.5 μM, and 1.25 μM), followed by the addition of LPS (500 ng mL−1). After being cultured for another 4 h, the cells were harvested and lysed with cell lysates, which contained IP buffer (Beyotime, P0013) and 1 mM phenylmethanesulfonyl fluoride (PMSF: Beyotime, ST506) for 30 min at 4 °C. The cell supernatant was collected by centrifugation at 14000 g for 10 min at 4 °C. Protein concentration was determined using the BCA protein assay kit (Thermo Scientific, 23227). Furthermore, each protein sample was mixed with a quarter volume of 5× SDS-PAGE sample loading buffer and the mixture was boiled for 10 min. Equal amounts of total cellular protein were separated in a 12.5% SDS-PAGE gel and then transferred to polyvinylidenedifluoride membranes (Bio-Rad). After wet transfer, the membranes were blocked for 4 h at room temperature in 5% non-fat dry milk in TBS containing 0.1% Tween 20 (TBST), washed thrice in TBST, incubated with the primary antibody (anti-phosphorylation of ERK1/2 (Thr202/Tyr204), anti-ERK1/2, anti-phosphorylation of p38 (Thr180/Tyr182), anti-p38, anti-phosphorylation of IκBα (Ser32/36), anti-IκBα, anti-phosphorylation of NF-κB p65, and anti-NF-κB p65) at 4 °C overnight (all the primary antibodies were purchased from Cell Signaling Technology and diluted in TBST at a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), washed thrice in TBST, incubated with anti-rabbit or anti-mouse secondary antibody (1
1000), washed thrice in TBST, incubated with anti-rabbit or anti-mouse secondary antibody (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 in TBST, Cell Signaling Technology) for 1.5 h at room temperature, washed in TBST and exposed to ECL reagents.
1000 in TBST, Cell Signaling Technology) for 1.5 h at room temperature, washed in TBST and exposed to ECL reagents.
Statistical analysis
Results are presented as mean ± standard error (SD). Statistical analysis was performed using Student's t-test (two-tailed, paired) versus vehicle controls and regarded as being significant when p < 0.05.Conflicts of interest
The authors declare no competing interests.Acknowledgements
We express our gratitude to the Key Laboratory of Zebrafish Modeling and Drug Screening for Human Diseases of Guangdong Higher Education Institutes, Southern Medical University, for supplements of zebrafish larvae. This research was supported financially by the National Natural Science Foundation of China (Grant: 81503031 and 21405013), the Science and Technology Project of Guangdong Province (Grant: 2016A020226004), and the Initial Scientific Research Fund of Doctor in Foshan University (Grant: gg040952).References
- P. Sakthivel, N. Sharma, P. Klahn, M. Gereke and D. Bruder, Curr. Med. Chem., 2016, 23, 1549–1570 CrossRef CAS PubMed.
- S. Cohen and E. Flescher, Phytochemistry, 2009, 70, 1600–1609 CrossRef CAS PubMed.
- Y. Ishii, S. Sakai and Y. Honma, Biochim. Biophys. Acta, Mol. Cell Res., 2003, 1643, 11–24 CrossRef CAS.
- P. Mlejnek, J. Cell. Biochem., 2001, 83, 678–689 CrossRef CAS PubMed.
- P. B. Brewer, H. Koltai and C. A. Beveridge, Mol. Plant, 2013, 6, 18–28 CrossRef CAS PubMed.
- Y. Kapulnik, P. M. Delaux, N. Resnick, E. Mayzlish-Gati, S. Wininger, C. Bhattacharya, N. Séjalon-Delmas, J. P. Combier, G. Bécard and E. Belausov, Planta, 2011, 233, 209–216 CrossRef CAS PubMed.
- C. Rameau, C. R. Biol., 2010, 333, 344–349 CrossRef CAS PubMed.
- V. Gomez-Roldan, S. Fermas, P. B. Brewer, V. Puech-Pagès, E. A. Dun, J. P. Pillot, F. Letisse, R. Matusova, S. Danoun and J.-C. Portais, Nature, 2008, 455, 189–194 CrossRef CAS PubMed.
- M. Umehara, A. Hanada, S. Yoshida, K. Akiyama, T. Arite, N. Takeda-Kamiya, H. Magome, Y. Kamiya, K. Shirasu and K. Yoneyama, Nature, 2008, 455, 195–200 CrossRef CAS PubMed.
- C. Pollock, H. Koltai, Y. Kapulnik, C. Prandi and R. Yarden, Breast Cancer Res. Treat., 2012, 134, 1041–1055 CrossRef CAS PubMed.
- C. B. Pollock, S. McDonough, V. S. Wang, H. Lee, L. Ringer, X. Li, C. Prandi, R. J. Lee, A. S. Feldman and H. Koltai, Oncotarget, 2014, 5, 1683–1698 Search PubMed.
- E. Mayzlish-Gati, D. Laufer, C. F. Grivas, J. Shaknof, A. Sananes, A. Bier, S. Ben-Harosh, E. Belausov, M. D. Johnson and E. Artuso, Cancer Biol. Ther., 2015, 16, 1682–1688 CrossRef CAS PubMed.
- J. K. Kundu and Y. J. Surh, Mutat. Res., Rev. Mutat. Res., 2008, 659, 15–30 CrossRef CAS PubMed.
- S. I. Grivennikov, F. R. Greten and M. Karin, Cell, 2010, 140, 883–899 CrossRef CAS PubMed.
- P. Libby, P. M. Ridker and A. Maseri, Circulation, 2002, 105, 1135–1143 CrossRef CAS PubMed.
- P. Libby, Arterioscler., Thromb., Vasc. Biol., 2012, 32, 2045–2051 CrossRef CAS PubMed.
- A. S. Major and D. G. Harrison, Circulation, 2011, 124, 2809–2811 CrossRef PubMed.
- W. Xie and L. Du, Diabetes, Obes. Metab., 2011, 13, 289–301 CrossRef CAS PubMed.
- X. Xie, K. Yoneyama and K. Yoneyama, Annu. Rev. Phytopathol., 2010, 48, 93–117 CrossRef CAS PubMed.
- E. Dor, D. M. Joel, Y. Kapulnik, H. Koltai and J. Hershenhorn, Planta, 2011, 234, 419–427 CrossRef CAS PubMed.
- C. Bhattacharya, P. Bonfante, A. Deagostino, Y. Kapulnik, P. Larini, E. G. Occhiato, C. Prandi and P. Venturello, Org. Biomol. Chem., 2009, 7, 3413–3420 CAS.
- A. S. Mwakaboko and B. Zwanenburg, Bioorg. Med. Chem., 2011, 19, 5006–5011 CrossRef CAS PubMed.
- G. H. Nefkens, J. W. J. Thuring, M. F. Beenakkers and B. Zwanenburg, J. Agric. Food Chem., 1997, 45, 2273–2277 CrossRef CAS.
- L. S. Tedesco, J. Fuseler, M. Grisham, R. Wolf and S. C. Roerig, Pain, 2002, 95, 215–223 CrossRef CAS PubMed.
- H. Y. Tseng, S. H. Wu, W.-H. Huang, S. F. Wang, Y. N. Yang, N. Mahindroo, T. Hsu, W. T. Jiaang and S. J. Lee, Bioorg. Med. Chem. Lett., 2005, 15, 2027–2032 CrossRef CAS PubMed.
- M. Taira, K. Sasaki, S. Saitoh, T. Nezu, M. Sasaki, S. Kimura, K. Terasaki, K. Sera, T. Narushima and Y. Araki, Dent. Mater. J., 2006, 25, 726–732 CrossRef CAS.
- R. Sharma, S. Agrawal, A. Saxena and R. Sharma, J. Interferon Cytokine Res., 2013, 33, 384–391 CrossRef CAS PubMed.
- K. Popko, E. Gorska, A. Stelmaszczyk-Emmel, R. Plywaczewski, A. Stoklosa, D. Gorecka, B. Pyrzak and U. Demkow, Eur. J. Med. Res., 2010, 15, 120–122 Search PubMed.
- J. Bienvenu, L. Coulon, C. Doche, M. Gutowski and G. Grau, Eur. Cytokine Network, 1993, 4, 447–451 CAS.
- M. Y. Huang, J. Lin, Z. J. Huang, H. G. Xu, J. Hong, P. H. Sun, J. L. Guo and W. M. Chen, MedChemComm, 2016, 7, 658–666 RSC.
- W. Li, H. Huang, Y. Zhang, T. Fan, X. Liu, W. Xing and X. Niu, Eur. J. Pharmacol., 2013, 715, 62–71 CrossRef CAS PubMed.
- C. Xie, J. Kang, M. E. Ferguson, S. Nagarajan, T. M. Badger and X. Wu, Mol. Nutr. Food Res., 2011, 55, 1587–1591 CAS.
- C. Xie, J. Kang, Z. Li, A. G. Schauss, T. M. Badger, S. Nagarajan, T. Wu and X. Wu, J. Nutr. Biochem., 2012, 23, 1184–1191 CrossRef CAS PubMed.
- S. Chen, J. Xiong, Y. Zhan, W. Liu and X. Wang, Cell. Mol. Neurobiol., 2015, 35, 523–531 CrossRef CAS PubMed.
- D. B. Reddy and P. Reddanna, Biochem. Biophys. Res. Commun., 2009, 381, 112–117 CrossRef CAS PubMed.
- T. D. Gilmore, Semin. Cancer Biol., 1997, 8, 61–62 CrossRef PubMed.
- Y. Yamamoto and R. B. Gaynor, J. Clin. Invest., 2001, 107, 135–142 CrossRef CAS PubMed.
- K. Ashikawa, S. Majumdar, S. Banerjee, A. C. Bharti, S. Shishodia and B. B. Aggarwal, J. Immunol., 2002, 169, 6490–6497 CrossRef CAS.
- S. M. Wurgler-Murphy and H. Saito, Trends Biochem. Sci., 1997, 22, 172–176 CrossRef CAS PubMed.
- J. M. Kyriakis and J. Avruch, Physiol. Rev., 2012, 92, 689–737 CrossRef CAS PubMed.
- E. Herlaar and Z. Brown, Mol. Med. Today, 1999, 5, 439–447 CrossRef CAS PubMed.
- S. Wigchert, E. Kuiper, G. Boelhouwer, G. Nefkens, J. Verkleij and B. Zwanenburg, J. Agric. Food Chem., 1999, 47, 1705–1710 CrossRef CAS PubMed.
- J. W. J. Thuring, G. H. Nefkens and B. Zwanenburg, J. Agric. Food Chem., 1997, 45, 2278–2283 CrossRef CAS.
- E. M. Mangnus, F. J. Dommerholt, R. L. De Jong and B. Zwanenburg, J. Agric. Food Chem., 1992, 40, 1230–1235 CrossRef CAS.
- K. Ueno, M. Fujiwara, S. Nomura, M. Mizutani, M. Sasaki, H. Takikawa and Y. Sugimoto, J. Agric. Food Chem., 2011, 59, 9226–9231 CrossRef CAS PubMed.
- M. Y. Huang, J. Lin, K. Lu, H. G. Xu, Z. Z. Geng, P. H. Sun and W. M. Chen, J. Agric. Food Chem., 2016, 64, 2893–2900 CrossRef CAS PubMed.
- H. B. Zhou, D. T. Yang, N. E. Lvleva, N. E. Mircescu, R. Niessner and C. Haisch, Anal. Chem., 2014, 86, 1525–1533 CrossRef CAS PubMed.
- H. B. Zhou, D. T. Yang, N. P. Lvleva, N. E. Mircescu, S. Schubert, R. Niessner, A. Wieser and C. Haisch, Anal. Chem., 2015, 87, 6553–6561 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c7md00461c |
| ‡ These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2018 |