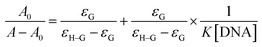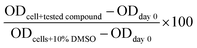DOI:
10.1039/C6RA19469A
(Paper)
RSC Adv., 2016,
6, 110517-110524
Monofunctional platinum(II) dithiocarbamate complexes: synthesis, characterization and anticancer activity†
Received
1st August 2016
, Accepted 8th November 2016
First published on 11th November 2016
Abstract
Heteroleptic platinum(II) dithiocarbamates, of general formula [Pt(DTC)LCl], where DTC = 4-(4-methoxyphenyl)piperazine-1-carbodithioate (1 and 2) and 4-(furan-2-carbonyl)piperazine-1-carbodithioate (3) and L = tri(4-flourophenylphosphine) (1 and 3) and tri(4-chlorophenylphosphine) (2) have been synthesized and characterized by different analytical techniques. These complexes are square planar with picoplatin or phenanthriplatin type steric hindrance from aromatic C–H groups of the phosphine ligand as shown by single-crystal analysis. In 1, the Pt square plane is hindered by two axially oriented hydrogens, whereas by only one in 2 and 3. DNA-binding studies by UV/visible spectroscopy revealed a stronger electrostatic interaction of 1 compared to 2 and 3, and the results are further supported by viscometry and cyclic voltammetric measurements. Their in vitro anticancer activity against five different cancer cell lines using a MTT assay revealed high potency of the complexes. The higher activity of 1 than both 2 and 3 is consistent with DNA binding strength and we speculate that it may be due to the relatively inert nature of platinum towards off-target biomolecules ensured by the hindrance caused by the two axially oriented hydrogens.
Introduction
The well-known anticancer classical platinum drugs, cisplatin, carboplatin and oxaliplatin are bifunctional square-planar platinum(II) complexes differ relatively in the ligand structure (Fig. 1). In oxaliplatin, the presence of N,N-(1,2-diaminocyclohexane) and O,O-(oxalate)chelates1 make it more stable and hence lowers ototoxicity and nephrotoxicity as compared to both cisplatin and carboplatin.2 It has been observed that the CTR1 copper transporter is important in the accumulation of platinum in the cell. Less dependency of oxaliplatin on the CTR1 copper transporter for the cellular uptake imply the seminal role of 1,2-diaminocyclohexane carrier ligand.3 Cisplatin and carboplatin form bifunctional intra-strand cross links in DNA owing to the presence of same carrier ligand (ammine),4 while in contrast, oxaliplatin forms both inter- and intra-strand cross links in the DNA.5 Furthermore, the 1,2-diaminocyclohexane ligand enables oxaliplatin to form bulkier and more hydrophobic oxaliplatin–DNA adducts than both those with cisplatin and carboplatin, such bulkier adducts inhibiting DNA replication more effectively.6 The less labile leaving groups in carboplatin and oxaliplatin also renders then less toxic, although toxicity and resistance remain major hurdles in their clinical use.7–9
 |
| | Fig. 1 Structures of bifunctional and monofunctional platinum(II) complexes planned as anticancer agents. | |
Consequently, new efficient platinum drugs with a mechanism of action dissimilar to that of cisplatin are needed, a scenario that favours the move from bifunctional to monofunctional drugs in order to overcome toxicity, resistance and inactivation of platinum drugs. In this context, monofunctional cationic platinum(II) complexes having a single labile chloride, such as in pyriplatin and phenanthriplatin, have received attention10 owing to their higher activity over classic bifunctional cross-linking agents.11–13 The former give monofunctional adducts with DNA being characterized by a single covalent bond between the platinum center and a nitrogen base of DNA, as manifested in the crystal structure of a dodecamer of site-specifically pyriplatin-platinated duplex DNA.14 Contrary to classical platinum drugs, these complexes do not form DNA cross-links and as a result cause little distortion of the DNA double helix, yet they are still able to block transcription.12 The crystal structure confirmed the perpendicular orientation of the phenanthridine ligand to the platinum coordination plane, thus producing picoplatin-type steric hindrance by partially blocking an open face of the platinum center, thus protecting it from deactivation by off-target biological nucleophiles such as thiols,11,15 glutathione and metallothionein. Furthermore, effective binding of phenanthriplatin to 5′-deoxyguanosine monophosphate indicates that this steric hindrance does not affect the phenanthriplatin reaction with DNA nucleobases. A further advantage is that the hydrophobic phenanthridine is also responsible for greater cellular uptake of phenanthriplatin as compared to cisplatin or pyriplatin.
Motivated by these observations we herein report the synthesis and anticancer activity of three new monofunctional platinum(II) dithiocarbamate complexes with a structural resemblance to phenanthriplatin. These complexes were anticipated to have low reactivity towards off-target sulfur containing biomolecules owing to the strong binding of the chelating dithiocarbamate ligand and also present the possibility of shielding of the platinum centre by aromatic C–H moieties of the phosphine ligand, thus potentially reducing nucleophilic attack by surrounding biomolecules.
Experimental
Materials and methods
Reagents were purchased from; Sigma-Aldrich {1-(4-methoxyphenyl)piperazine and 1-(2-furoyl)piperazine}, Wako Japan {tris(4-flourophenyl)phosphine, tris(4-chlorophenyl)phosphine and platinum(II) chloride}, Riedel-de Haën (CS2) and sodium hydroxide from a local supplier. All the solvents were purchased from Daejung, Sigma-Aldrich and Scharlau. The chemicals used in anticancer study (ethylenediaminetetraacetic acid disodium salt dihydrate (Na2EDTA)), acetic acid, pyruvic acid, L-glutamine, sodium chloride, penicillin-G, sodium sarcosinate, sodium dodecyl sulfate (SDS), Triton X-100, streptomycin sulfate, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) and trizma-base were purchased from Sigma-Aldrich (USA). Fetal bovine serum (FBS) and Dulbecco's Modified Eagle Medium (DMEM) were purchased from GibcoBRL, Gaithersburg, MD. Calf thymus (CT) DNA was procured from Sigma Aldrich. A Gallenkamp (UK) electrothermal melting point apparatus was used for melting points determination. FT-IR spectra were recorded on a Thermo Scientific Nicolet 6700 FT-IR Spectrophotometer (4000–200 cm−1). UV-visible measurements were carried out by UV-1601 UV-visible Spectrophotometer in a pair of quartz cells of 1 cm path length. Elemental analysis was carried out using a CE-440 Elemental Analyzer (Exeter Analytical, Inc.). 300 MHz Bruker instrument was used for obtaining 1H, 13C NMR spectra (TMS as an internal reference) and 31P NMR (85% H3PO4 in water as reference). Chemical shifts are given in ppm and multiplicities of proton signals are presented; s = singlet, d = doublet, t = triplet, dd = doublet of a doublets.
Synthesis of ligands (L1, L2)
Sodium salts of dithiocarbamate ligands were prepared by stirring methanolic solution of the corresponding piperazine {1-(4-methoxyphenyl)piperazine (2 g, 10.40 mmol) (for L1)} and 1-(2-furoyl)piperazine (1.87 g, 10.40 mmol) (for L2)} and sodium hydroxide (0.42 g, 10.40 mmol) for 1 hour followed by the dropwise addition of CS2 (0.79 g, 10.40 mmol) at 0 °C. After two hours stirring, the resultant solution was filtered off and the solvent was rotary evaporated to yield light brown (L1) and light green (L2) products (Scheme 1). Ligands were dried and recrystallized in dry methanol.
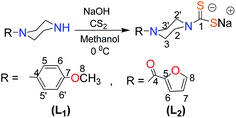 |
| | Scheme 1 Synthetic methodology and numbering scheme for ligands L1 and L2. | |
Ligand (L1). Yield: 2.95 g, 92%. Mp 214 °C. Elemental analysis, % calculated (found) for C12H15N2OS2Na (mol. wt: 290.4 g mol−1): C, 49.63 (49.60); H, 5.21 (5.20); N, 9.65 (9.63); S, 22.08 (22.07). FTIR (4000–400 cm−1): 1510 ν(N-CSS), 1217 ν(C-NC2), 1018 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S), 1001 ν(C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.45 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 2.93 (t, 4H, H3,3′, 3J 1H1H = 5.1 Hz), 3.39 (s, 3H, H8), 6.80–6.92 (m, 4H, benzene, H5,6). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.6 (C1); 50.3 (C2,2′), 51.3 (C3,3′); 145.9 (C4), 118.0 (C5,5′), 114.6 (C6,6′), 153.4 (C7), 55.6 (C8).
S), 1001 ν(C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.45 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 2.93 (t, 4H, H3,3′, 3J 1H1H = 5.1 Hz), 3.39 (s, 3H, H8), 6.80–6.92 (m, 4H, benzene, H5,6). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.6 (C1); 50.3 (C2,2′), 51.3 (C3,3′); 145.9 (C4), 118.0 (C5,5′), 114.6 (C6,6′), 153.4 (C7), 55.6 (C8).
Ligand (L2). Yield: 2.93 g, 95%. Mp 212 °C. Elemental analysis, % calculated (found) for C10H11N2O2S2Na (mol. wt: 278.3 g mol−1): C, 43.15 (43.10); H, 3.98 (3.96); N, 10.06 (10.02); S, 23.04 (23.00). FTIR (4000–400 cm−1): 1645 ν (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1572 ν (N-CSS), 1209 ν (C-NC2), 1015 ν (C
O), 1572 ν (N-CSS), 1209 ν (C-NC2), 1015 ν (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S), 1003 ν (C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.39 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 3.62 (s, br, 4H, H3,3′), 7.02 (d, 1H, H6, 3J 1H1H = 3.6 Hz), 6.62 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.84 (t, 1H, H8, 3J 1H1H = 0.9 Hz). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.9 (C1); 49.2 (C2,2′); 40.8 (C3,3′); 159.0 (C4), 147.4 (C5), 116.1 (C6), 111.8 (C7), 145.2 (C8).
S), 1003 ν (C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.39 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 3.62 (s, br, 4H, H3,3′), 7.02 (d, 1H, H6, 3J 1H1H = 3.6 Hz), 6.62 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.84 (t, 1H, H8, 3J 1H1H = 0.9 Hz). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.9 (C1); 49.2 (C2,2′); 40.8 (C3,3′); 159.0 (C4), 147.4 (C5), 116.1 (C6), 111.8 (C7), 145.2 (C8).
Synthesis of complexes (1–3)
The complexes (1–3) were synthesized by the simultaneous and dropwise addition of the corresponding organophosphine (tris(4-flourophenyl)phosphine) (0.316 g, 1 mmol (1, 3)), (tris(4-chlorophenyl)phosphine) (0.366 g, 1 mmol (2)) in chloroform and sodium salt of corresponding dithiocarbamate (sodium 4-(4-methoxyphenyl)piperazine-1-carbodithioate) (0.290 g, 1 mmol (1, 2)), (sodium 4-(furan-2-carbonyl)piperazine-1-carbodithioate) (0.278 g, 1 mmol (3)) in methanol to the suspension of platinum(II) chloride (0.266 g, 1 mmol) in chloroform and refluxing the reaction mixture for 5 hours (Scheme 2). The resulting solutions were filtered and evaporated on rotary evaporator to get the solid products. The products were dried and recrystallized in chloroform and methanol. Crystals were obtained on slow evaporation.
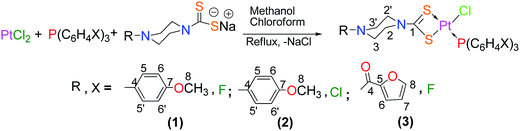 |
| | Scheme 2 Complexes 1–3 synthesis and numbering scheme. | |
Complex 1. Yield: 0.66 g, 82%. Mp 255 °C. Elemental analysis, % calculated (found) for C30H27ClF3N2OPPtS2 (mol. wt: 814.2 g mol−1): C, 44.26 (44.24); H, 3.34 (3.33); N, 3.44 (3.42); S, 7.88 (7.86). FTIR (4000–200 cm−1): 1530 ν(N-CSS), 1225 ν(C-NC2), 1013 ν(C–S), 359 ν(Pt–S), 289 ν(Pt–Cl), 239 ν(Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.91 (t, 4H, H2,2′, 3J 1H1H = 4.8 Hz), 3.14 (t, 4H, H3,3′, 3J 1H1H = 4.8 Hz), 3.42 (s, 3H, H8), 6.81–6.99 (m, 4H, H5,6′), 7.66–7.72 (m, 6H, Hb), 7.35–7.48 (m, 6H, Hc). 13C NMR {(CDCl3, 75 MHz) δ (ppm)}: 201 (C1); 50.5 (C2,2′), 47.6 (C3,3′); 144.3 (C4), 119.5 (C5,5′), 114.5 (C6,6′), 155 (C7), 55.5 (C8), 134.5 (dd, Ca, 4JFC = 2.25 Hz), 136.2–136.6 (m, Cb), 115.6–116.9 (m, Cc), 166.3 (d, Cd, 1JFC = 34.5 Hz). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 13.1.
Complex 2. Yield: 0.69 g, 81%. Mp 250 °C. Elemental analysis, % calculated (found) for C30H27Cl4N2OPPtS2 (mol. wt: 863.5 g mol−1): C, 41.73 (41.71); H, 3.15 (3.12); N, 3.24 (3.21); S, 7.43 (7.41). FTIR (4000–200 cm−1): 1526 ν (N-CSS), 1219 ν (C-NC2), 1011 ν (C–S), 361 ν (Pt–S), 293 ν (Pt–Cl), 239 ν (Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.83 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 3.14 (t, 4H, H3,3′, 3J 1H1H = 5.1 Hz), 3.79 (s, 3H, H8), 6.85–6.92 (m, 4H, H5,6), 7.42–7.47 (m, Hb), 7.57–7.64 (m, Hc). 13C NMR {(CDCl3-75 MHz) δ (ppm)}: 205.4 (C1); 50.5 (C2,2′); 47.0 (C3,3′); 144.3 (C4), 119.5 (C5,5′), 114.7 (C6,6′), 155.0 (C7), 55.6 (C8), 127.0 (d, Ca),135.4 (d, Cb), 128.9 (d, Cc), 137.9 (d, Cd). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 13.0.
Complex 3. Yield: 0.66 g, 82%. Mp 245 °C. Elemental analysis, % calculated (found) for C28H23ClF3N2O2PPtS2 (mol. wt: 802.1 g mol−1): C, 41.93 (41.91); H, 2.89 (2.88); N, 3.49 (3.47); S, 8.00 (7.98). FTIR (4000–200 cm−1): 1636 ν (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1585 ν (N-CSS), 1223 ν (C-NC2), 1009 ν (C–S), 361 ν (Pt–S), 287 ν (Pt–Cl), 266 ν (Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.93 (m, 4H, H2,2′), 3.77 (m, 4H, H3,3′), 7.23 (d, 1H, H6, 3J 1H1H = 3.4 Hz), 6.54 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.50 (d, 1H, H8, 3J 1H1H = 0.9 Hz), 7.64–7.72 (m, Hb,b′), 7.10–7.22 (m, Hc,c′). 13C NMR {(CDCl3-75 MHz) δ (ppm)}: 207.2 (C1); 46.8 (C2,2′), 46.6 (C3,3′); 158.9 (C4), 147.2 (C5), 118.2 (C6), 111.9 (C7), 144.3 (C8), 162.8 (d, Ca, 4JFC = 3 Hz), 136.4 (dd, Cb, 3JFC = 4.5, 8.25 Hz); 116 (dd, Cc, 2JFC = 9, 12.75 Hz), 124.7 (dd, Cd, 1JFC = 3.75, 57.75 Hz). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 9.6.
O), 1585 ν (N-CSS), 1223 ν (C-NC2), 1009 ν (C–S), 361 ν (Pt–S), 287 ν (Pt–Cl), 266 ν (Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.93 (m, 4H, H2,2′), 3.77 (m, 4H, H3,3′), 7.23 (d, 1H, H6, 3J 1H1H = 3.4 Hz), 6.54 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.50 (d, 1H, H8, 3J 1H1H = 0.9 Hz), 7.64–7.72 (m, Hb,b′), 7.10–7.22 (m, Hc,c′). 13C NMR {(CDCl3-75 MHz) δ (ppm)}: 207.2 (C1); 46.8 (C2,2′), 46.6 (C3,3′); 158.9 (C4), 147.2 (C5), 118.2 (C6), 111.9 (C7), 144.3 (C8), 162.8 (d, Ca, 4JFC = 3 Hz), 136.4 (dd, Cb, 3JFC = 4.5, 8.25 Hz); 116 (dd, Cc, 2JFC = 9, 12.75 Hz), 124.7 (dd, Cd, 1JFC = 3.75, 57.75 Hz). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 9.6.
X-ray single crystal analysis
X-ray analysis of 1–3 was carried out on a Bruker Kapa APEX-II CCD diffractometer having a CCD detector set 40.0 mm from crystal. From a sealed ceramic diffraction tube (SIEMENS) having graphite monochromatic Mo-Kα radiation, different intensities were determined. Then by using Patterson and DIRDIF method, the structure of the compound was solved. While further refinement on F2 was carried out by full matrix least square technique using SHELXL-97.16
DNA-binding studies
CT-DNA (20 mg) was dissolved in doubly distilled water to prepare stock solution and concentration was determined spectrophotometrically at 260 nm using an epsilon value of 6600 M−1 cm−1.17 Purity in terms of protein free nature of the DNA was confirmed by A260/A280 that was ∼1.9. The UV absorption titrations were performed by keeping the concentrations of the complexes fixed whereas varying the DNA concentration. In the course of titration the same DNA concentrations were added to both reference and sample cuvettes to eliminate the DNA absorbance itself. After the solutions were mixed for ∼5 min, the absorption spectra were recorded in a range of 200–800 nm.
Benesi–Hildebrand equation was used to determine the association/binding constants of the Pt(II) complexes with DNA.18
where
K is the association/binding constant,
A0 and
A are the absorbances of the compound and its adduct with DNA respectively, and
εG and
εH–G are the absorption coefficients of the compound and the compound–DNA adduct respectively. The Gibb's free energy (Δ
G) was determined from the following equation
ΔG = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K K |
where
R is general gas constant 8.314 J K
−1 mol
−1 and
T is the temperature (298 K).
Viscosity measurements were carried out using an Ubbelohde viscometer at room temperature. A 100 μM solution of DNA and a series of solutions with this constant concentration of DNA and varying concentration of the complexes (50 μM, 100 μM, 150 μM, 200 μM, 250 μM) were made in DMSO. A digital stopwatch was used to measure flow time and an average flow time was calculated from three readings for each solution. First relative viscosity “η” of DNA to solvent was calculated. Then relative viscosities (η) of series of solutions containing constant concentration of DNA and varying concentrations of complexes were measured. The values of relative specific viscosities (η/(η0)) were plotted versus [complex]/[DNA] ratio.19
Cyclic voltammetry was performed for 2 on Biologic SP-300 cyclic voltammeter running with EC-lab Express V 5.40 software japan with three electrodes system i.e. Pt-disc as working electrode, Pt-wire as counter electrode and Ag/AgCl as reference electrode. Analytical grade TBAP was used as supporting electrolyte. Before every reading working electrode was polished with alumina powder and rinsed with distilled water. Nitrogen gas (99.9%) was purged through the working solution to avoid oxygen interference. Drug–DNA binding constant was determined with equation:
log(1/[DNA]) = log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K + log(I/Io − I) K + log(I/Io − I) |
where
K is the binding constant,
Io and
I are the peak currents of free drug and DNA-bound drug respectively. Following form of Randles–Sevcik equation was used for the calculation of diffusion coefficients:
20| Ipa = 2.99 × 105n(αn)1/2ACoDo1/2V1/2 |
where
Ipa is the anodic peak current,
Co is the reductant's concentration in mol cm
−3,
A is the geometric area of the electrode in cm
2,
n is the number of electrons involved in the process and
Do is the diffusion coefficient in cm
2 s
−1.
The cytotoxic potential of complexes 1–3 against five cancer cell lines
The sulforhodamine B (SRB) cellular protein-staining method as described,21 was used to evaluate the cytotoxic potential of the three complexes (1–3) towards proliferation of five different human cancer cell lines. Cancer cells (1 × 104 cells in 190 μL of the complete media) were plated in 96-well plates containing the tested compounds dissolved in 100% dimethylsulfoxide (DMSO) and incubated at 37 °C, 5% CO2 in humidified air for 72 h. The incubation was stopped with tri-chloroacetic acid. The cells were washed, air dried and stained with SRB solution. Optical densities (ODs) were measured at 515 nm using a microplate reader. A zero-day control was performed in each case by adding an equivalent number of cells to several wells, incubating at 37 °C for 30 min and processing as described above. The percentage cell survival was calculated using the following formula:
Results and discussion
Synthesis and characterization
Ligands (L1, L2) and complexes (1–3) were synthesized in good yields by standard methods (see Experimental section). The ligands were highly soluble in methanol, while all complexes were soluble in DMSO, chloroform, acetone and acetonitrile. The presence of Pt–S (361 cm−1) and Pt–P {239 cm−1 (1–2) and 266 cm−1 (3)} stretches which were absent in the precursors indicate the attachment of dithiocarbamate and phosphine respectively,22 absorptions between 1430–1580 cm−1 are also characteristic of dithiocarbamates.23–26 The 1H NMR spectra of 1–3 all show the upfield shift of methylene protons of the piperazine ring at position 2,2′ (3.91, 3.83 and 3.93 ppm) as compared to the free ligands {4.45 (L1) and 4.38 ppm (L2)}.27 In the 13C{1H} NMR spectra a ca. 8 ppm upfield shift of CS2 carbon as compared to the free ligands confirmed the dithiocarbamate attachment to platinum through sulfur atoms,28 while in the 31P{1H} NMR spectrum a singlet (13.1–9.6 ppm) was observed in each case.
Solid-state structures
Crystal data and refinement parameters for the complexes (1–3) are given in Table 1, while the selected bond lengths and bond angles are given in Tables 2 and 3. Each show the expected pseudo square-planar PtS2ClP geometry, the S(1)–Pt(1)–S(2) chelate angles being smaller than ideal value of 90° {74.91(4)° 1, 74.97(14)° 2 and 75.04(6)° 3} which reduces the trans S(1)–Pt(1)–Cl(1) bond angle {170.37(4)° 1, 168.52(15)° 2 and 168.16(6)° 3}. Two sulfurs slightly asymmetrically with one short {2.2766(12) 1; 2.293(4) 2; 2.2923(16) 3} and one long {2.3574(12) 1; 2.343(4) 2; 2.3527(15) 3} Pt–S bond being be attributed to the trans-influence of the phosphine.
Table 1 Crystal data and structure refinement for the complexes (1–3)
| Crystal data |
1 |
2 |
3 |
| Chemical formula |
C30H27ClF3N2OPPtS2 |
C30H27Cl4N2OPPtS2 |
C28H23ClF3N2O2PPtS2 |
| Crystal size (mm3) |
0.350 × 0.220 × 0.160 |
0.400 × 0.230 × 0.160 |
0.38 × 0.20 × 0.15 |
| Formula weight (g mol−1) |
814.16 |
863.51 |
802.11 |
| Crystal system |
Monoclinic |
Monoclinic |
Orthorhombic |
| Space group |
P21/c |
P21/c |
Pbca |
| Temperature (K) |
296(2) |
296(2) |
296(2) |
| Radiation |
MoKα (λ = 0.71073) |
MoKα (λ = 0.71073) |
MoKα (λ = 0.71073) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
| Cell parameters |
| a (Å) |
19.2669(11) |
20.5863(15) |
9.4060(4) |
| b (Å) |
9.8920(6) |
9.3450(7) |
17.1884(10) |
| c (Å) |
16.4584(9) |
18.4610(13) |
36.1133(19) |
| α (°) |
90 |
90 |
90 |
| β (°) |
96.837(3) |
114.952(3) |
90 |
| γ (°) |
90 |
90 |
90 |
| Volume (Å3) |
3114.5(3) |
3220.0(4) |
5838.6(5) |
| Z |
4 |
4 |
8 |
| μ (mm−1) |
4.821 |
4.897 |
5.145 |
| F(000) |
1592 |
1688 |
3120 |
| 2θ range for data collection |
4.636 to 56.034 |
4.414 to 55.202 |
4.74 to 55.046 |
| Reflections collected |
27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 678 678 |
26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 012 012 |
27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 915 915 |
| Independent reflections |
7449 |
7384 |
6716 |
| Goodness-of-fit on F2 |
0.976 |
1.130 |
1.110 |
| Final R indexes [I ≥ 2σ(I)] |
R1 = 0.0386, wR2 = 0.0663 |
R1 = 0.0815, wR2 = 0.1982 |
R1 = 0.0458, wR2 = 0.0770 |
| Final R indexes [all data] |
R1 = 0.0752, wR2 = 0.0760 |
R1 = 0.1195, wR2 = 0.2127 |
R1 = 0.0914, wR2 = 0.0877 |
Table 2 Selected bond lengths of complexes (1–3)
| Type of bond |
Bond length (Å) |
| 1 |
2 |
3 |
| Pt1–P1 |
2.2361(12) |
2.231(3) |
2.2482(15) |
| Pt1–S1 |
2.2766(12) |
2.293(4) |
2.2923(16) |
| Pt1–S2 |
2.3574(12) |
2.343(4) |
2.3527(15) |
| Pt1–Cl1 |
2.3340(12) |
2.334(4) |
2.3283(16) |
| S1–C1 |
1.718(4) |
1.741(18) |
1.728(6) |
| S2–C1 |
1.717(5) |
1.712(14) |
1.724(6) |
| N1–C1 |
1.306(5) |
1.31(2) |
1.314(7) |
Table 3 Selected bond angles of complexes (1–3)
| Angle type |
Bond angle (°) |
| 1 |
2 |
3 |
| P1–Pt1–S1 |
96.37(4) |
97.98(12) |
97.61(6) |
| P1–Pt1–Cl1 |
93.16(5) |
92.47(15) |
94.03(6) |
| S1–Pt1–Cl1 |
170.37(4) |
168.52(15) |
168.16(6) |
| P1–Pt1–S2 |
170.23(4) |
172.50(13) |
171.00(5) |
| S1–Pt1–S2 |
74.91(4) |
74.97(14) |
75.04(6) |
| Cl1–Pt1–S2 |
95.69(4) |
94.78(17) |
93.56(6) |
It is known that square planar platinum(II) complexes undergo ligand substitution via an associative mechanism the incoming ligand coordinating to an axial position11 and generating a trigonal–bipyramidal transition state. In 1–3, the substituents on the phosphine are oriented such that two of the aromatic rings are placed approximately perpendicular to the coordination plane, while the third lies in the plane. Consequently, one of the aromatic C–H bonds occupies an axial position in the platinum coordination plane (Fig. 2a–c). This potentially causing steric hindrance for an approaching nucleophile, a phenomena observed in picoplatin and phenanthriplatin.11,29 Complex 1 is protected at both axial sites by ArC–H moieties, while in 2 & 3 only one side is shielded. The distance of these C-atoms from Pt on both axial sites varies in 1 (i.e. 3.435 Å and 3.635 Å), distances that are slightly longer than those observed in picoplatin (3.224 Å) and phenanthriplatin (3.220 Å).11 These Pt⋯C distances in 2–3 are longer at 3.616 and 3.670 Å respectively. Thus 1 might expected to be more inert to substitution reactions as a result of the greater steric hindrance.11 There are also a number of interesting intermolecular interactions seen in the crystal structures of 1–3 but since they are not maintained in solution they will not be discussed further here (see ESI† for more detail).
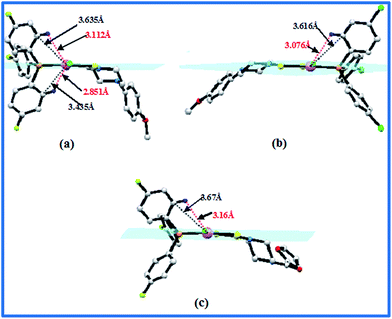 |
| | Fig. 2 Crystal structures of the complexes (1–3) showing steric hindrance from aromatic C–H groups of organophosphine (a) complex 1, (b) complex 2, (c) complex 3, C–Pt distance (black), H–Pt distance (red). | |
DNA binding
UV/visible spectroscopy is a useful technique to inspect the binding constants and binding modes of metal complexes with DNA. The low intensity spectra in Fig. 3 (also S3 and S4†) refer to the free metal complex electronic absorptions i.e. in the absence of DNA. The intense intra-ligand π–π* UV region peaks30 observed in the spectra of 1–3 undergone the obvious hyperchromic changes upon addition of increasing DNA concentration. The hyperchromism suggest a complex DNA interaction mode other than intercalation.31 This kind of spectral behavior can be assigned to electrostatic interaction (or may be covalent binding), a mode accompanied with distortion in the metal coordination core and consequently enhance the probability of intra-ligand π–π* transitions.32 The electrostatic binding mode suggests the generation of a cationic complex in solution via the detachment of Pt-attached chloride. The removal/substitution of halides with solvent molecules i.e. H2O within the cell and then target DNA is also well recognized.33 To further clarify the binding strength of 1–3 with DNA, the association/binding constants were calculated from the intercept-to-slope ratios of A0/(A − A0) vs. 1/[DNA] plots and were found to be 1.81 × 104 M−1 (1), 0.74 × 104 M−1 (2) and 1.12 × 104 M−1 (3). Furthermore, negative values of Gibb's free energy {−24.29 kJ mol−1 (1), −22.07 kJ mol−1 (2) and −23.10 kJ mol−1 (3)} suggest the spontaneous nature of the complex–DNA interaction.
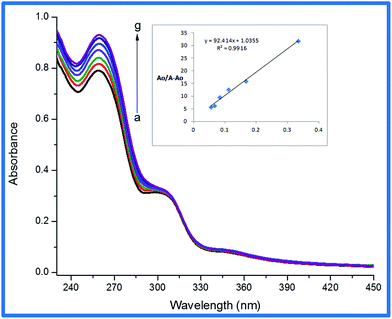 |
| | Fig. 3 Absorbance of 25 μM complex (3) in the absence (a) and presence of (b) 3 μM, (c) 6 μM, (d) 9 μM, (e) 12 μM, (f) 15 μM and (g) 18 μM DNA. The inset graph represents the plot of A0/A − A0 vs. 1/[DNA] (μM)−1 for calculation of binding constant (K) and Gibb's free energy (ΔG). | |
Viscosity measurement is a hydrodynamic method to evaluate drug–DNA interaction and is regarded as the most effective means of distinguishing intercalation and non-intercalation modes in the absence of crystallographic or NMR data.19 A decrease in relative viscosity of CT–DNA with increase in the complex concentration (Fig. 4) demonstrates that the complexes bend/kink the DNA helical structure decreasing its effective length and hence the viscosity.34,35 The decrease in relative viscosity can be rationalized in terms of non-intercalative binding, predominantly administered by the metal center instead of coordinated ligands.36,37 Since the metal complex–DNA binding via electrostatic interaction usually exhibit diminution in viscosity,38 it is a clue favoring the mode as proposed via electronic absorption measurements. In order to further confirm the complex–DNA interaction a representative complex i.e. 2 was also studied via cyclic voltammetric technique. The complex provides couple of well-defined redox peaks at 50 mV s−1 scan rate with an oxidation maximum at −0.67 V and a reduction maximum at −0.80 V, suggesting a quasi-reversible process (Fig. 5). Changes in the oxidation peak current were examined after adding the DNA solution. The decrease in oxidation current from 2.57 × 10−3 to 1.72 × 10−3 mA in the presence of 80 μM DNA signifies complex–DNA interaction. The association/binding constant calculated (0.74 × 104 M−1) from changes in the oxidation peak current by the successive addition of different DNA concentrations (Fig. S5 and S6†) is in agreement with K value obtained from electronic absorption data. The lower diffusion coefficients upon 20 μM DNA addition (5.07 × 10−7) than the free-complex (5.47 × 10−7) reveal the formation of high molecular weight complex–DNA adduct (Fig. 6, S7 and S8†).
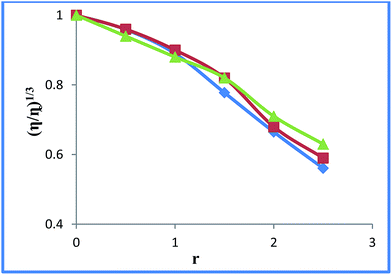 |
| | Fig. 4 Effects of increasing amounts of complexes 1 (blue), 2 (orange) and 3 (gray) on the relative viscosities of CT-DNA at room temperature, [DNA] = 100 μM, r = [complex]/[DNA]. | |
 |
| | Fig. 5 Representative cyclic voltammogram of 1 mM complex 2 in the absence of DNA (red) and in the presence of 80 μM DNA (black) in DMSO with 0.5 M TBAP as supporting electrolyte at 50 mV s−1 scan rate. | |
 |
| | Fig. 6 Representative plot of I/μA vs. (V s−1)1/2 for determination of diffusion coefficients of free drug (2) and 2-DNA adduct. | |
Anticancer activity of 1–3
Complexes 1–3 were examined for their in vitro cytotoxic potential against LU human lung carcinoma {University of Illinois at Chicago (UIC)}, MCF-7 human breast adenocarcinoma, MDA-MB-231 human breast adenocarcinoma, Hepa-IcIc7 mouse liver hepatoma and PC-3 human prostate adenocarcinoma {American Type Culture Collection (ATCC)} by MTT method using staurosporine as a standard drug. The IC50 values show that all are active against these five cancer cell lines (Table 4). The stable nature of Pt–P and Pt–S bonds (as indicated by single-crystal X-ray analysis) is suggestive of their safe carriage to the target DNA by lowering down the affinity of Pt center towards off-target biomolecules.11,15 On the basis of the complex–DNA study and relevant literature33 it can be assumed that these complexes remain intact in the outer cellular environment but dissociate within the cells due to change in Cl− ion concentration (from high to low). Complex 1 showed enhanced activity against four cell lines (LU, MDA-MB-231, Hepa-IcIc7 and PC-3), an observation consistent with higher DNA binding constant value. Steric hindrance imposed by two axially leaning aromatic C–H of the triorganophosphine might also play a role of molecular access to the target. The relatively lower activity of 2 and 3 can be attributed to their low DNA binding affinity and (possibly) single-sided axial hindrance and protection. Furthermore, the higher activity of 1 and 3 may be due to the presence of fluoro-moieties, which not only gives high lipophilicity to the complex to get into the cell39–41 but also have the potential to stabilize complex–DNA adducts through hydrogen bonding. Moreover, IC50 values are much closer to DNA-distorting anticancer drug cisplatin42–44 instead of kinase inhibitor staurosporine, the telltale sign of DNA-distortion being the reason for anticancer action. For the most active complex 1 activity varies against different cell lines in the sequence LU > IcIc7 > PC3 > MDA-231 > MCF7. From this variation, it can be concluded that anticancer action of a complex also depends on the type of cancer.
Table 4 IC50 values (μM) of the complexes (1–3) against five different cancer cell lines
| Complex |
LU |
MCF-7 |
Hepa-IcIc7 |
PC-3 |
MDA-MB-231 |
| (1) |
3.7 ± 1.5 |
14.2 ± 2.0 |
4.8 ± 0.8 |
6.2 ± 1.0 |
8.8 ± 2.9 |
| (2) |
49.27 ± 0.9 |
45.2 ± 1.3 |
48.4 ± 5.3 |
30.1 ± 1.6 |
45.6 ± 3.8 |
| (3) |
20.12 ± 7.4 |
4.9 ± 0.7 |
18.8 ± 3.6 |
21.2 ± 4.3 |
22.3 ± 2.0 |
| Staurosporine |
0.025 |
0.02 |
0.020 |
0.01 |
0.03 |
| Cisplatin |
— |
22.4 (ref. 42) |
— |
38 (ref. 43) |
39 ± 5.0 (ref. 44) |
Conclusions
Stimulated by the promising anticancer activity of phenanthriplatin, three new monofunctional platinum(II) dithiocarbamate complexes 1–3 have been synthesized and characterized using different spectroscopic techniques and X-ray crystallography. Notably, each demonstrates phenanthriplatin-type steric hindrance at a single (2 and 3) or both (1) axial positions resulting from binding of the aromatic C–H moieties of the phosphine ligand. This axial sheltering may lead to resistance to nucleophilic attack at the Pt(II) center, and thus enhance the chances of complex reaching its pharmacological target intact. These complexes are a few times more active than cisplatin against five cancer cell lines, and the activity varies in the sequence 1 > 3 > 2, in accordance with DNA binding strength. The high activity of 1 and 3 can be attributed to the fluoro groups owing to either its ability to penetrate the cell membrane efficiently or to stabilize complex–DNA adducts by hydrogen bonding. The DNA binding further revealed the non-intercalative mode of interaction. Overall, the anticancer activity seems to be governed by the nature of the phosphine and dithiocarbamate substituents and studies are underway to prepare and screen a larger library of such complexes in order to identify complexes with high anticancer activity, better understand the way(s) they get into the cell and also to determine the nature of their interaction with DNA inside the cell.
Acknowledgements
We acknowledge the financial support from TWAS (The World Academy of Sciences, project no. 11-143 RG/PHA/AS_C-UNESCO FR: 3240262658) and Higher education Commission of Pakistan.
References
- M. G. Apps, E. H. Choi and N. J. Wheate, Endocr.-Relat. Cancer, 2015, 22, R219–R233 CAS.
- L. M. Pasetto, M. R. D'Andrea, E. Rossi and S. Monfardini, Crit. Rev. Oncol. Hematol., 2006, 59, 159–168 CrossRef PubMed.
- E. Raymond, S. Faivre, S. Chaney, J. Woynarowski and E. Cvitkovic, Mol. Cancer Ther., 2002, 1, 227–235 CAS.
- R. M. Wing, P. Pjura, H. R. Drew and R. E. Dickerson, EMBO J., 1984, 3, 1201–1206 CAS.
- J. Graham, M. Muhsin and P. Kirkpatrick, Nat. Rev. Drug Discovery, 2004, 3, 11–12 CrossRef CAS PubMed.
- J. L. Misset, H. Bleiberg, W. Sutherland, M. Bekradda and E. Cvitkovic, Crit. Rev. Oncol. Hematol., 2000, 35, 75–93 CrossRef CAS PubMed.
- N. P. Johnson, J.-L. Butour, G. Villani, F. L. Wimmer, M. Defais, V. Pierson and V. Brabec, in Ruthenium and Other Non-Platinum Metal Complexes in Cancer Chemotherapy, Springer, 1989, pp. 1–24 Search PubMed.
- P. J. O'Dwyer, J. P. Stevenson and S. W. Johnson, Cisplatin: chemistry and biochemistry of a leading anticancer drug, 1999, pp. 29–69 Search PubMed.
- P. J. Morin, Drug Resist. Updates, 2003, 6, 169–172 CrossRef CAS PubMed.
-
(a) K. S. Lovejoy, M. Serova, I. Bieche, S. Emami, M. D'Incalci, M. Broggini, E. Erba, C. Gespach, E. Cvitkovic and S. Faivre, Mol. Cancer Ther., 2011, 10, 1709–1719 CrossRef CAS PubMed;
(b) I. A. Riddell, T. C. Johnstone, G. Y. Park and S. J. Lippard, Chem.–Eur. J., 2016, 22, 7574–7581 CrossRef CAS PubMed.
- G. Y. Park, J. J. Wilson, Y. Song and S. J. Lippard, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11987–11992 CrossRef CAS PubMed.
-
(a) T. C. Johnstone, G. Y. Park and S. J. Lippard, Anticancer Res., 2014, 34, 471–476 CAS;
(b) S. X. Chong, S. C. F. Au-Yeung and K. K. W. To, Curr. Med. Chem., 2016, 23, 1268–1285 CrossRef CAS PubMed.
- R. H. Shoemaker, Nat. Rev. Cancer, 2006, 6, 813–823 CrossRef CAS PubMed.
- R. C. Todd and S. J. Lippard, in Platinum and other heavy metal compounds in cancer chemotherapy, Springer, 2009, pp. 67–72 Search PubMed.
- J. Parkinson and P. Sadler, J. Chem. Soc., Dalton Trans., 1998, 3577–3586 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A, 2008, 64, 112–122 CrossRef CAS PubMed.
- M. Reichmann, S. Rice, C. Thomas and P. Doty, J. Am. Chem. Soc., 1954, 76, 3047–3053 CrossRef CAS.
- S. Shujha, A. Shah, N. Muhammad, S. Ali, R. Qureshi, N. Khalid and A. Meetsma, Eur. J. Med. Chem., 2010, 45, 2902–2911 CrossRef CAS PubMed.
- S. Shi, J. Liu, J. Li, K. C. Zheng, X. Huang, C. Tan, L. Chen and L. Ji, J. Inorg. Biochem., 2006, 100, 385–395 CrossRef CAS PubMed.
- R. Holze and B. Bunsen-Ges, J. Phys. Chem., 1994, 98, 1350 Search PubMed.
- M. You, D. M. Wickramaratne, G. L. Silva, H. Chai, T. E. Chagwedera, N. R. Farnsworth, G. A. Cordell, A. D. Kinghorn and J. M. Pezzuto, J. Nat. Prod., 1995, 58, 598–604 CrossRef CAS.
- V. Alverdi, L. Giovagnini, C. Marzano, R. Seraglia, F. Bettio, S. Sitran, R. Graziani and D. Fregona, J. Inorg. Biochem., 2004, 98, 1117–1128 CrossRef CAS PubMed.
- N. Manav, A. Mishra and N. Kaushik, Spectrochim. Acta, Part A, 2004, 60, 3087–3092 CrossRef CAS PubMed.
- N. Manav, A. Mishra and N. Kaushik, Spectrochim. Acta, Part A, 2006, 65, 32–35 CrossRef CAS PubMed.
- G. Hogarth, Prog. Inorg. Chem., 2005, 53, 71–561 CrossRef CAS.
- E. D. Risberg, J. Mink, A. Abbasi, M. Y. Skripkin, L. Hajba, P. Lindqvist-Reis, É. Bencze and M. Sandström, Dalton Trans., 2009, 1328–1338 RSC.
- P. A. Ajibade and O. G. Idemudia, Bioinorg. Chem. Appl., 2013, 2013, 1–8 CrossRef PubMed.
- H. Khan, A. Badshah, M. Said, G. Murtaza, J. Ahmad, B. J. Jean-Claude, M. Todorova and I. S. Butler, Appl. Organomet. Chem., 2013, 27, 387–395 CrossRef CAS.
- Y. Chen, Z. Guo, S. Parsons and P. J. Sadler, Chem.–Eur. J., 1998, 4, 672–676 CrossRef CAS.
- Q. L. Zhang, J. G. Liu, J. Liu, G. Q. Xue, H. Li, J. Z. Liu, H. Zhou, L. H. Qu and L. N. Ji, J. Inorg. Biochem., 2001, 85, 291–296 CrossRef CAS PubMed.
- V. A. Bloomfield, D. M. Crothers and I. Tinoco, Physical Chemistry of Nucleic Acids, Harper & Row, New York, 1974, p. 432 Search PubMed.
- F. Arjmand and M. Aziz, Eur. J. Med. Chem., 2009, 44, 834–844 CrossRef CAS PubMed.
- S. J. Dougan, A. Habtemariam, S. E. McHale, S. Parsons and P. J. Sadler, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 11628–11633 CrossRef CAS PubMed.
- S. Satyanarayana, J. C. Dabrowiak and J. B. Chairs, Biochemistry, 1993, 32, 2573–2584 CrossRef CAS PubMed.
- S. Shi, J. Liu, J. Li, K. C. Zheng, X. M. Huang, C. P. Tan, L. M. Chen and L. N. Ji, J. Inorg. Biochem., 2006, 100, 385–395 CrossRef CAS PubMed.
- W. Villarreal, L. Colina-Vegas, C. R. de Oliveira, J. C. Tenorio, J. Ellena, F. C. Gozzo, M. R. Cominetti, A. G. Ferreira, M. A. B. Ferreira, M. Navarro and A. A. Batista, Inorg. Chem., 2015, 54, 11709–11720 CrossRef CAS PubMed.
- M. M. Aleksi and V. Kapetanovi, Acta Chim. Slov., 2014, 61, 555–573 Search PubMed.
- S. Satyanarayana, J. C. Dabrowiak and J. B. Chaires, Biochemistry, 1992, 31, 9319–9324 CrossRef CAS PubMed.
- S. Sersen, J. Kljun, K. Kryeziu, R. Panchuk, B. Alte, W. Korner, P. Heffeter, W. Berger and I. Turel, J. Med. Chem., 2015, 58, 3984–3996 CrossRef CAS PubMed.
- M. Maschke, H. Alborzinia, M. Lieb, S. Woelfl and N. Metzler-Nolte, ChemMedChem, 2014, 9, 1188–1194 CrossRef CAS PubMed.
- A. K. Renfrew, R. Scopelliti and P. J. Dyson, Inorg. Chem., 2010, 49, 2239–2246 CrossRef CAS PubMed.
- M. Altaf, M. Monim-ul-Mehboob, A. A. Seliman, M. Sohail, M. I. Wazeer, A. A. Isab, L. Li, V. Dhuna, G. Bhatia and K. Dhuna, Eur. J. Med. Chem., 2015, 95, 464–472 CrossRef CAS PubMed.
- R. McPherson, P. Galettis and P. De Souza, Br. J. Cancer, 2009, 100, 649–655 CrossRef CAS PubMed.
- A. J. Pope, C. Bruce, B. Kysela and M. J. Hannon, Dalton Trans., 2010, 39, 2772–2774 RSC.
Footnote |
| † Electronic supplementary information (ESI) available: Single-crystal X-ray diffraction data for the structural analysis have been deposited with the Cambridge Crystallographic Data Center. CCDC 1497036 (1), 1497035 (2) and 1497034 (3). For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/c6ra19469a |
|
| This journal is © The Royal Society of Chemistry 2016 |
Click here to see how this site uses Cookies. View our privacy policy here.  *a,
Faisal Hayata,
Shahan Zeb Khanab,
Graeme Hogarthc,
Tamara Kondratyukd,
John M. Pezzutode and
Muhammad Nawaz Tahirf
*a,
Faisal Hayata,
Shahan Zeb Khanab,
Graeme Hogarthc,
Tamara Kondratyukd,
John M. Pezzutode and
Muhammad Nawaz Tahirf

![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S), 1001 ν(C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.45 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 2.93 (t, 4H, H3,3′, 3J 1H1H = 5.1 Hz), 3.39 (s, 3H, H8), 6.80–6.92 (m, 4H, benzene, H5,6). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.6 (C1); 50.3 (C2,2′), 51.3 (C3,3′); 145.9 (C4), 118.0 (C5,5′), 114.6 (C6,6′), 153.4 (C7), 55.6 (C8).
S), 1001 ν(C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.45 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 2.93 (t, 4H, H3,3′, 3J 1H1H = 5.1 Hz), 3.39 (s, 3H, H8), 6.80–6.92 (m, 4H, benzene, H5,6). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.6 (C1); 50.3 (C2,2′), 51.3 (C3,3′); 145.9 (C4), 118.0 (C5,5′), 114.6 (C6,6′), 153.4 (C7), 55.6 (C8).![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1572 ν (N-CSS), 1209 ν (C-NC2), 1015 ν (C
O), 1572 ν (N-CSS), 1209 ν (C-NC2), 1015 ν (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S), 1003 ν (C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.39 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 3.62 (s, br, 4H, H3,3′), 7.02 (d, 1H, H6, 3J 1H1H = 3.6 Hz), 6.62 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.84 (t, 1H, H8, 3J 1H1H = 0.9 Hz). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.9 (C1); 49.2 (C2,2′); 40.8 (C3,3′); 159.0 (C4), 147.4 (C5), 116.1 (C6), 111.8 (C7), 145.2 (C8).
S), 1003 ν (C–S). 1H NMR {(DMSO-d6, 300 MHz), δ (ppm)}: 4.39 (t, 4H, H2,2′, 3J 1H1H = 5.1 Hz), 3.62 (s, br, 4H, H3,3′), 7.02 (d, 1H, H6, 3J 1H1H = 3.6 Hz), 6.62 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.84 (t, 1H, H8, 3J 1H1H = 0.9 Hz). 13C NMR {(DMSO-d6, 75 MHz), δ (ppm)}: 214.9 (C1); 49.2 (C2,2′); 40.8 (C3,3′); 159.0 (C4), 147.4 (C5), 116.1 (C6), 111.8 (C7), 145.2 (C8).![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1585 ν (N-CSS), 1223 ν (C-NC2), 1009 ν (C–S), 361 ν (Pt–S), 287 ν (Pt–Cl), 266 ν (Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.93 (m, 4H, H2,2′), 3.77 (m, 4H, H3,3′), 7.23 (d, 1H, H6, 3J 1H1H = 3.4 Hz), 6.54 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.50 (d, 1H, H8, 3J 1H1H = 0.9 Hz), 7.64–7.72 (m, Hb,b′), 7.10–7.22 (m, Hc,c′). 13C NMR {(CDCl3-75 MHz) δ (ppm)}: 207.2 (C1); 46.8 (C2,2′), 46.6 (C3,3′); 158.9 (C4), 147.2 (C5), 118.2 (C6), 111.9 (C7), 144.3 (C8), 162.8 (d, Ca, 4JFC = 3 Hz), 136.4 (dd, Cb, 3JFC = 4.5, 8.25 Hz); 116 (dd, Cc, 2JFC = 9, 12.75 Hz), 124.7 (dd, Cd, 1JFC = 3.75, 57.75 Hz). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 9.6.
O), 1585 ν (N-CSS), 1223 ν (C-NC2), 1009 ν (C–S), 361 ν (Pt–S), 287 ν (Pt–Cl), 266 ν (Pt–P). 1H NMR {(CDCl3, 300 MHz), δ (ppm)}: 3.93 (m, 4H, H2,2′), 3.77 (m, 4H, H3,3′), 7.23 (d, 1H, H6, 3J 1H1H = 3.4 Hz), 6.54 (dd, 1H, H7, 3J 1H1H = 3.3 Hz and 1.8 Hz), 7.50 (d, 1H, H8, 3J 1H1H = 0.9 Hz), 7.64–7.72 (m, Hb,b′), 7.10–7.22 (m, Hc,c′). 13C NMR {(CDCl3-75 MHz) δ (ppm)}: 207.2 (C1); 46.8 (C2,2′), 46.6 (C3,3′); 158.9 (C4), 147.2 (C5), 118.2 (C6), 111.9 (C7), 144.3 (C8), 162.8 (d, Ca, 4JFC = 3 Hz), 136.4 (dd, Cb, 3JFC = 4.5, 8.25 Hz); 116 (dd, Cc, 2JFC = 9, 12.75 Hz), 124.7 (dd, Cd, 1JFC = 3.75, 57.75 Hz). 31P NMR {(CDCl3-161.97 MHz) δ (ppm)}: 9.6.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K
K![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K + log(I/Io − I)
K + log(I/Io − I)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif)
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 678
678![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 012
012![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 915
915