Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Quantitative evaluation and in vivo visualization of mercury ion bioaccumulation in rotifers by novel aggregation-induced emission fluorogen nanoparticles†
Yusheng
Jiang‡
abc,
Tao
He‡
bd,
Yuncong
Chen
e,
Yinlan
Ruan
f,
Yabin
Zhou
c,
Ben Zhong
Tang
*e,
Jianguang
Qin
*b and
Youhong
Tang
 *c
*c
aCollege of Aquaculture and Life Sciences, Dalian Ocean University, Liaoning 116023, China
bCollege of Science and Engineering, Flinders University, South Australia 5042, Australia. E-mail: jian.qin@flinders.edu.au; Tel: +61 8 82013045
cCentre for NanoScale Science and Technology, Flinders University, South Australia 5042, Australia. E-mail: youhong.tang@flinders.edu.au; Tel: +61 8 82012138
dCollege of Animal Science and Technology, Southwest University, Chongqing 400716, China
eDepartment of Chemistry, The Hong Kong University of Science and Technology, Hong Kong, China. E-mail: tangbenz@ust.hk; Tel: +852 2358 7375
fInstitute for Photonics and Advanced Sensing, School of Physical Sciences, University of Adelaide, South Australia 5005, Australia
First published on 12th September 2017
Abstract
In this study, a specifically-designed aggregation-induced emission fluorogen (AIEgen) with nanoparticle aggregates was used to quantitatively evaluate the bioaccumulation of Hg2+ and visualize Hg2+ kinetics in vivo within the rotifer Brachionus plicatilis for the first time. Quantitative results showed that a sharp drop in Hg2+ concentration occurred at the very beginning in the medium containing rotifers and Hg2+, showing a quick initial uptake of Hg2+ by the rotifers, and then the concentration in the medium plateaued after 5 min. With an increase in rotifer density, the amount of bioaccumulation increased in the rotifer. However, the bioaccumulation efficiency of Hg2+ decreased from 5.28 μg mg−1 h−1 at a low rotifer density of 0.093 mg ml−1 to 2.61 μg mg−1 h−1 at a high rotifer density of 0.375 mg mL−1. Moreover, the fluorescence images and spectra results illustrate that the ingestion of Hg2+ by the rotifer was via its mouth surrounded by the ciliary corona to the digestive tract, and Hg2+ could not permeate into the body integument through diffusion during the study period. Hg2+-induced fluorescence in rotifers dissipated in 6 h after staining, possibly through defecation and excretion. This study indicates that inorganic mercury can be quickly ingested by a rotifer via feeding, but is unlikely deposited as methylated mercury in rotifer tissues.
Environmental significanceThe bioaccumulation of Hg2+ in aquatic organisms is astonishing. However, traditional methods are unable to quantify mercury dynamics inside small aquatic invertebrates because these methods only provide a “snapshot” of organisms for a particular time and life stage. In this study, a specifically-designed aggregation-induced emission fluorogen with nanoparticle aggregates was used to quantitatively evaluate the bioaccumulation of Hg2+ and visualize Hg2+ kinetics in vivo within a rotifer for the first time. The results illustrated that Hg2+ could not permeate into the body integument through diffusion, and Hg2+-induced fluorescence in rotifers dissipated after staining, possibly through defecation and excretion during the study period. This study indicates that inorganic mercury can be quickly ingested by a rotifer via feeding, but is unlikely deposited as methylated mercury in rotifer tissues. |
Introduction
Bioaccumulation is the process of gradual build-up of a chemical in a living organism within a trophic level, through which the concentration of the chemical in the organism exceeds that in the medium.1 Some aquatic organisms are able to accumulate, retain and transform toxic chemicals within the body through biochemical and physiological processes especially in a habitat where pollutants prevail due to human activity.2 Toxic substance accumulation can reduce growth and reproductive fitness, and cause genetic mutation and even death of aquatic organisms.2 Therefore, toxic chemicals in algae can be transferred along the food chains to zooplankton, fish, sea mammals, and humans.3Zooplankton is an important trophic link between primary producers and predators in an aquatic system. Rotifers are widely-distributed small invertebrates in water, mainly consuming microalgae and organic detritus, and serve as food for fish, shrimp and crab.4 In nature, rotifers are first level consumers that can accumulate toxic substances by grazing algae at the base of a food chain.5 Rotifers are also important live food for the culture of many aquatic animals in aquaculture farms. In the laboratory, rotifers have been used as a model organism in aquatic toxicological research due to their short reproduction cycle, fast population growth and ease of culture.4
Among all the toxic substances accumulated in aquatic organisms, mercury (Hg) is one of the most dangerous and ubiquitous heavy metal pollutants in water, and is ranked the third after arsenic and lead in the list of hazardous substances.6 A major concern with Hg contamination is its transfer and biomagnification in aquatic food chains. Mercury exists in many compounds, bonding with other elements such as chlorine, carbon and oxygen, and all these mercury compounds are toxic.6 The mercury ion (Hg2+) can easily pass through the skin and respiratory and gastrointestinal tissues into the human body, causing damage to the central nervous system and endocrine system.7 Moreover, Hg2+ can be converted to methyl mercury (MeHg) by a variety of microorganisms in water to interfere with the normal development of the fetal brain in humans.8
The bioaccumulation of Hg2+ in aquatic organisms is astonishing. However, traditional methods are unable to quantify mercury dynamics inside small aquatic invertebrates because these methods only provide a “snapshot” of organisms for a particular time and life stage, and the transfer mechanism of toxic substances along the food chain is largely unknown, especially in small aquatic organisms.9 Moreover, traditional methods are limited for use in well-established laboratories, due to the requirements of sophisticated instrumentation and labor-intensive processes for sample preparation.10 Therefore, there is a compelling need to develop a new methodology to detect and quantify toxic substances in aquatic organisms in vivo to understand the mechanisms of enrichment and transfer of Hg2+.
Many previous studies have tested Hg2+ transformation in aquatic biota (phytoplankton, zooplankton, and fish) and analyzed the respective Hg2+ concentrations.11 However, these studies did not provide information regarding the uptake and removal kinetics of Hg2+, which are important parameters for interpreting and predicting the food-chain transfer of Hg2+. It remains essentially unknown whether the accumulated Hg2+ in zooplankton originates from medium absorption, ingestion from food, or parental transfer. Empirical evidence exists for the dominance of food for MeHg exposure in fish, but no information is available with regard to zooplankton species.12
Aggregation-induced emission (AIE) refers to a photophysical effect whereby light emission of a fluorogen is activated by aggregate formation to nanoparticles, which may be used as a sensing method in biological applications.13 Fluorogens with AIE effects are known as AIEgens that luminesce more efficiently in the practically useful aggregate state (for example, forming nanoparticles) than in the solution state.14 In an AIE system, the aggregates shine brighter than their individual parts, following the general collective quantity-effect rule. The AIE effect permits the use of highly concentrated solutions of fluorogens and their aggregates in aqueous media for sensing and imaging applications, leading to the development of fluorescence turn-on or light-up nanoprobes with a wide molecular diversity, ready structural tenability, and good biological compatibility.15 The discovery of AIE has made it possible to track trace substances in small organisms through fluorescence quantification and visualization.16–18
AIEgen fluorescent probes have been developed19,20 using a novel methodology based on an AIEgen for detection and quantification of Hg2+ bioaccumulation in algae and Hg2+ release from algae after bioaccumulation.21 Such applications can be further explored to detect Hg2+ dynamics in other organisms in aquatic ecosystems. In this study, we used a specially designed AIEgen to quantitatively evaluate the bioaccumulation of Hg2+ within a rotifer, and for the first time used this AIEgen to visualize Hg2+ kinetics in vivo within the rotifer in salt water, to better understand the bioaccumulation process using a novel and simple method. This study provides further understanding of the mechanism of interaction between heavy metal ions and small organisms in aquatic systems.
Materials and methods
Materials
All reagents were obtained from Sigma-Aldrich (Australia) unless otherwise specified. Stock solutions of AIEgen (m-TPE-RNS) and HgCl2 at a concentration of 1.0 mM were prepared and stored as previously reported.21The rotifer Brachionus plicatilis was obtained from College of Science and Engineering, Flinders University, with a mean length of 80–180 μm. Microalgae Nannochloropsis oculata and baker's yeast were used to feed the rotifers at a density >400 ind ml−1. Before the test, only yeast was used to minimize the effects of algal pigment on the AIEgen's fluorescence. Stock suspensions of yeast were made by vortexing 3 g dry yeast in 60 ml distilled water, stored at 4 °C, and fed at 0.1 g per million rotifers. Nannochloropsis oculata were cultured according to a reported method.21
The AIE test solution for staining rotifers contained 10 μM m-TPE-RNS. The test solutions for Hg2+ determination contained 10 μM m-TPE-RNS and various Hg2+ concentrations in the form of HgCl2. The photoluminescence (PL) intensity was read on a fluorescence spectrometer (Varian, Australia) with the excitation wavelength at 355 nm.
Optimization for staining Hg2+ with AIE in salt water
A salinity ladder (0, 5, 10, 15, 20, 25, 30, and 35 practical salinity unit) was prepared with filtered sea water and distilled water. The salinity was confirmed by a refractometer. After mixing 10 μL of the m-TPE-RNS stock solution with 390 μL acetonitrile (ACN) and 590 μL water at different salinities, 10 μL of Hg2+ stock solution was added. The PL intensity was measured at 1, 5, 15, 30, and 45 min on the fluorescence spectrometer to determine the peak absorption value over time. The effect of salinity on the PL intensity in AIE and Hg2+ reaction is shown in Fig. S1† and the effect of elapsed time on the PL intensity in AIE and Hg2+ reaction is shown in Fig. S2.†To develop a master curve for Hg2+ determination, a series of Hg2+ concentrations (1, 5, 10, 20, and 30 μM) were detected at a constant AIE concentration (10 μM) over the abovementioned time periods. After the PL intensity was obtained, the PL master curve was developed to show the relationships between PL intensities and Hg2+ concentrations. The Hg2+ concentration could be quantified by the corresponding PL intensity measured using the fluorescence spectrometer.
Quantitative test of Hg2+ bioaccumulation by rotifers
Rotifers were harvested from an aquarium tank with a 50 μm mesh net and rinsed with tap water, and then introduced into a series of water samples containing different HgCl2 concentrations (1, 2.5, 5, and 10 μM) with a rotifer density of 1000 ind mL−1 for 1 h, followed by a period of recovery in clean water for 24 h. Live and dead rotifers were counted from each treated sample using an optical microscope (Leica, USA) and the survival rate was calculated to determine the toxic response of rotifers to Hg2+. The survival rate of rotifers after 1 h of incubation at different concentrations of Hg2+ is shown in Fig. S3† and the typical optical images of live, dead and live rotifers with eggs are shown in Fig. S4.†For bioaccumulation of Hg2+ in rotifers, B. plicatilis were harvested and washed using the method above, and were adjusted to a density of 600 ind mL−1 with clean water, to which HgCl2 was added to reach a final concentration of 5 μM. The rate of Hg2+ absorption by algae was measured at 5, 10, 15, 20, 25, 30, and 45 min. Before each measurement of Hg2+ concentration in the solution, the rotifers were removed using the mesh net. Briefly, the Hg2+ concentration in 600 μL of supernatant was determined at 400 μL AIE in the ACN solution to reach a water![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ACN ratio of 3
ACN ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. AIEgens formed nanoparticles with a mean size of 159 nm, as shown in Fig. S5.† The PL intensity of the stained solution was read on the fluorescence spectrometer.
2. AIEgens formed nanoparticles with a mean size of 159 nm, as shown in Fig. S5.† The PL intensity of the stained solution was read on the fluorescence spectrometer.
To determine the Hg2+ absorption by rotifers, a series of rotifer densities (120, 240, 360, 480, and 500 ind mL−1) were prepared by diluting the stock rotifer culture with clean water. HgCl2 was added to each sample to reach a final Hg2+ concentration of 5 μM. After incubation for 20 min, the rotifers were removed using a mesh net. The Hg2+ remaining in the solution was determined by the AIE method with an AIE concentration of 5 μM. Then, the amount of Hg2+ residue in the culture medium was measured by the AIE method using the fluorescence spectrometer and the PL readings were converted to Hg2+ concentrations using the developed master curve.
In vivo visualization of the dynamics of Hg2+ within rotifers by AIEgens
Rotifers were harvested as per the method above, and were adjusted to a density of 1000 ind mL−1 with clean water and incubated with HgCl2 at a concentration of 5 μM for 20 min. They were then transferred into clean water after 20, 60, 180, and 240 min, and stained using the m-TPE-RNS working solution at a final concentration of 1 μM. An AIE working solution was prepared by diluting the AIE stock solution in ACN and mixing it with water (v/v, 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3) to a concentration of 50 μM. After rinsing twice, the rotifers were subjected to imaging using a Leica TCS SP5 scanning confocal microscope. For the fluorescence imaging, the wavelength of blue and red channels was set at 460–500 nm, and 570–610 nm, respectively, but the excitation wavelength was 405 nm.
3) to a concentration of 50 μM. After rinsing twice, the rotifers were subjected to imaging using a Leica TCS SP5 scanning confocal microscope. For the fluorescence imaging, the wavelength of blue and red channels was set at 460–500 nm, and 570–610 nm, respectively, but the excitation wavelength was 405 nm.
As a supplement to the fluorescence images, spectral analysis of Hg2+ was conducted on the rotifers. Our optical characterization system consisted of an in-house scanning fluorescence confocal microscope22 using a 100× objective lens (NA = 0.9) with low background fluorescence for excitation and signal collection, and diffraction-limited spatial resolution (∼300 nm). A 532 nm diode laser was used for excitation of the samples with an excitation power of 1 mW. The illuminating position of the laser beam on the samples was controlled by a scanner, and the maximum image area of the samples was 200 μm × 200 μm with the scan step size down to 300 nm. The fluorescence signal was collected through a 532 nm notch filter and then a long-pass filter with two short cut-off wavelengths (540 nm and 630 nm), and coupled to a multimode fiber for spectral analysis using a commercial spectrometer. Thus, the background fluorescence from the tissue with its high percentage located in the short wavelength range (<630 nm) was removed. The schematic drawing of the scanning confocal microscope configuration is shown in Fig. S6.†
Results and discussion
Physiology of the rotifer and mechanisms of AIE for Hg2+ sensing
In the present study, the rotifer Brachionus plicatilis was used to characterize Hg2+ absorption in small zooplankton that are widely distributed in brackish water; rotifers occasionally numerically dominate zooplankton communities due to their high reproductive rates.23 The rotifer B. plicatilis is an efficient grazer of microalgae and bacteria, linking the primary producers with secondary consumers in an ecosystem, and is one of the most studied rotifer species in ecotoxicology and aquaculture.19 Like other rotifer species, B. plicatilis reproduces by cyclical parthenogenesis, a lifecycle asexual reproduction without males. Typically, the population of female rotifers grows parthenogenetically (asexual reproduction), whereby amictic (asexual) females produce mitotic diploid eggs. These eggs hatch into genetically identical amictic females. Despite their minute size in the range of 40–200 μm, female rotifers are anatomically complex (Fig. 1). | ||
| Fig. 1 Structure of the rotifer Brachionus plicatilis.24 | ||
The detection mechanism of a specific AIEgen for Hg2+ has been reported.19 For application in chemical and biological sensing, a highly specific reaction of thiosemicarbazides to form 1,3,4-oxadiazoles triggered by Hg2+ is adopted for recognition of Hg2+. In the absence of Hg2+, the sensors are hydrophobic and tend to form aggregates in water. Due to the non-missive spirolactam form of rhodamine, only the blue emission of the tetraphenylethene (TPE) aggregate is expected. After treatment with Hg2+, positively charged rhodamine fluorophores are generated, and their solubility in water is greatly increased. In consequence, TPE emission cannot be observed due to the process of dark through-bond energy transfer (DTBET) and non-radiative decay, whereas emission of rhodamine is intensified. Thus, ratiometric Hg2+ detection could be realized by this rational design strategy (Fig. 2).
 | ||
| Fig. 2 Schematic illustration of the ratiometric Hg2+ sensing mechanism of TPE–RNS.19 | ||
Quantitative evaluation of bioaccumulation of Hg2+ by the rotifer
Numerous studies have shown that Hg is highly toxic. Concentrations at or lower than 0.005 mg L−1 (0.018 μM) cause reduction in both survival and reproduction of various species of Brachionus including B. rubens25 and B. patulus.26 However, the severity of heavy metal toxicity is influenced by various factors such as temperature and food concentration.Bioaccumulation of Hg2+ within rotifers can be quantitatively evaluated by the AIEgen method.21 Several interesting observations were obtained during the evaluation of rotifer bioaccumulation. Rotifers could quickly absorb Hg2+ and the remaining Hg2+ ions in the solution after rotifer absorption were quantified using the master curve (Fig. S7†). Fig. 3 shows that a significant drop in Hg2+ concentration in the medium occurred at the very beginning at a Hg2+ concentration of 5 μM, indicating the significant initial uptake of Hg2+ by rotifers. The concentration of Hg2+ levelled off and plateaued after 5 min. Therefore, 20 min was used as the incubation time in the subsequent study.
Rotifer density is another important parameter influencing the process of bioaccumulation. With an increase in rotifer density, the amount of Hg2+ bioaccumulation increased in the rotifer (Fig. S8†). In the solution with 1.355 μg mL−1 HgCl2 (5 μM), the bioaccumulation efficiency of Hg2+ decreased from 5.28 μg mg−1 h−1 at a rotifer density of 0.093 mg mL−1 to 2.61 μg mg−1 h−1 when the rotifer density increased to 0.375 mg mL−1 (Fig. 4).
 | ||
| Fig. 4 Bioaccumulation efficiency of Hg2+ (HgCl2, 5 μM, 1.355 μg mL−1) at different rotifer densities (mg mL−1) within 20 min. | ||
In vivo visualization of Hg2+ dynamics within rotifers by AIEgens
The m-TPE–RNS based AIEgens have been used for Hg2+ detection in living cells.19 The potential Hg2+ sensing performance of m-TPE–RNS in solution motivated us to evaluate its application in Hg2+ imaging in living rotifers. The rotifers were incubated in a 5 μM HgCl2 solution for 1 h. After washing with clean water, they were stained with 1 μM m-TPE–RNS for 1 h and imaged using confocal laser scanning microscopy. A strong emission in the red channel 570–610 nm was observed along the rotifer digestive tract, especially around the esophagus and mastax (a tooth-like structure for chewing). However, a moderate emission intensity in the blue channel 460–500 nm was detected only in rotifers incubated with the AIEgen, in which the majority of fluorescence was also excited from the mastax. Rotifers in the absence of Hg2+ and AIEgen showed weak fluorescence signals in the blue and red channels 570–610 nm. The blue to red color of the merged image indicates a distinct increment of Hg2+ at the intracellular level after Hg2+ incubation (Fig. 5). Rotifers incubated with the AIEgen, or Hg2+ and AIEgen showed fluorescence along the digestive tract, especially around the mastax, suggesting that the AIEgen in the solution could enter the animal body through feeding.The accumulation of Hg2+ in phytoplankton occurs by passive diffusion across the cell membrane.27 In contrast to algae, the manner of Hg2+ transfer into zooplankton is not clear. In the present study, the fluorescence results showed that Hg2+ was visualized around the rotifer body surface (Fig. 6a1 and a2) and within the digestive tract (Fig. 6b). It is reasonable to infer that inorganic Hg2+ can enter the digestive organ of the rotifer by ingestion, and Hg2+ cannot permeate into the rotifer by diffusion through the integument. Presumably, this is due to the particular structure of the rotifer integument, which is composed of a thick external layer and a thin internal layer with dense material.28,29
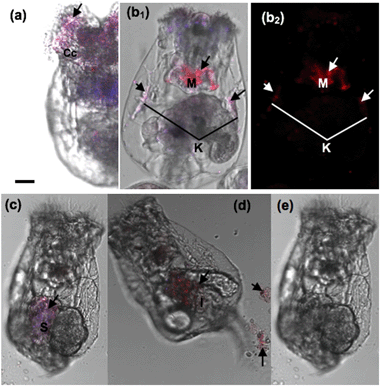
Our results demonstrated that the rotifers incubated with Hg2+ and stained with the AIEgen showed obvious fluorescence along the digestive organ (Fig. 6b). A clearing test was performed to monitor the kinetic transferring of Hg2+ within the rotifer body. Rotifers could ingest Hg2+via their ciliary corona (Fig. 7a). After the rotifers had been transferred into clean water for 20 min, Hg2+ could be viewed in the mastax (Fig. 7b1 and 8b) and flame cells (functional kidney, Fig. 7b2 and 8b). Our result is supported by a previous study on fish where most of the ingested inorganic and organic Hg2+ ions were distributed in the muscle, kidney, gonad, liver and gut.30 However, the distribution of mercury in the body of zooplankton has not been reported. This study was the first to use AIEgen fluorescence as a tool to visualize Hg2+ distribution in rotifer tissues after Hg2+ intake.
The fluorescence in the rotifers decreased over time from 40 min after the rotifers were transferred from the Hg2+ solution into clean water and almost disappeared in 360 min (Fig. 7c–e), indicating that Hg2+ could dissipate from the rotifer body via the processes of defecation and excretion. A scanning fluorescence confocal microscope with a spectral analysis system was used for the validation of Hg2+ distribution in the rotifer body, using the AIEgen for in vivo visualization for the first time, as shown in Fig. 8. The inserts in Fig. 8(c) and (d) (detailed information shown in ESI† Fig. S10) show the PL spectra of two selected points in the fluorescence images of Fig. 8(c) and (d), respectively. The spectrum of the point with red color in Fig. 8(d) shows more than 10 times higher PL intensity at a wavelength of 590 nm than that of the spectrum of the point with blue color in Fig. 8(c), which indicates that the red color in the fluorescence image in Fig. 8 represents the accumulation areas of Hg2+ in the rotifer.
Interestingly, Hg2+ could no longer be observed within the kidneys during this period (Fig. 7c–e, 8c and d). Firstly, the Hg2+ was distributed in the stomach and intestine, but not in the flame cells (kidneys) after 40 min. This phenomenon is similar to a report on fish where the intestinal wall is an effective barrier to HgCl2 but is permeable to MeHg.31 Secondly, besides defecation through the digestive tract, Hg2+ could be excreted through the kidney, as fluorescence (Hg2+) totally disappeared from within the rotifers after 360 min (Fig. 7e). Consequently, we can infer that inorganic mercury is not easily retained in rotifer tissues, though it can quickly enter rotifers via feeding (Fig. 3).
Previous studies have confirmed that in comparison with Hg2+, MeHg is more toxic and more readily accumulated by aquatic biota because of its lipophilic and protein-binding properties.32,33 Methylation of mercury in the aquatic environment is a critical step towards the accumulation of mercury in the aquatic food chain, and MeHg is produced in the environment primarily by anaerobic bacteria34 or by some specific macroalgae.35 According to our fluorescence results, it seems that mercury methylation is unlikely in rotifers, which is one trophic link above algae. Further research is warranted to determine if the direct intake of MeHg would lead to mercury deposition in rotifers.
Conclusions
In this study, we used a specially designed AIEgen to quantitatively evaluate the bioaccumulation of Hg2+ in rotifers, which is the first time that an AIEgen is used to visualize Hg2+ kinetics in vivo within rotifers in salt water to better understand the bioaccumulation process using a simple procedure. Our quantitative results showed that a significant drop in PL intensity occurred at the very beginning after Hg2+ addition, indicating a fast initial uptake of Hg2+ by the rotifers. The amount of bioaccumulation increased with the rotifer density, but the bioaccumulation efficiency of Hg2+ decreased with the rotifer quantity. The distribution of Hg2+ was visualized within the digestive tract, excretory organ and body surface of the rotifers using fluorescence images and spectrum analyses. It seems that inorganic Hg2+ is accumulated in the rotifer digestive organ via food ingestion, and Hg2+ could not diffuse into the rotifer via the body integument. Although inorganic mercury can enter rotifers though food ingestion, it is unlikely that Hg2+ can be methylated and deposited in rotifers. Future study should further investigate the difference between bioaccumulation of inorganic and organic mercury.Conflicts of interest
There are no conflicts to declare.Acknowledgements
Y. Jiang and T. He are grateful for the support of the China Scholarship Council with visiting scholar programs to Flinders University. Y. Tang is grateful for the support of Flinders University through a Faculty of Science and Engineering Establishment Grant.Notes and references
- M. F. Gutierrez, A. M. Gagneten and J. C. Paggi, Ecotoxicology, 2012, 21(1), 37–47 CrossRef CAS PubMed.
- R. A. Lavoie, T. D. Jardine, M. M. Chumchal, K. A. Kidd and L. M. Campbell, Environ. Sci. Technol., 2013, 47(23), 13385–13394 CrossRef CAS PubMed.
- O. Dalman, A. Demirak and A. Balci, Food Chem., 2006, 95(1), 157–162 CrossRef CAS.
- U. Daewel, S. S. Hjollo, M. Huret, R. Ji, M. Maar, S. Niiranen, M. Travers-Trolet, M. A. Peck and K. E. van de Wolfshaar, ICES J. Mar. Sci., 2014, 71(2), 254–271 CrossRef.
- J. S. Gray, Mar. Pollut. Bull., 2002, 45(1–12), 46–52 CrossRef CAS PubMed.
- K. A. Kidd, D. C. G. Muir, M. S. Evans, X. Wang, M. Whittle, H. K. Swanson, T. Johnston and S. Guidford, Sci. Total Environ., 2012, 438, 135–143 CrossRef CAS PubMed.
- L. M. Campbell, R. J. Norstrom, K. A. Hobson, D. C. G. Muir, S. Backus and A. T. Fisk, Sci. Total Environ., 2005, 351, 247–263 CrossRef PubMed.
- C. J. Watras, R. C. Back, S. Halvorsen, R. J. M. Hudson, K. A. Morrison and S. P. Wente, Sci. Total Environ., 1998, 219(2–3), 183–208 CrossRef CAS PubMed.
- F. A. P. C. Gobas, W. de Wolf, L. P. Burkhard, E. Verbruggen and K. Plotzke, Integr. Environ. Assess. Manage., 2009, 5, 624–637 CrossRef CAS PubMed.
- B. C. Kelly, M. G. Ikonomou, J. D. Blair, A. E. Morin and F. A. P. C. Gobas, Science, 2007, 317, 236–239 CrossRef CAS PubMed.
- C. J. Watras, R. C. Back, S. Halvorsen, R. J. M. Hudson, K. A. Morrison and S. P. Wente, Sci. Total Environ., 1998, 219, 183–208 CrossRef CAS PubMed.
- M. T. K. Tsui and W.-X. Wang, Environ. Sci. Technol., 2004, 38(3), 808–816 CrossRef CAS PubMed.
- J. Mei, N. L. C. Leung, R. T. K. Kwok, J. W. Y. Lam and B. Z. Tang, Chem. Rev., 2015, 115(21), 11718–11940 CrossRef CAS PubMed.
- J. Mei, Y. Hong, J. W. Y. Lam, A. Qin, Y. Tang and B. Z. Tang, Adv. Mater., 2014, 26(31), 5429–5479 CrossRef CAS PubMed.
- D. Ding, K. Li, B. Liu and B. Z. Tang, Acc. Chem. Res., 2013, 46(11), 2441–2453 CrossRef CAS PubMed.
- F. Guo, W. P. Gai, Y. Hong, B. Z. Tang, J. Qin and Y. Tang, Chem. Commun., 2015, 51(97), 17257–17260 RSC.
- S. Chen, H. Wang, Y. Hong and B. Z. Tang, Mater. Horiz., 2016, 3, 283–293 RSC.
- Y. Hong, Methods Appl. Fluoresc., 2016, 4, 022003, DOI:10.1088/2050-6120/4/2/022003.
- Y. Chen, W. Zhang, Y. Cai, R. T. K. Kwok, Y. Hu, J. W. Y. Lam, X. Gu, Z. He, Z. Zhao, X. Zheng, B. Chen, C. Gui and B. Z. Tang, Chem. Sci., 2017, 8(3), 2047–2055 RSC.
- Z. Ruan, C. Li, J. Li, J. Qin and Z. Li, Sci. Rep., 2015, 5, 15987, DOI:10.1038/srep15987.
- Y. Jiang, Y. Chen, M. Alrashdi, W. Luo, B. Z. Tang, J. Qin and Y. Tang, RSC Adv., 2016, 6(102), 100318–100325 RSC.
- D. M. Shotton, J. Cell Sci., 1989, 94, 175–206 Search PubMed.
- K. Venetia, C. M. Joséand and D. Pascal, J. Biol. Res., 2012, 17, 97–112 Search PubMed.
- W. Zhao, L. Q. Wang, G. X. Wang, X. Q. Wang, Q. Liu, Z. J. Li, S. L. Zhang and Y. Han, Hydrobiology, China Agriculture Press, China, 2005 Search PubMed.
- S. S. S. Sarma, P. S. L. Jurado and S. Nandini, Rev. Biol. Trop., 2001, 49(1), 77–84 CAS.
- S. S. S. Sarma, H. F. Nunez-Cruz and S. Nandini, Dongwu Xuebao, 2005, 51(1), 46–52 CAS.
- J. Gutknecht, J. Membr. Biol., 1981, 61, 61–66 CrossRef CAS.
- K. Bender and W. Kleinow, Comp. Biochem. Physiol., Part B: Biochem. Mol. Biol., 1988, 89(3), 483–487 CrossRef.
- J. Yu and S. Cui, Hydrobilogia, 1997, 358, 95–103 CrossRef.
- D. Kasper, E. F. A. Palermo, A. C. M. I. Dias, G. L. Ferreira, R. P. Leitao, C. W. C. Branco and O. Malm, Neotrop. Ichthyol., 2009, 7(4), 751–758 CrossRef.
- World Health Organization (WHO), Mercury-environmental aspects, WHO, Switzerland, 1989 Search PubMed.
- D. W. Boening, Chemosphere, 2000, 40(12), 1335–1351 CrossRef CAS PubMed.
- S. M. Ullrich, T. W. Tanton and S. A. Abdrashitova, Crit. Rev. Environ. Sci. Technol., 2001, 31(3), 241–293 CrossRef CAS.
- H. Hsu-Kim, K. H. Kucharzyk, T. Zhang and M. A. Deshusses, Environ. Sci. Technol., 2013, 47(6), 2441–2456 CrossRef CAS PubMed.
- R. Pongratz and K. G. Heumann, Chemosphere, 1998, 36(9), 1935–1946 CrossRef CAS.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c7en00599g |
| ‡ These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2017 |