Guided inquiry in a biochemistry laboratory course improves experimental design ability
Nina M.
Goodey
*a and
Cigdem P.
Talgar
*b
aDepartment of Chemistry and Biochemistry, Montclair State University, 1 Normal Avenue, Montclair, New Jersey 07043, USA. E-mail: goodeyn@mail.montclair.edu; Fax: +1 973 655 7772; Tel: +1 973 666 1368 Tel: +1 973 655 3410
bCenter for Advancing Teaching and Learning through Research, Northeastern University, 360 Huntington Ave., Boston, Massachusetts 02115, USA. E-mail: c.talgar@neu.edu; Fax: +1 617 373 7779; Tel: +1 617 373 2000
First published on 23rd August 2016
Abstract
Many biochemistry laboratory courses expose students to laboratory techniques through pre-determined experiments in which students follow stepwise protocols provided by the instructor. This approach fails to provide students with sufficient opportunities to practice experimental design and critical thinking. Ten inquiry modules were created for a one-semester undergraduate biochemistry laboratory course; these are freely available on the project website. A slightly modified version of the Experimental Design Ability Test (EDAT) was used to assess the impact of inquiry-based learning on student experimental design ability in four experimental (inquiry) and four control (cookbook) sections. EDAT is a published tool that has been validated for use in undergraduate populations. The results, measured by pre- and post-tests, showed a significant positive impact on the experimental design ability of students in sections that employed the inquiry approach, when compared to those in control sections that employed the cookbook approach. A follow-up conversation with students in a sequel course suggested that the inquiry-based approach also benefited students by promoting self-directed learning.
Introduction
Students entering the workforce are perceived by employers to be insufficiently prepared in skills including problem solving and critical thinking (Casner-Lotto and Barrington, 2006; Jaschik, 2015). Many of these skills overlap with science process skills, including the ability to design experiments. Indeed, recommendations of faculty committees on core skills and concepts in biochemistry were summarized in multiple valuable articles (Boyer, 2003; Voet et al., 2003; Tansey et al., 2013; White et al., 2013; Talgar and Goodey, 2015) and emphasize the need to move away from content-driven curricula and increase focus on process skills (Coil et al., 2010). Furthermore, a recent survey of research-active biochemistry professionals in both industry and academia indicated that employers feel students in biochemistry laboratory courses must master process skills related to designing experiments in order to be successful in research positions after graduation (Table 1) (Talgar and Goodey, 2015).| Rank | Skill |
|---|---|
| 1 | A mastery of a limited number of basic math manipulations (solution calculations, dilutions, and serial dilutions). |
| 2 | Understanding the concepts behind the methods, procedures, and assays one works with. |
| 3 | The ability to look up information. |
| 4 | Possessing problem solving skills. |
| 5 | The ability to learn new methods. |
In the authors' opinion, biochemistry laboratory courses are, in many instances, taught in a way that does not provide opportunities for students to practice the above skills. Many research method courses in undergraduate curricula are designed around understanding concepts and thus address skill #2, but may not focus equally on skills #1, 3, 4, and 5 (Handelsman et al., 2004). Biochemistry laboratory courses often employ a cookbook style approach where students perform pre-defined laboratory exercises and are given very detailed instructions on how to accomplish a particular task or experiment. The step-by-step protocols provided in cookbook style courses can deprive students of the opportunities to develop the key skills that these experimental environments have the potential to foster and which employers actively seek. Yet cookbook-style laboratory modules are ubiquitous in biochemistry laboratory courses, perhaps because they are easily assigned to adjunct or graduate student instructors with less experience to guide a student-directed experimental design and because faculty may lack the know how to prepare the necessary materials for inquiry style labs (Cracolice and Monteyne, 2004).
Several studies have indicated that inquiry-based learning in biochemistry increases conceptual understanding and improved student attitudes toward their learning (Basaga et al., 1994; Bailey, 2009; Jensen and Lawson, 2011; Knutson et al., 2011; Goldey et al., 2012; Beck et al., 2014). In such learning environments, research skills are taught through inquiry style and/or project-focused labs, where the process is more generative and students are challenged to integrate and apply their declarative knowledge to develop experimental protocols. By designing experiments, in addition to gaining the conceptual knowledge, students better understand the scientific concepts and the math necessary to set up the design. Furthermore, this approach provides critically important opportunities for students to develop flawed experiments, and then redesign their protocols after receiving feedback from the instructor.
While the number of studies of well-developed inquiry approaches in biochemistry laboratory courses continues to grow (Kirk et al., 2008; Murthy et al., 2014), the assessment of their efficacy must be improved (Ruiz-Primo et al., 2011). In a literature review of metadata on inquiry learning, Beck and coworkers emphasize the importance of using published assessment tools and employing control groups when assessing the impacts of inquiry based learning (Beck et al., 2014).
One goal of the present article is to contribute to the debate concerning the relative merits of inquiry and cookbook learning in laboratory courses (Ault, 2002, 2004; Cracolice and Monteyne, 2004) by providing meaningful data. Such data may encourage departments to consider constructively aligning their biochemistry laboratory courses with the learning goals discussed above by incorporating opportunities for students to gain research skills in addition to conceptual knowledge. To further understand the value of inquiry-based learning environments in teaching experimental design, we fundamentally transformed half of the sections of an introductory Biochemistry laboratory course “Experimental Biochemistry I” to more closely simulate the modern research environment. Specifically, the inquiry sections were designed to emphasize research skills, such as critical and creative thinking, and experimental design. We compared changes in students' experimental design ability in these sections to those of students in sections utilizing more traditional instruction that consisted of pre-designed cookbook experiments in which the experimental protocol was provided by the instructor rather than the students. Three months after the course, we evaluated student responses to questions about their impressions of the class. The evaluations indicated a lasting perceived value of this inquiry-based intervention for students. This work addresses the following questions:
(1) Does replacing cookbook type laboratory experiments by inquiry-based modules in a biochemistry laboratory course improve students' ability to design experiments?
(2) Do students in inquiry-based sections of Experimental Biochemistry I feel more or less prepared for a research/project oriented second semester course, Experimental Biochemistry II?
Based on prior studies, we hypothesized that there would be a larger improvement in students' experimental design ability in the inquiry sections relative to the cookbook sections, and that students in the inquiry sections of Experimental Biochemistry I would be better prepared for a research/project oriented second semester course.
Methods
Participants
Participants for the study were recruited from the first semester offering of the Experimental Biochemistry I course offered at Montclair State University (MSU). Students in Experimental Biochemistry I were juniors and seniors majoring in Biochemistry, Chemistry, Molecular Biology, or Biology. All students in the courses were invited to participate in the study with an in-person-plea. Students (n = 146) enrolled in the first semester offering of the course in two separate first semesters voluntarily participated in the study in return for extra credit points; all students elected to take the assessment, rather than the provided alternative activity, and received extra credit. Students randomly enrolled into the four different sections (2 inquiry and 2 cookbook). Students were not aware of the different teaching approaches in these sections and the inquiry and cookbook sections were randomly assigned only at the beginning of the semester. Therefore, students were not pre-screened based on interest. During the study, we did not know whether the cookbook or inquiry approach would prove to be more effective and did not intentionally disadvantage any group. Approval was obtained from the MSU Internal Review Board for this study (Protocol # 001272). Only the scores from students that had longitudinal data (both pre- and post-EDAT test scores) were included in the analysis; the analyzed sample size was 103 students. Some of the students (39) in the study continued into Experimental Biochemistry II during the second semester.Experimental design
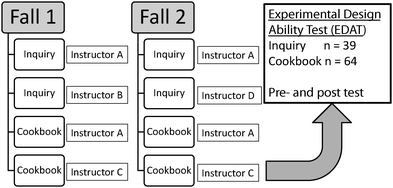
In this 2 (pedagogical intervention) × 2 (pre-post test) mixed methods quasi-experimental study, participants were derived from two different iterations of the Experimental Biochemistry I course (Inquiry vs. Cookbook). A total of 39 participants, for whom we had longitudinal data, were in the inquiry-based courses while 64 participants were in the cookbook version. All participants were administered the EDAT within the first two weeks of the course. Both groups were exposed to the same experiments and biochemistry techniques, just in a different educational format. Two sections each semester used the new, inquiry-based modules and two sections used the cookbook modules. This allowed us to compare results from the control group (cookbook labs) to the group that used the new inquiry modules (Fig. 1) and also compare results between the inquiry modules being implemented by different faculty members. Each semester, there were a total of 3 instructors. One instructor (co-author NG) instructed two sections (one inquiry and one cookbook) which the remaining two sections were instructed by two different instructors. One of these sections was inquiry and one was cookbook.Design of inquiry modules
Prior to the first implementation of the Inquiry modules (Fall 1), experiments used at MSU in Experimental Biochemistry I were a set of predefined exercises where students followed step-by-step instructions on how to conduct an experiment. We created a revised set of laboratory modules that were hypothesis driven and focused on fundamental concepts in biochemistry while maintaining the topic matter of the cookbook experiments (Table 2). The modules follow a logical sequence and address different aspects of the same enzyme, dihydrofolate reductase; a sample module is provided in the appendix (Appendix D500) and the remaining modules can be freely downloaded from the project website (http://www.montclair.edu/csam/nsf-tues-grant/). Some of the modules use materials created in previous modules. For example, students purify the protein dihydrofolate reductase and then do kinetic and inhibition studies using their purified sample in later modules. A laboratory sheet was prepared for each experiment and contained sections shown in Table 3.| Inquiry module title | Module purpose/goal |
|---|---|
| Module 1: checking the calibration of a micropipette | Design and conduct an experiment to determine the precision and accuracy of a micropipette using the gravimetric method. |
| Module 2: determination of unknown NADPH concentration using absorbance spectrophotometry | Design and conduct an experiment to determine the concentration (in mM) in the provided unknown NADPH sample. |
| Module 3: establishing standard curve (Bradford assay) | Design and implement a protocol to establish a standard curve for BSA that shows the relationship between absorbance and protein concentration. |
| Module 4: protein purification by chromatography | Separate the protein Dihydrofolate Reductase (DHFR) from a mixture of many proteins. |
| Module 5: determination of protein purity by SDS-PAGE | Design and conduct an experiment to look at the purity of your DHFR sample and the relationship between amount of protein loaded in a well and the appearance of the resulting band in an SDS-PAGE gel. |
| Module 6: protein concentration determination | Design and implement a protocol to establish the standard curve for BSA and determine the concentration of protein in the samples from Module 4. |
| Module 7: measuring DHFR catalytic activity – effect of enzyme concentration of reaction rate. | Design and implement a protocol to examine the effect of DHFR (enzyme) concentration on the time dependence of the DHFR catalyzed reaction. You will also use the data to determine the catalytic activity of DHFR. |
| Module 8: effect of the DHFR inhibitor Trimethoprim on DHFR catalytic activity | Your assignment is to plan and conduct an experiment to examine the effect of TMP (inhibitor) concentration on the time dependence of the DHFR catalyzed reaction. You will determine an IC50 value for the inhibitor. |
| Module 9: extraction of plasmid DNA, restriction digest, and DNA gel electrophoresis | You will receive a frozen pellet of E. coli cells containing a plasmid. Design and implement a protocol to determine the size of the plasmid and the insert within the plasmid. |
| Module 10: bioinformatics presentations | Design an in silico protocol to (1) determine number of BamHI and NdeI restriction sites in B. stearothermophilus DHFR DNA plasmid and the size of the insert in base pairs; (2) determine the protein sequence of the B. stearothermophilus DHFR; (3) predict the molecular weight, number of amino acids, the pI, and molar extinction coefficient of B. stearothermophilus DHFR; (4) determine the % identity of B. stearothermophilus DHFR with DHFRs human, B. amyloliquefaciens and G. thermodenitrificans. Groups give presentations on their favorite module. |
| Order | Section in laboratory sheet |
|---|---|
| 1 | Introduction to module (in easy to understand terms) |
| 2 | Purpose/goal of the module (see Table 2) |
| 3 | Background on the methods/instruments students might use and references for associated readings |
| 4 | A list of the available tools, chemicals, instrumentation and other supplies |
| 5 | Math moment: a set of related lab math exercises that lead students to the right path with the design |
| 6 | Vocabulary |
| 7 | Advice for experimental design |
| 8 | Common mistakes (list of common pitfalls while designing/doing the experiment) |
| 9 | Safety information |
The modules were formatively revised after each semester they were implemented. Before implementation an undergraduate student worker was employed during the summer before the first and second implementation to test the modules and the modules were revised based on student feedback.
Corresponding cookbook modules
The cookbook modules followed the same sequence of related experiments as the modules in the inquiry set but, unlike the inquiry modules, the cookbook module documents provided a step-by-step protocol instead of the “advice on experimental design” section. Students in the cookbook sections did not have to turn in experimental designs as assignments each week. In place of writing an experimental plan, the cookbook cohorts were expected to study the provided experimental protocol before coming to lab. They submitted all other materials including answers to math moment questions and data sheets and reports, just like the inquiry group. All exams used in the different sections were identical.Implementation of inquiry modules
Students received the inquiry assignment sheets a week before the laboratory session to allow time for groups (2–3 members per group) to design an experiment (or set of experiments) that would answer the question posed or accomplish the stated goal (Table 2). Students also completed math moment questions individually before coming to lab. Students then carried out the experiment/s and recorded their process and analyzed data. Time was built into each module for students to fail, receive instructor feedback, understand limitations of experimental methods, and try again without judgment. During the semester, students prepared five data sheets with figures and calculations (one per module for modules 1–3 and 9–10) individually and two comprehensive laboratory reports spanning modules 4–6 and 7–8 in groups. This distribution of individual and group work forced each student to perform calculations, data analysis, and figure/table preparation individually but also gave students exposure to working on preparing and revising laboratory reports in groups, as they often are likely to do in a work environment. Each group chose one module and presented a 15 minute talk on their approach and findings at the end of the semester. Both experimental and control groups were exposed to the same biochemistry lectures, delivered in traditional style (lecture with occasional group problem solving).Experimental design ability test (EDAT)
We used the EDAT, which measures students' understanding of the criteria for good experimental design through open-ended responses to a prompt grounded in an everyday life science problem (Sirum and Humburg, 2001). EDAT was administered for each section of Experimental Biochemistry I at the beginning and end of the course. There were two different scenario versions of the EDAT test. The administration of the test was counterbalanced such that half of the participants received the “Iron is good for women” version as the pre-test and the “Ginseng” as the post-test and the other half of the participants received the reverse order (See Appendices A and B for EDAT tests and rubric). The study was originally planned for three years employing a control group (2 sections) where the cookbook approach was used and an experimental group (2 sections) where the inquiry approach was used. Once we analyzed the data and found that the inquiry approach resulted in improved student learning, we decided to only run inquiry sections during the third year.The EDAT responses were scored based on a rubric provided by the developers of the EDAT test; the prompts provided to students are available for review in Appendix A. The authors needed to make minor revisions to the rubric to clarify grading and better capture the intricacies of experimental design ability in the biochemistry laboratory. These changes are described in Appendix B and the slightly revised rubric is available for review in Appendix B. Scoring was performed by three independent graders (two faculty members and one graduate assistant). Student responses were provided to the graders as a list of randomized responses, without codes, names, or section information. The graders did not know whether a particular response originated from a student in an inquiry section or a cookbook section and whether the response was written at the beginning or at the end of the semester. All student responses were graded independently by the three different graders and the final scores were an average of these grades. As there is an element of subjectivity in the grading protocol, the graders, to the best of their ability, standardized the grading by discussing the individual grades assigned to a sample set of answers as a group to ensure that they were in agreement on how to apply the rubric. Originally, a grading manual was created, and later on, the original rubric was modified to add brief instructions for graders on some of the grading items to increase consistency (as described in Appendix B).
Follow-up conversations
Following the completion of the second semester laboratory courses, we had four follow-up conversations with students to better understand students' perceptions of their preparation for a classroom-based research experience in an advanced Experimental Biochemistry II course. This advanced biochemistry laboratory course met twice a week for three hours and was only required for biochemistry majors. Student groups (2–4 students) were given different mutants of B. stearothermophilus DHFR and a goal for their semester long project. For example, a group was given the goal to determine whether the mutation impacts the catalytic activity of the enzyme. With instructor support, students designed a sequence of experiments (expression and purification of wildtype and mutant dihydrofolate reductase, protein analysis, and enzyme kinetics) that they conduct in the laboratory. Once they met their first project goal, students came up with a research question and designed more experiments. Students had to do significant problem solving and troubleshooting in the process.During the follow-up conversations, our goal was to gain insight into whether students, who had the inquiry-based experience in Experimental Biochemistry I, felt better prepared for the research experience compared to students who had been in one of the cookbook sections. These groups consisted of 5–8 participants from two sections of the follow-up course Experimental Biochemistry II enrolled during the semester following the Experimental Biochemistry I course. Students were recruited by in-person invitation during class in Experimental Biochemistry II; all students enrolled in Experimental Biochemistry II elected to participate in the conversations. The participants were thus selected based on their continuation in Experimental Biochemistry II. As only biochemistry majors are required to take the second course in the sequence, all participants were biochemistry majors at the time of the follow-up conversation. The participants were grouped as a function of which version of the previous course they took. There were a total of 26 students who participated in the sessions: 11 students from previous inquiry and 16 students from previous cookbook sections. Each session (∼45 minutes) was facilitated by a skilled moderator who recorded student comments. The facilitator did not know which group (inquiry or cookbook) she was working with at each session. The aims for the follow-up conversations were to obtain feedback on how prepared students felt for the advanced biochemistry laboratory course (Experimental Biochemistry II) and whether this depended on whether a student was enrolled in an inquiry or cookbook style course during the previous semester. We did not thoroughly quantitate the numbers of times particular issues were mentioned or conduct in depth thematic analysis; consequently, the findings from the follow-up conversations must be interpreted with caution.
Results and findings
EDAT data
The mean of the EDAT scores for the inquiry groups increased significantly from 3.8 ± 0.40 to 4.9 ± 0.39 while the mean of the EDAT scores for all cookbook students decreased from 5.4 ± 0.25 to 5.0 ± 0.25 (Fig. 2). Mixed Analysis of Variance revealed an interaction between type of condition (inquiry vs. cookbook) and scores pre and post to be significant (F(1,101) = 9.00, p = 0.003, d = 0.45). These data show that the inquiry approach resulted in higher gains in experimental design ability than the cookbook approach in our course and in our setting.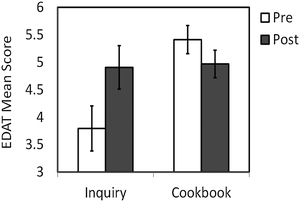 | ||
| Fig. 2 EDAT pre- and posttest mean scores with standard errors are shown for the inquiry and cookbook sections. | ||
The EDAT results were not significantly different between the two Fall semesters when data was obtained. Mixed Analysis of Variance revealed no significant interaction between semester (Fall 1 vs. Fall 2) and pre and post-test EDAT scores. There was also no significant difference in the changes in EDAT scores between the instructors who taught the inquiry courses: analysis of variance showed no significant interaction between instructor and pre and post-test EDAT scores. Similarly there was no significant different in the changes (pre- to post-test) in EDAT scores between different instructors who taught the cookbook sections. Using the rubric associated with the EDAT, students' EDAT responses were independently scored by three raters. The scores were then compared for inter-rater reliability. Each rater was able to score each of the responses that were analyzed and the inter rater reliability was determined to have a Pearson's Correlation Coefficient of 0.84 (p < 0.000). This indicates that the three raters' responses were correlated.
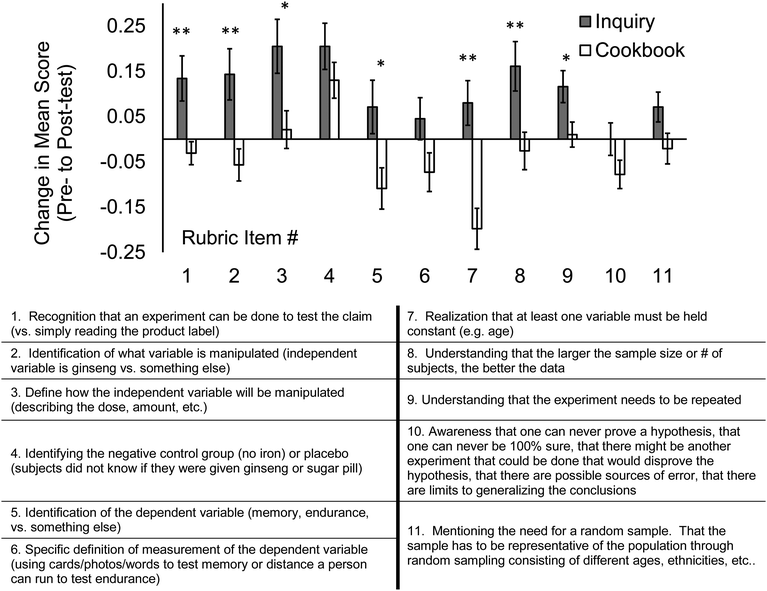
Fig. 3 shows the inquiry- and cookbook section changes in pre- to post-test mean scores for each individual EDAT rubric item with associated standard errors. The mean changes in pre- to post-test scores for items #1, 2, 3, 5, 7, 8, and 9 were significantly different between the inquiry and cookbook groups; mean values, standard errors, t and p values are listed in Table 4. These data suggest that the inquiry approach, compared to the cookbook approach, more effectively prepares students to identify independent and dependent variables (#2 and #5) and understand constants (#7), sample size (#8), and replicates (#9). The absolute inquiry- and cookbook section pre- to post-test mean scores for each individual EDAT rubric item are shown in Appendix C for reference.
| Rubric item # | Mean change pre-to post score ± SE (inquiry sections) | Mean change pre-to post score ± SE (cookbook sections) | t-value | p-value |
|---|---|---|---|---|
| 1 | 0.13 ± 0.050 | −0.03 ± 0.026 | 2.95 | 0.004** |
| 2 | 0.14 ± 0.057 | −0.06 ± 0.036 | 3.00 | 0.003** |
| 3 | 0.21 ± 0.060 | 0.02 ± 0.042 | 2.53 | 0.012* |
| 4 | 0.21 ± 0.051 | 0.13 ± 0.040 | 1.16 | 0.248 |
| 5 | 0.07 ± 0.059 | −0.11 ± 0.046 | 2.41 | 0.017* |
| 6 | 0.05 ± 0.046 | −0.07 ± 0.043 | 1.86 | 0.064 |
| 7 | 0.08 ± 0.049 | −0.20 ± 0.045 | 4.16 | 0.000** |
| 8 | 0.16 ± 0.055 | −0.02 ± 0.041 | 2.73 | 0.007** |
| 9 | 0.12 ± 0.035 | 0.01 ± 0.028 | 2.36 | 0.019* |
| 10 | 0.00 ± 0.036 | −0.08 ± 0.031 | 1.64 | 0.102 |
| 11 | 0.07 ± 0.033 | −0.02 ± 0.034 | 1.81 | 0.071 |
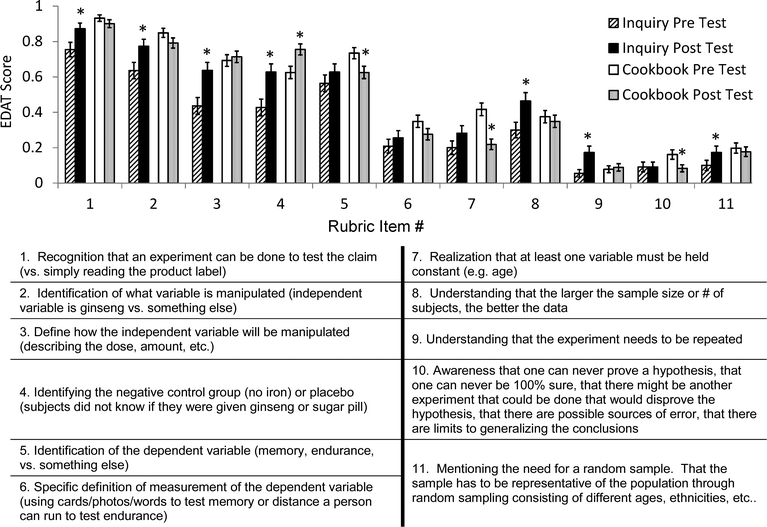
To our surprise, for rubric items #5 and #7, there was also a significant pre- to post-test decrease in the cookbook group's mean scores. For item #5, the mean pre-test score in the cookbook sections (0.73 ± 0.032 (SE)) was significantly higher than the post-test mean score (0.63 ± 0.035); t = 2.39, p = 0.018. Similarly, within the cookbook sections, for items #7 and 10, the mean pre-test scores (0.42 ± 0.036 and 0.16 ± 0.027) were significantly higher compared to the post-test mean scores (0.22 ± 0.030 and 0.083 ± 0.020); t = 4.39 and 2.50, p = 0.000 and 0.013, respectively, for questions #7 and #10 (Appendix C).
Follow-up conversations
Student retrospective reflections are listed in Table 5 and are divided into two columns, comments from students who had had the cookbook vs. inquiry experience. Anecdotally, students who benefited from the inquiry sections felt better prepared for the classroom based research course and emphasized that they benefited from learning from their own mistakes. Students who took the cookbook lab felt less well prepared for the research based course but also expressed that it challenged and motivated them more than the previous semester's course (cookbook experience).| Students from Cookbook sections said | Students from Inquiry sections said |
|---|---|
|
On How Prepared they Feel:
Feel that in terms of the present class (Exp. Biochem. II), they are left on their own “It would be nice to have the goal clearer in our minds, we did not do the planning enough (in Exp. Biochem. II)” It's the first time we are doing this, last semester was more structured Feels like we are “jumping in without our swimmies” Feel insecure if they are doing it right Kept asking “Am I doing this right?” On Technique: Technique wise they feel they are ready Now we have a general idea On Exp. Biochem II Experience: We are going a little further than last semester Feels like what we are doing is more important More emphasis, more interesting, feels like it matters more than going through the procedures like how many drops you can feed on a pen This semester is challenging, you have to decide what procedure to following Before we were told what to do, but now depending on the result you have to decide what to do |
On How Prepared they Feel:
Math modules were very helpful Even general modules were very helpful Very prepared A lot of techniques are repeats we are just now getting to the mastery level (in Exp. Biochem. II) We liked writing our own procedures On Learning from Failure in Exp. Biochem I: We weren't sure what we had to do, and messed up a lot (in Experimental Biochemistry I) Huge learning curve We learned the hard way You do it wrong, you learn from your mistakes If you get everything up front, you don't know what you don't know Last semester we had to write our own protocols |
Discussion
This work is founded on the educational theory, constructive alignment, which is an approach to teaching and learning where a student constructs his or her own knowledge and learning though a set of appropriately designed learning activities. To accomplish this, the teacher creates a learning environment and learning – and assessment activities that support and align with the intended learning outcomes. Constructive alignment suggests that a course's outcomes should align with the activities in which students engage and with assessments of student performance (Biggs, 1996, 2003). In our courses, we want students to learn research skills that will enable their success in modern research positions, including experimental design, laboratory math skills, data analysis, ability to look up information, understanding of the concepts behind methods, troubleshooting, understanding the way a research project progresses, collaboration, and oral and written presentation skills. The present study investigated the impact of embedding inquiry-driven activities, in which the students develop their own protocols, on ability to design experiments, as compared to cookbook models.This is, to our knowledge, the first investigation of the impact of inquiry labs in a biochemistry laboratory course on student learning using both a control group and a validated experimental design ability assessment, although a slightly modified version (see Appendix B for details). The results of the EDAT assessment demonstrate significant advances in students' experimental design thinking in inquiry sections relative to the cookbook condition. These differences suggest that the inquiry-based educational model better, more tightly, aligns the course activities and some of the desired outcomes, specifically the ability to design experiments, than does the cookbook based model.
As discussed above, the data show that students in the cookbook sections, taught in a traditional way, did not make equally large gains as students in the inquiry sections (Fig. 1 and 3). Surprisingly, students in the cookbook groups also performed worse in the EDAT after the course, compared to how they performed before the course. It is possible that weekly exposure to cookbook experiences, where students may not think about experimental design as a part of laboratory experimentation, hinders their ability to perform tasks related to designing experiments such as identifying dependent variables (revised EDAT Rubric #5), understanding the need for keeping experimental variables (other than the independent variable) constant (#7), and considering experimental limitations (#10). This is perhaps analogous to the finding that student exposure to multiple choice only exams (as compared to a mix of multiple choice with constructed-response questions) presents an obstacle for critical thinking in an introductory biology course (Stanger-Hall, 2012). Moreover, the finding that exposure to cookbook-style experiences may decrease students' ability to design experiments parallels a disturbing findings by Barbera and coworkers where they applied the Colorado CLASS-chem instrument that looks at students' versus experts' thinking. The study showed that students, after traditional introductory chemistry courses, tend to think less rather than more like experts compared to before the course (Adams et al., 2008); similar observations were made in an introductory physics course (Adams et al., 2006). These findings, and ours, suggest that aspects of traditional teaching methods can be not only ineffective but also potentially harmful.
Retrospective reflections from the subsequent semester provide some hints for why students in the cookbook sections may have become less apt at experimental design during Experimental Biochemistry I. Students reported that the cookbook version of Experimental Biochemistry I was an unstimulating learning environment compared to the advanced course Experimental Biochemistry II: “Feels like what we are doing is more important (in the advanced course)”, and “More emphasis, more interesting, feels like it matters more than going through the procedures like how many drops can you feed on a pen” (Table 5). We think it is possible that this reported lack of intellectual engagement and the rote following of protocols in the cookbook courses caused students to think less like scientists, damaging their ability or willingness to design their own experiments.
While students self-selected the section they enrolled in, without knowing that there were different teaching approaches, and while the sections were randomly assigned as inquiry or cookbook in the beginning of the semester, one must be cautious when considering whether students were truly randomly assigned into the sections. The mean pre-course EDAT score of students in the cookbook sections was higher than the mean pre-course EDAT score for students in the inquiry sections (Fig. 1); students in the cookbook sections, on average, thus started at a higher level in experimental design ability compared to students in the inquiry groups. Students in the study were from different majors and it is possible that they chose their section based on their course timetable and that there could be a cohort-bias. To the best of our ability, we addressed this by looking at the change in student performance between pre- and post-assessments to account for the possibility that students in one section were more or less prepared compared to another in the beginning of the course.
We sought feedback on the long term benefits of the emphasis on experimental design thinking in the inquiry-based sections three months following the conclusion of the course by way of follow-up conversations conducted with students who had completed those courses and were now enrolled in an advanced experimental biochemistry course. These retrospective reflections suggested that graduates of the inquiry-based course believed they were “very prepared (for the advanced course)” (Table 5). These students were able to identify how the advanced course was giving them an opportunity to further master skills they learned in the first course. These students possessed a clear understanding of the role that each course played in their development “we learned the hard way”, and commented on the “huge learning curve”; they noted that Experimental Biochemistry I provided them a valuable opportunity to become experts in experimental design. On the other hand, those who completed the cookbook version of Experimental Biochemistry I felt under-prepared for Experimental Biochemistry II, students stated that they “felt insecure”, “not sure if they are doing it right”. They discussed how it was the latter course, which instructed them about experimental design for the first time: “It's the first time we are doing this, last semester was more structured”, “Feels like we are “jumping in without our swimmies”, and “…feel that in terms of the present class (Experimental Biochemistry II), we are left on our own” (Table 5). These comments underscore how relatively unprepared the cookbook experience left students to tackle their research projects in Experimental Biochemistry II.
Lastly, students who had taken the course utilizing inquiry-based instruction, in addition to understanding experimental methodology, reported having acquired skills such as learning from failure (self-directed learning) which was number five on the list of skills that both industry and academic professionals felt were central to students' preparation (Talgar and Goodey, 2015). Students from inquiry group in the follow-up conversation reflected on Experimental Biochemistry I as follows: “We weren't sure what we had to do and messed up a lot”, “You do it wrong, you learn from your mistakes”, and “If you get everything up front, you don't know what you don't know”. These comments suggest that students from the inquiry sections were able to spend time in Experimental Biochemistry II improving previously-developed experimental design skills and may thus have benefited more from their research project experiences in the advance course. The ability to practice experimental design, over time, in two settings may have consolidated student understanding; it can be useful to look at inquiry experiments in the context of a curriculum in addition to focusing on individual courses.
The finding that the inquiry approach results in students feeling better prepared for a class-based research experience may be particularly significant for those institutions specifically interested in preparing their undergraduates for research. The modules we developed surrounding a common theme, dihydrofolate reductase enzymatic activity and inhibition are available online at http://www.montclair.edu/csam/nsf-tues-grant/. A sample of cookbook and inquiry modules is shown in Appendix D. We expect instructors at other institutions will adopt these modules for their courses. In the modules, we chose to use a single enzyme to (dihydrofolate reductase) to allow students to see the longitudinal process of a research project. We think that the benefits of using a single enzyme throughout the semester outweighed the disadvantages of being exposed to only one enzyme and one catalytic reaction. As faculty may prefer to use their own favorite enzyme, we are currently preparing a manuscript, which discusses our experience in converting existing cookbook biochemistry labs to inquiry modules. Brickman and coworkers reported higher gains in science literacy skills in inquiry sections compared to traditional sections in a non-majors introductory biology laboratory course but noted that students expressed some resistance to the inquiry approach (Brickman et al., 2009). We agree that balancing student independent activities with instructor support should be considered when engaging in inquiry teaching (Brown et al., 2006) and Vygotsky defines the zone of proximal development to be where students are challenged not too much nor too little (Vygotsky, 1980). Accordingly, we have provided some suggestions to facilitate inquiry-style biochemistry laboratory teaching in Table 6.
| Inquiry lab feature | Possible challenge | Recommendations for instructor |
|---|---|---|
| Students design their own experiments. | Due to lack of experience, students may struggle with experimental design, not know where to start, and feel overwhelmed or discouraged. |
Explain the value of experimental design ability in research positions post-graduation.
Provide support and encouragement; have students do the first experimental design in class. Offer constructive feedback on students’ experimental plans in office hours or on-line before they are due. Direct students to “Advice for experimental design” section in modules for ideas on how to get started. |
| Experiments often fail; students must redesign and repeat experiments. | Students feel worried about “completing the experiment” or running out of time. Students, who are used to failure-proofed, confirmatory experiments, can feel upset when an experiment fails. |
Explain the professional value of learning troubleshooting.
Emphasize the importance of the research process over completing a task. Have students analyze their data in lab directly after performing an experiment and provide ample time to redesign and repeat the experiment during lab; allow students to reflect before offering feedback. Encourage students to view instructor as a coach rather than evaluator. |
| Lab focused on unanswered questions rather than confirming known information. | Students may question what and whether they are learning because they are not used to open-ended inquiry or the teacher not having all the answers. |
Explain that biochemical research deals with unanswered questions and that it is “normal and exciting” to not know all the answers.
Be open about not knowing an answer but explain how one can proceed to find out answers. Listen to student questions with attention. It is helpful for instructor to have research experience. |
| Lab reports are based on individual experimental designs. | Grading can be time consuming due to non-standard lab reports with different experimental methods. |
For some modules, have students prepare datasheets instead of full reports; employ peer-review, combine multiple, related modules into one lab report and have students work on select reports in groups.
Utilize rubrics that are general enough for different reports. |
| Requires understanding of concept behind the lab. | Students do not understand the concept the question is based on and struggle with experimental design. |
Emphasize the importance of studying the concepts before experimental design.
Use on-line videos as pre-lab lectures. Use a book that provides the theory behind different experimental methods (for example, Modern Experimental Biochemistry by Rodney Boyer). |
| Students decide what instrumentation to use | Different students/groups need training on different instruments. |
Provide background information on instruments on the experimental modules.
Prepare videos on different instruments; make these and others available on-line; include safety information. |
| Calculations are needed for experimental design. | Students do not know how to perform particular calculations and struggle with the design. |
Provide sample math problems in module that relate to the experiment to “prime” students for experimental design.
Consider grading select math problems to ensure students practice. |
| Students' experimental designs vary. | Students may encounter different experimental “pitfalls”. |
Provide students a list of common mistakes (pitfalls) in the module.
Discuss possible challenges individually with different groups as they are getting started. |
We hope inquiry learning through our modules improves student gains in other competencies that have an overlap with experimental design ability. We have not yet directly assessed the impact of inquiry labs on the ability to look up information, to problem solve, and to learn new methods. We plan to next examine the effectiveness of the inquiry approach on gains in laboratory math skills, ranked as the most important skill based on a recent survey of biochemistry professionals (Talgar and Goodey, 2015). Research that employs the same inquiry modules at a range of different institutions is still needed to determine whether the findings can be generalized.
The results presented here may encourage skeptical members of the biochemistry teaching community to consider adopting an inquiry style approach to teaching biochemistry laboratory courses, facilitate peer acceptance of faculty members who wish to implement inquiry style learning, or generate discussion on the pros and cons of inquiry based learning in the biochemistry laboratory setting. This study sets a standard for assessments of inquiry approaches in biochemistry laboratory courses and may encourage other researchers to employ appropriate assessments. Our students in the inquiry style laboratory setting are now required to define questions, design experiments, and recognize that there are many paths to the same goal in research and that considerable analysis and decision making must occur in this process.
Appendix A. Prompts and grading rubric for EDAT used in this study
EDAT prompt 1
Advertisements for magnesium supplements claim that it promotes endurance. To determine if the claim is fraudulent and prior to accepting this claim, what type of evidence would you like to see? Provide details of an investigative design.EDAT prompt 2
The claim has been made that women may be able to achieve significant improvements in memory by taking iron supplements. To determine if the claim is fraudulent and prior to accepting this claim, what type of evidence would you like to see? Provide details of an investigative design.Appendix B. Grading rubric for EDAT used in this study
Note: the rubric was slightly modified from what was published previously (Sirum and Humburg, 2001). We added an item (#3: “Define how the independent variable will be manipulated (describing the dose, amount, etc.”)) because the ability to consider how to manipulate the independent variable is an important skill in biochemistry research. We also added another item (#11: “Mentioning the need for a random sample. That the sample has to be representative of the population through random sampling consisting of different ages, ethnicities, etc.…”). We removed original rubric item #7 (“Realization that there are many variables that must be held constant (vs. only one or no mention)”) because it was somewhat repetitive with original rubric item #5 (“Realization that there is one other variable that must be held constant (vs. no mention)”) and caused confusion between graders. To increase the consistency of grading, we changed the wording of number original rubric item #4, which is now #6, to be “Specific definition of measurement of the dependent variable (using cards/photos/words to test memory). Not enough to say conduct a memory test. They must describe some approach to measure memory/ endurance”.Rubric: Student response can receive either 1 or 0 points for each item.
____ 1. Recognition that an experiment can be done to test the claim (vs. simply reading the product label)
____ 2. Identification of what variable is manipulated (independent variable is ginseng vs. something else)
____ 3. Define how the independent variable will be manipulated (describing the dose, amount, etc.)
____ 4. Identifying the negative control group (no iron) or placebo (subjects did not know if they were given ginseng or sugar pill)
____ 5. Identification of the dependent variable (memory, endurance, vs. something else)
____ 6. Specific definition of measurement of the dependent variable (using cards/photos/words to test memory or distance a person can run to test endurance)
____ 7. Realization that at least one variable must be held constant (e.g. age)
____ 8. Understanding that the larger the sample size or # of subjects, the better the data
____ 9. Understanding that the experiment needs to be repeated
____ 10. Awareness that one can never prove a hypothesis, that one can never be 100% sure, that there might be another experiment that could be done that would disprove the hypothesis, that there are possible sources of error, that there are limits to generalizing the conclusions
____ 11. Mentioning the need for a random sample. That the sample has to be representative of the population through random sampling consisting of different ages, ethnicities, etc.
Appendix C
Fig. 4.Appendix D. Sample inquiry and cookbook modules
The remaining modules can be found at the following web address: http://www.montclair.edu/csam/nsf-tues-grant/.Module 7. Measuring DHFR catalytic activity – effect of enzyme concentration of reaction rate. Inquiry version
1. Introduction
Enzymes are biocatalysts that speed up chemical reactions. The catalytic activity of an enzyme is the number of substrate molecules converted to product per second per 1 molecule of enzyme.Note in the above equation that the units for catalytic activity will be s−1. You could of course use a different unit of time, for example minutes. In this case the units of catalytic activity would be min−1.
In this module you will determine the catalytic activity for the enzyme DHFR.
You will also examine how enzyme concentration affects your results. The enzyme is a catalyst that speeds up the process of converting substrate to product. The more enzyme molecules in the reaction vessel, the faster the reaction proceeds. This is analogous to a factory: the more employees working, the more pieces of leather converted to shoes per hour (or per minute, or per second). The more enzymes “working” in your tube, the more molecules of substrate are converted to product per unit time.
To determine the catalytic activity, you will need to know the number of substrate molecules converted to product molecules per unit time (see the equation above). To do this, you will mix the enzyme with substrate and then see how fast the substrate is converted to product.
In a typical experiment, you will measure the change (decrease) in substrate concentration or the increase in product concentration over time. In this experiment you will measure the disappearance of substrate and cofactor by recording the decrease in absorbance at 340 nm over time. In your data file, the Y-axis will be absorbance340nm and the X-axis will be time. Pay attention to the units of the X-axis, be sure to write them down. The slope of the initial linear decrease will represent the change in absorbance over time (ΔAbs/min) and can be used to calculate initial velocity (v0). To convert the slope to Δ[substrate]/time, you will use a special extinction coefficient (see below).
To obtain the catalytic activity values, you can divide Δ[substrate]/time by the concentration of enzyme in the experiment. Just be sure to have the units of enzyme concentration be the same as the units of substrate concentration so that the units cancel out.
Please bring a memory stick where the measured data can be stored! The data can be shared later between all lab members via email or everyone can save the data to their own memory stick.
2. Purpose of the lab
Your lab assignment is to run the reaction (DHF + DHFR + NADPH → THF + DHFR + NADP+) and monitor the disappearance of the substrate (DHF) and cofactor (NADPH). This data will allow you to determine the catalytic activity of the enzyme you purified. You will also experiment with different enzyme (DHFR) concentrations to optimize the experiment. DHFR concentration is the independent variable, all other parameters are kept constant. The dependent variable will be initial velocity.Your assignment is to design and implement a protocol to examine the effect of DHFR (enzyme) concentration on the time dependence of the DHFR catalyzed reaction. You will also use the data to determine the catalytic activity of DHFR.
3. Agenda for the day
• You must have worked through Math Moment problems before coming to class. Upon entry, each student shows their work to instructor to receive points.• Instructor presentation on measuring catalytic activity.
• In class, groups review Math Moment Problems to prepare for experimental design.
• Experimental design: groups discuss and finalize experimental protocol. Each student writes down a step-by-step protocol for the experiment. Note: you must have done most of the experimental design before coming to lab.
• Conduct the experiment in groups. Each student individually records data in their personal laboratory notebook.
• Clean up.
4. Background
• Useful information can be found in Chapter 8B. Please, read this section.• The enzyme that catalyzes the reaction is DHFR (dihydrofolate reductase) (∼18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 Da). DHFR catalyzes the conversion of substrate DHF (dihydrofolate) to product THF (tetrahydrofolate).
600 Da). DHFR catalyzes the conversion of substrate DHF (dihydrofolate) to product THF (tetrahydrofolate).
• NADPH (nicotinamide adenine dinucleotide phosphate) is the cofactor required for the reaction. NADPH donates a hydride (H−) to DHF. NADPH is converted to NADP+ in the reaction. The reaction thus has two starting materials (DHF and NADPH) and two products (THF and NADP+).
• Detecting the reaction: DHF (substrate) and NADPH (cofactor) absorb more light at 340 nm than THF (product) and NADP+. Today, the plate reader will be set so that it takes many absorbance measurements as the reaction proceeds at specific time intervals. This is the “kinetic setting”. The plate reader will only give the time values and the absorbance values, you will use Excel to graph and analyze your data.
• The initial velocity for the DHFR reaction will be determined by measuring the rate of enzyme-dependent decrease in absorbance at 340 nm using the extinction coefficient of 13.2 mM−1 cm−1 and Beer’s law. This means that the values you obtain for the ΔAbs/time (using a 0.5 cm pathlength in the microtiter plate well) will be divided by this extinction coefficient and pathlength (0.5 cm) as stated in Beer’s law. Note what the units are when you do this division when extinction coefficient is expressed in units of mM−1 cm−1 → min−1 (mM−1 cm−1 × 0.5 cm)−1
• The assay will be conducted in a microtiterplate. The pathlength will be 0.5 cm. Different wells will be identical experiments except that each one will have a different enzyme concentration.
• Provide your kcat value in units of s−1. Note that the instrument gives the data in minutes and you will need to do a conversion.
• Concentrations we will use in the assay (final in microtiterplate well) are NADPH (100 μM) and DHF (100 μM). A Mastermix is provided that contains buffer, NADPH, and DHF at appropriate concentrations.
• DHF is a suspension and you must extensively mix Mastermix before each time you add it to another well.
• DHF is light sensitive and must be protected from light as much as possible during the experiment (use aluminum foil for this purpose).
5. Math moment
1. What three molecules will be needed in the microtiterplate well, in addition to the buffer, to start the DHFR catalyzed reaction?2. What are the products of the reaction catalyzed by DHFR?
3. What wavelength must the plate reader be set to in order to monitor the reaction? Do the starting materials DHF and NADPH or the products THF and NADP+ absorb more light at this wavelength?
4. What molecules are present in the beginning of the reaction (time = 0 seconds) in the microtiterplate well?
5. Which molecules are being consumed during the reaction? Which molecules are being made during the reaction?
6. You will be provided a Mastermix solution containing DHF at 118 μM and NAPDH also at 118 μM in buffer. If you mix 170 μL of mastermix with 30 μL of enzyme solution in a microtiterplate well, what is the final concentration of DHF in the well? What is the final concentration of NADPH in the well?
7. Do you expect to see increase or decrease in absorbance at 340 nm as the reaction proceeds? Why?
8. What is the extinction coefficient you will use to convert the enzyme-dependent decrease in absorbance at 340 nm to change in substrate/cofactor concentration? Include units in your answer. See background section.
9. We want to measure kcat, which is equal to Vmax/[E]. Explain why in our experiment the concentration of the substrate and the cofactor must be much greater than then concentration of the enzyme (saturating concentrations). Think about the shape of the Michaelis Menten curve.
10. What will happen to the absorbance vs. time slope if you double the amount of enzyme in the assay?
11. If the initial, linear portion of the line has a slope (Δabsorbance/Δtime × cm) of 0.02 s−1, use the extinction coefficient above to convert this to a slope that gives the Δ[DHF]/Δtime. Remember to include units in your answer. Note: you can use Beer’s law in the following way:
| Δ(absorbance) = ε × 0.5 cm × Δ[DHF] |
note: solve for Δ[DHF]!
12. If the enzyme concentration in the assay in the question above was 30 nM, what is the catalytic activity? Note: you will want to divide the Δ[DHF]/Δtime value by the enzyme concentration (see the equation in the introduction). The units of the [DHF] and [DHFR] will cancel out as long as you make sure that they are in the same units. You can do a unit conversion.
13. Let’s say that you are provided a Mastermix of NADPH and DHF at appropriate concentrations to get the correct final concentrations where enzyme is saturated with NADPH and DHF. You will add 30 μM of enzyme solution to the well for a total concentration of 200 μL. You want to make 8 solutions with different concentrations of enzyme. You want each one to be a 2-fold dilution of the previous one. Please, write into the table how you will make the solutions.
| Name of | Volume of buffer to add (μL) | Total volume (μL) | Final dilution factor | |
|---|---|---|---|---|
| D1 | 100 μL of DHFR stock | 0 | 100 | 1-fold (no dilution) |
| D2 | 50 μL of D1 | 50 | 100 | 2-fold dilution |
| D3 | 100 | |||
| D4 | 100 | |||
| D5 | 100 | |||
| D6 | 100 | |||
| D7 | 100 | |||
| D8 | 100 |
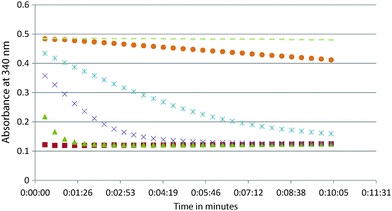
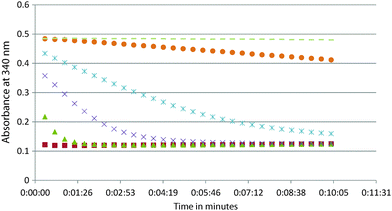
14. See the possible data below. On the graph, label the data 1–6 from the experiment with the HIGHEST enzyme concentration (data 1) to the lowest enzyme concentration (data 6). Remember that the slope of the initial decrease is the initial velocity. Think about what happens to the initial velocity as you increase enzyme concentration. Also, label the 3 sets of data that would be best for determining the kcat. When you analyze your experimental data for this module, do it for all the data sets that have a good initial slope and then take the average. Remember that you will want to measure the initial velocity (the initial decrease). Once the reaction runs out of substrate, there is no longer a decrease and you see a horizontal line (plateau). Provide your answer in units of s−1 (note that x-axis here is in minutes). You will need to do the same conversion for your experimental data.
15. Use the data shown in filled circles in the figure in the question above to determine the kcat for the experiment in units of s−1. Use the extinction coefficient provided in the Background Section. Assume that the enzyme concentration in this experiment was 2 nM.
16. Below in the graph, draw a reaction curve that you would like to see (absorbance vs. time). Now add a curve for what the data would look like if you use way too much enzyme (use thicker line). Think about how, in practice, it will take you a few moments to mix your solutions in a well, then press the button to start measuring, so your time 0 on the graph is actually a little bit after the reaction was initiated. Hint: is it possible that the data looks like a straight line. Where would the line be? Finally, add a line for what the data would look like if you had way too little enzyme (dotted line).
6. Supplies provided
• Ice buckets and ice.• Micropipetters and tips.
• Multichannel pipettor.
• 96-Well microtiter plates.
• Eppendorf tubes.
• Plate shaker.
• UV-Vis plate reader.
• Buffer (40 mM HEPES at pH 6.8).
• Mastermix solution containing DHF and NAPDH both at 118 μM in 40 mM Hepes pH 6.8 buffer.
• Enzyme (DHFR)R solution (your purified enzyme from Module 6). Note: it will be necessary to convert the enzyme concentration into units of μM.
• Aluminum foil.
7. Experimental design
• There are three molecules that must be mixed together, along with buffer, to start the reaction. In practice you will mix 170 μL of Mastermix with 30 μL of enzyme solution in the wells.• You will need to make 8 different dilutions (D1–D8) of the enzyme solution (2× serial dilutions (also known as 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 serial dilutions)). The first solution (D1) will be the undiluted enzyme. For this well, you will simply add 30 μL of the enzyme stock into the well.
2 serial dilutions)). The first solution (D1) will be the undiluted enzyme. For this well, you will simply add 30 μL of the enzyme stock into the well.
• When making your 2× serial dilutions of enzyme (D1–D8), mix each dilution completely before making the next dilution.
• You will need a negative control well where there is no enzyme. In this well, add buffer in place of enzyme.
• In your experiment, you should have 9 wells (8 different enzyme concentrations and one negative control).
• You do not want your reaction to “go” before you are ready to measure so that you will be able to record the time dependent decrease in absorbance before it is “over”. The reaction will not happen until the enzyme is added to the well. For this reason, add enzyme at the very last possible moment (when everything else is ready to go and the plate is in already in the machine, ready to go in).
• For the reason above, you should think about the order of your additions to the microtiterplate. You should have 9 wells (8 different enzyme concentrations and one negative control). You should first add mastermix to all 9 wells. You should then add buffer to the negative control well (nothing should happen). Only then should you add enzyme to the remaining 8 wells. When adding the different enzyme solutions (D1–D8) into the different wells, you should start with the lowest concentration D8 because the reaction should be slowest and not likely go to completion before you measure. You need to complete adding the enzyme solutions into the different wells as quickly as possible and then read the plate as quickly as possible so that you do not miss the reaction for any of the wells. Be completely prepared before doing this part – you will need to think through the steps and practice them with your group. This will be a challenging part of the experiment and you will succeed only if you are prepared.
• One great option for adding the enzyme quickly is to place the different enzyme solutions D1–D8 and buffer for the control into the wells just below the experimental wells. Then you can use a multichannel pipettor to transfer 30 μL of the appropriate solution to the experimental wells, all at once. This way you will minimize the delay between the beginning of the reaction and starting the measurement. The plate can be already in the plate reader plate holder when you do this. You might want to practice this step at your bench with water before you actually do it.
• This is a delicate reaction. Think what is happening at every step and perform your very best to be successful with this experiment.
• Remember to keep all reagents in the ice. Cover the ice bucket with foil because the substrate is sensitive to light. Keep the plate covered with a piece of foil as much as possible once the mastermix is in the well.
8. Common mistakes and some advice
• Remember good pipetting technique. Use the correct pipettor (P20, P200, or P1000) for the volume you measuring. Please, handle pipettors gently.• In solution, DHF is a suspension. If the suspension sits still for a while the solid and liquid phases separate. Therefore, before you use the mastermix, mix it by pipetting it up and down.
• DHF is sensitive to light, keep mastermix covered with foil when not using it.
• Make sure to label tubes and record the wells.
• Be careful with pipetting. If too forcefully pipetted, reagents may splash into other wells.
• Think in which order to fill the wells.
• Be sure to use a clean tip when necessary. Do not contaminate your samples or solutions.
9. Vocabulary
Catalytic activity, substrate, cofactor, reaction rate, kcat.10. Safety
You must wear safety glasses when conducting the experiment. You must never eat or drink in the laboratory. You will need to wait in line to use the plate reader. Please, be patient when waiting for your turn to use the plate shaker and plate reader. Be gentle with the plate reader and plate shaker, these are delicate instruments. Any observed violations of these rules will result in lower final grade and/or removal from the lab. These safety items are solely the responsibility of the student.11. Clean up
For clean-up, return the remaining original solution to the instructor. Discard dilutions in the sink and eppendorf tubes in the regular trash. Do not throw anything in the biohazard waster. Wash 96-well plates and leave them at the sink to dry. Mark the well that you used with a marker. Return pipettors in the correct boxes, the last person puts the boxes in the cabinet in the back of the lab. Dry your ice buckets and place them back in the cabinet. Place all other items where you got them from. Make sure they are clean. Leave your bench ready for the next class to start working.12. Homework
Data from modules 7 and 8 will combined into a formal lab report (one per group). When analyzing data, you should be able to use a few of your slopes (a few wells) to calculate kcat. The ones that finished too quickly or too slowly will not be useful. Remember to use the initial slope (initial linear decrease) to obtain your slope (Δabsorbance/Δtime). Then convert it to Δ[DHF]/Δtime. Finally, divide by [E] to determine kcat. Make sure to convert your kcat to units of s−1.Module 7. Measuring DHFR catalytic activity – effect of enzyme concentration of reaction rate. Cookbook version
13. Introduction
Enzymes are biocatalysts that speed up chemical reactions. The catalytic activity of an enzyme is the number of substrate molecules converted to product per second per 1 molecule of enzyme.Note in the above equation that the units for catalytic activity will be s−1. You could of course use a different unit of time, for example minutes. In this case the units of catalytic activity would be min−1.
In this module you will determine the catalytic activity for the enzyme DHFR.
You will also examine how enzyme concentration affects your results. The enzyme is a catalyst that speeds up the process of converting substrate to product. The more enzyme molecules in the reaction vessel, the faster the reaction proceeds. This is analogous to a factory: the more employees working, the more pieces of leather converted to shoes per hour (or per minute, or per second). The more enzymes “working” in your tube, the more molecules of substrate are converted to product per unit time.
To determine the catalytic activity, you will need to know the number of substrate molecules converted to product molecules per unit time (see the equation above). To do this, you will mix the enzyme with substrate and then see how fast the substrate is converted to product.
In a typical experiment, you will measure the change (decrease) in substrate concentration or the increase in product concentration over time. In this experiment you will measure the disappearance of substrate and cofactor by recording the decrease in absorbance at 340 nm over time. In your data file, the Y-axis will be absorbance340nm and the X-axis will be time. Pay attention to the units of the X-axis, be sure to write them down. The slope of the initial linear decrease will represent the change in absorbance over time (ΔAbs/min) and can be used to calculate initial velocity (v0). To convert the slope to Δ[substrate]/time, you will use a special extinction coefficient (see below).
To obtain the catalytic activity values, you can divide Δ[substrate]/time by the concentration of enzyme in the experiment. Just be sure to have the units of enzyme concentration be the same as the units of substrate concentration so that the units cancel out.
Please bring a memory stick where the measured data can be stored! The data can be shared later for each lab member via email or everyone can save the data to their own memory stick.
14. Purpose of the lab
Your lab assignment is to run the reaction (DHF + DHFR + NADPH → THF + DHFR + NADP+) and monitor the disappearance of the substrate (DHF) and cofactor (NADPH). This data will allow you to determine the catalytic activity of the enzyme you purified. You will also do the experiment with different enzyme (DHFR) concentrations to optimize the experiment. DHFR concentration is the variable, all other parameters are kept constant.Your assignment is to examine the effect of DHFR (enzyme) concentration on the time dependence of the DHFR catalyzed reaction. You will also use the data to determine the catalytic activity of DHFR.
15. Agenda for the day
• You must have worked through Math Moment problems before coming to class. Upon entry, each student shows their work to instructor to receive points.• Instructor presentation on measuring catalytic activity.
• In class, groups review Math Moment Problems to prepare for experimental design.
• Conduct the experiment in groups. Each student individually records data in their personal laboratory notebook.
• Clean up.
16. Background
• Useful information can be found in Chapter 8B. Please, read this section.• The enzyme that catalyzes the reaction is DHFR (dihydrofolate reductase) (∼18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 Da). DHFR catalyzes the conversion of substrate DHF (dihydrofolate) to product THF (tetrahydrofolate).
600 Da). DHFR catalyzes the conversion of substrate DHF (dihydrofolate) to product THF (tetrahydrofolate).
• NADPH (nicotinamide adenine dinucleotide phosphate) is the cofactor required for the reaction. NADPH donates a hydride (H−) to DHF. NADPH is converted to NADP+ in the reaction. The reaction thus has two starting materials (DHF and NADPH) and two products (THF and NADP+).
• Detecting the reaction: DHF (substrate) and NADPH (cofactor) absorb more light at 340 nm than THF (product) and NADP+. Today, the plate reader will be set so that it takes many absorbance measurements as the reaction proceeds at specific time intervals. This is the “kinetic setting”. The plate reader will only give the time values and the absorbance values, you will use Excel to graph and analyze your data.
• The initial velocity for the DHFR reaction will be determined by measuring the rate of enzyme-dependent decrease in absorbance at 340 nm using the extinction coefficient of 13.2 mM−1 cm−1 and Beer’s law. This means that the values you obtain for the ΔAbs/time (using a 0.5 cm pathlength in the microtiter plate well) will be divided by this extinction coefficient and pathlength (0.5 cm) as stated in Beer’s law. Note what the units are when you do this division when extinction coefficient is expressed in units of mM−1 cm−1 → min−1 (mM−1 cm−1 × 0.5 cm)−1
• The assay will be conducted in a microtiterplate. The pathlength will be 0.5 cm. Different wells will be identical experiments except that each one will have a different enzyme concentration.
• Provide your kcat value in units of s−1. Note that the instrument give the data in minutes and you will need to do a conversion.
• Concentrations we will use in the assay (final in microtiterplate well) are NADPH (100 μM) and DHF (100 μM). A Mastermix is provided that contains buffer, NADPH, and DHF at appropriate concentrations.
• DHF is a suspension and you must extensively mix Mastermix before each time you add it to another well.
• DHF is light sensitive and must be protected from light as much as possible during the experiment (use aluminum foil for this purpose).
17. Math moment
1. What three molecules will be needed in the microtiterplate well, in addition to the buffer, to start the DHFR catalyzed reaction?2. What are the products of the reaction catalyzed by DHFR?
3. What wavelength must the plate reader be set to in order to monitor the reaction? Do the starting materials DHF and NADPH or the products THF and NADP+ absorb more light at this wavelength?
4. What molecules are present in the beginning of the reaction (time = 0 seconds) in the microtiterplate well?
5. Which molecules are being consumed during the reaction? Which molecules are being made during the reaction?
6. You will be provided a Mastermix solution containing DHF at 118 μM and NAPDH also at 118 μM in buffer. If you mix 170 μL of mastermix with 30 μL of enzyme solution in a microtiterplate well, what is the final concentration of DHF in the well? What is the final concentration of NADPH in the well?
7. Do you expect to see increase or decrease in absorbance at 340 nm as the reaction proceeds? Why?
8. What is the extinction coefficient you will use to convert the enzyme-dependent decrease in absorbance at 340 nm to change in substrate/cofactor concentration? Include units in your answer. See background section.
9. We want to measure kcat, which is equal to Vmax/[E]. Explain why in our experiment the concentration of the substrate and the cofactor must be much greater than then concentration of the enzyme (saturating concentrations). Think about the shape of the Michaelis Menten curve.
10. What will happen to the absorbance vs. time slope if you double the amount of enzyme in the assay?
11. If the initial, linear portion of the line has a slope (Δabsorbance/Δtime × cm) of 0.02 s−1, use the extinction coefficient above to convert this to a slope that gives the Δ[DHF]/Δtime. Remember to include units in your answer. Note: you can use Beer’s law in the following way:
| Δ(absorbance) = ε × 0.5 cm × Δ[DHF] |
note: solve for Δ[DHF]!
12. If the enzyme concentration in the assay in the question above was 30 nM, what is the catalytic activity? Note: you will want to divide the Δ[DHF]/Δtime value by the enzyme concentration (see the equation in the introduction). The units of the [DHF] and [DHFR] will cancel out as long as you make sure that they are in the same units. You can do an unit conversion.
13. Let’s say that you are provided a Mastermix of NADPH and DHF at appropriate concentrations to get the correct final concentrations where enzyme is saturated with NADPH and DHF. You will add 30 μM of enzyme solution to the well for a total concentration of 200 μL. You want to make 8 solutions with different concentrations of enzyme. You want each one to be a 2-fold dilution of the previous one. Please, write into the table how you will make the solutions.
| Name of | Volume of buffer to add (μL) | Total volume (μL) | Final dilution factor | |
|---|---|---|---|---|
| D1 | 100 μL of DHFR stock | 0 | 100 | 1-fold (no dilution) |
| D2 | 50 μL of D1 | 50 | 100 | 2-fold dilution |
| D3 | 100 | |||
| D4 | 100 | |||
| D5 | 100 | |||
| D6 | 100 | |||
| D7 | 100 | |||
| D8 | 100 |
14. See the possible data below. On the graph, label the data 1–6 from the experiment with the HIGHEST enzyme concentration (data 1) to the lowest enzyme concentration (data 6). Remember that the slope of the initial decrease is the initial velocity. Think about what happens to the initial velocity as you increase enzyme concentration. Also, label the 3 sets of data that would be best for determining the kcat. When you analyze you data experimental data for this module, do it for all the data sets that have a good initial slope and then take the average. Remember that you will want to measure the initial velocity (the initial decrease). Once the reaction runs out of substrate, there is no longer decrease and you see a horizontal line (plateau). Provide your answer in units of s−1 (note that x-axis here is in minutes). You will need to do the same conversion for your experimental data.
15. Use the data shown in filled circles in the figure in the question above to determine the kcat for the experiment in units of s−1. Use the extinction coefficient provided in the Background Section. Assume that the enzyme concentration in this experiment was 2 nM.
16. Below in the graph, draw a reaction curve that you would like to see (absorbance vs. time). Now add a curve for what the data would look like if you use way too much enzyme (use thicker line). Think about how, in practice, it will take you a few moments to mix your solutions in a well, then press the button to start measuring, so your time 0 on the graph is actually a little bit after the reaction was initiated. Hint: is it possible that the data looks like a straight line. Where would the line be? Finally, add a line for what the data would look like if you had way too little enzyme (dotted line).
18. Supplies provided
• Ice buckets and ice.• Micropipetters and tips.
• Multichannel pipettor.
• 96-Well microtiter plates.
• Eppendorf tubes.
• Plate shaker.
• UV-Vis plate reader.
• Buffer (40 mM HEPES at pH 6.8).
• Mastermix solution containing DHF and NAPDH both at 118 μM in 40 mM Hepes pH 6.8 buffer.
• Enzyme (DHFR)R solution (the purified enzyme from Module 6). Note: it will be necessary to convert the enzyme concentration into units of μM.
• Aluminum foil.
19. Experimental protocol
1. Label 8 tubes D1–D8.2. Make serial dilutions of the DHFR enzyme stock in small Eppendorf tubes as shown in the table below. Mix each dilution vigorously before making the next dilution. For example, mix D2 vigorously before making D3. Fill in the final concentrations in the table below (they are based on stock concentration of enzyme).
| Dilution | Volume of buffer to add (μL) | Total volume (μL) | Final concentration | |
|---|---|---|---|---|
| D1 | 100 μL of DHFR stock | 0 | 100 | |
| D2 | 50 μL of D1 | 50 | 100 | |
| D3 | 50 μL of D2 | 50 | 100 | |
| D4 | 50 μL of D3 | 50 | 100 | |
| D5 | 50 μL of D4 | 50 | 100 | |
| D6 | 50 μL of D5 | 50 | 100 | |
| D7 | 50 μL of D6 | 50 | 100 | |
| D8 | 50 μL of D7 | 50 | 100 |
3. Mix the MASTERMIX (provided) vigorously. You must mix MASTERMIX each time before you pipet from it. Place 170 μL of MASTERMIX in wells A1–A9 in a microtiter plate. Keep plate covered with foil (DHF is sensitive to light).
4. Add 30 μL of buffer to well A9. This is your “no enzyme” control. Since there is no enzyme in well A9, do you expect to see any change in absorbance over time?
5. Add 50 μL solution D8 to well B8. (Note, not the experiment well A8 but next to it!)
Add 50 μL solution D7 to well B7.
Add 50 μL solution D6 to well B6.
Add 50 μL solution D5 to well B5.
Add 50 μL solution D4 to well B4.
Add 50 μL solution D3 to well B3.
Add 50 μL solution D2 to well B2.
Add 50 μL solution D1 to well B1.
6. You must do the next steps (up to when you hit read on the plate reader) as quickly as you can because the reaction will start when you add enzyme (DHFR). So go to the instrument, make sure all settings are ready to go. Then place plate on the tray. Only then do the following. Use a multichannel pipettor to transfer 30 μL of solutions from wells B1–B8 to wells A1–A8.
7. Read the plate (10 minutes).
8. Look at your data. Be prepared to repeat the experiment if necessary.
20. Common mistakes and some advice
• Remember good pipetting technique. Use the correct pipettor (P20, P200, or P1000) for the volume you measuring. Please, handle pipettors gently.• In solution, DHF is a suspension. If the suspension is left sitting for a while the solid and liquid phases separate. Therefore, before you use the mastermix, mix it by pipetting it up and down.
• DHF is sensitive to light, keep mastermix covered with foil when not using it.
• Make sure to label tubes and record the wells.
• Be careful with pipetting. If too forcefully pipetted, reagents may splash into other wells.
• Think in which order to fill the wells.
21. Vocabulary
Catalytic activity, substrate, cofactor, reaction rate, kcat.22. Safety
You must wear safety glasses when conducting the experiment. You must never eat or drink in the laboratory. You will need to wait in line to use the plate reader. Please, be patient when waiting for your turn to use the plate shaker and plate reader. Be gentle with the plate reader and plate shaker, these are delicate instruments. Any observed violations of these rules will result in lower final grade and/or removal from the lab. These safety items are solely the responsibility of the student.23. Clean up
For clean-up, return the remaining original solution to the instructor. Discard dilutions in the sink and eppendorf tubes in the regular trash. Do not throw anything in the biohazard waster. Wash 96-well plates and leave them at the sink to dry. Mark the well that you used with a marker. Return pipettors in the correct boxes, the last person puts the boxes in the cabinet in the back of the lab. Dry your ice buckets and place them back in the cabinet. Place all other items where you got them from. Make sure they are clean. Leave your bench ready for the next class to start working.24. Homework
Data from modules 7 and 8 will combined into a formal lab report (one per group). When analyzing data, you should be able to use a few of your slopes (a few wells) to calculate kcat. The ones that finished too quickly or too slowly will not be useful. Remember to use the initial slope (initial linear decrease) to obtain your slope (Δabsorbance/Δtime). Then convert it to Δ[DHF]/Δtime. Finally, divide by [E] to determine kcat.Acknowledgements
This work is supported by a grant from the National Science Foundation: NSF-TUES grant (DUE-1245630) titled “Incorporation of Research Skills into the Undergraduate Biochemistry Curriculum to Create Extraordinary Scientists for the Modern Research Environment” to Nina M. Goodey, PI, Cigdem P. Talgar, Co-PI, James Dyer, Co-PI and John Siekierka, Co-PI. We thank Dr Jim Dyer, Dr John Siekierka, and Dr Ueli Gubler for serving as excellent instructors in the Experimental Biochemistry course and their educational insights. We thank students Diane Hagman and Tiina Sivonen for trialing the inquiry modules during summers 2013 and 2014, respectively, and their helpful feedback. We are grateful to Dr Lynn Schneemeyer for her insights for making the inquiry modules successful. We thank Livia Lazzaro for her expert assistance with administering and coding the assessments for this project. We are grateful to Dr Adrian Goodey for a critical reading of this manuscript.References
- Adams W. K., Perkins K. K., Podolefsky N. S., Dubson M., Finkelstein N. D. and Wieman C. E., (2006), New instrument for measuring student beliefs about physics and learning physics: the Colorado Learning Attitudes about Science Survey, Phys. Rev. ST Phys. Educ. Res., 2(1), 010101.
- Adams W. K., Wieman C. E., Perkins K. K. and Barbera J., (2008), Modifying and validating the Colorado Learning Attitudes about Science Survey for use in chemistry, J. Chem. Educ., 85(10), 1435.
- Ault A., (2002), What's Wrong with Cookbooks? J. Chem. Educ., 79, 1177.
- Ault A., (2004), What's Wrong with Cookbooks? J. Chem. Educ., 81, 1569.
- Bailey C. P., (2009), RNase one gene isolation, expression, and affinity purification models research experimental progression and culminates with guided inquiry-based experiments, Biochem. Mol. Biol. Educ., 37, 44–48.
- Basaga H., Geban O. and Tekkaya C., (1994), The Effect of the Inquiry Teaching Method on Biochemistry and Science Process Skill Achievements, Biochem. Educ., 22, 29–32.
- Beck C., Butler A. and da Silva K. B., (2014), Promoting inquiry-based teaching in laboratory courses: are we meeting the grade? CBE Life Sci. Educ., 13, 444–452.
- Biggs J. B., (1996), Enhancing teaching through constructive alignment, High. Educ., 32, 347–364.
- Biggs J. B., (2003), Teaching for quality learning at university, 2nd edn, Buckingham: Open University Press/Society for Research into Higher Education.
- Boyer R., (2003), Concepts and Skills in the Biochemistry/Molecular Biology Lab, Biochem. Mol. Biol. Educ., 31, 102–105.
- Brickman P., Gormally C., Hallar B. and Armstrong N., (2009), Effects of inquiry-based learning on students' science literacy skills and confidence, International Journal for the Scholarship of Teaching and Learning, 3(2), 16.
- Brown P. L., Abell S. K., Demir A. and Schmidt F. J., (2006), College science teachers' views of classroom inquiry, Sci. Educ., 90(5), 784–802.
- Casner-Lotto J. and Barrington L., (2006), Are They Really Ready to Work? Employers' Perspectives on the Basic Knowledge and Applied Skills of New Entrants to the 21st Century US Workforce (ERIC).
- Coil D., Wenderoth M. P., Cunningham M. and Dirks C., (2010), Teaching the Process of Science: Faculty Perceptions and an Effective Methodology, CBE Life Sci. Educ., 9, 524–535.
- Cracolice M. S. and Monteyne K., (2004), What's wrong with cookbooks? A reply to Ault, J. Chem. Educ., 81, 1559.
- Goldey E. S., Abercrombie C. L., Ivy T. M., Kusher D. I., Moeller J. F., Rayner D. A., Smith C. F. and Spivey N. W., (2012), Biological inquiry: a new course and assessment plan in response to the call to transform undergraduate biology, CBE Life Sci. Educ., 11, 353–363.
- Handelsman J., Ebert-May D., Beichner R., Bruns P., Chang A., DeHaan R., Gentile J., Lauffer S., Stewart J., Tilghman S. M., et al., (2004), Education. Scientific teaching, Science, 304, 521–522.
- Jaschik S., (2015), Well-Prepared in Their Own Eyes, Retrieved from Inside Higher Ed website: http://www.insidehighered.com/news 2015/01/20.
- Jensen J. L. and Lawson A., (2011), Effects of collaborative group composition and inquiry instruction on reasoning gains and achievement in undergraduate biology, CBE Life Sci. Educ., 10, 64–73.
- Kirk S. R., Silverstein T. P., Holman K. L. M. and Taylor B. L. H., (2008), Probing changes in the conformation of tRNA(Phe): an integrated biochemistry laboratory course, J. Chem. Educ., 85, 666–673.
- Knutson K., Smith J., Nichols P., Wallert M. A. and Provost J. J., (2011), Bringing the excitement and motivation of research to students; using inquiry and research-based learning in a year-long biochemistry laboratory: part II-research-based laboratory-a semester-long research approach using malate dehydrogenase as a research model, Biochem. Mol. Biol. Educ., 38, 324–329.
- Murthy P. P. N., Thomspon M. and Hungwe K., (2014), Development of a Semester-Long, Inquiry-Based Laboratory Course in Upper-Level Biochemistry and Molecular Biology, J. Chem. Educ., 91, 1909–1917.
- Ruiz-Primo M. A., Briggs D., Iverson H., Talbot R. and Shepard L. A., (2011), Impact of undergraduate science course innovations on learning, Science, 331, 1269–1270.
- Sirum K. and Humburg J., (2001), The experimental design ability test (EDAT), Bioscene, 37, 8–16.
- Stanger-Hall K. F., (2012), Multiple-choice exams: an obstacle for higher-level thinking in introductory science classes, CBE Life Sci. Educ., 11(3), 294–306.
- Talgar C. P. and Goodey N. M., (2015), Views from academia and industry on skills needed for the modern research environment, Biochem. Mol. Biol. Educ., 43, 324–332.
- Tansey J., Teaster B. J., Cox M., Fox K., Knight J., Sears D. and Bell E., (2013), Foundational Concepts and Underlying Theories for Majors in “Biochemistry and Molecular Biology”, Biochem. Mol. Biol. Educ., 41, 289–296.
- Voet J. G., Bell E., Boyer R., Boyle J., O'Leary M. and Zimmerman J. K., (2003), Recommended Curriculum for a Program in Biochemistry and Molecular Biology, Biochem. Mol. Biol. Educ., 31, 161–162.
- Vygotsky L. S., (1980), Mind in society: the development of higher psychological processes, Harvard University Press.
- White H. B., Benore M. A., Sumter T. F., Caldwell B. D. and Bell E., (2013), What Skills Should Students of Undergraduate Biochemistry and Molecular Biology Programs Have Upon Graduation? Biochem. Mol. Biol. Educ., 41, 297–301.
| This journal is © The Royal Society of Chemistry 2016 |