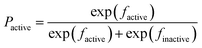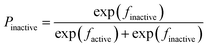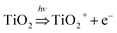Development of nanostructure–activity relationships assisting the nanomaterial hazard categorization for risk assessment and regulatory decision-making†
Guangchao Chen*a,
Willie J. G. M. Peijnenburgab,
Vasyl Kovalishyn c and
Martina G. Vijvera
c and
Martina G. Vijvera
aInstitute of Environmental Sciences CML, Leiden University, Einsteinweg 2, 2333 CC, Leiden, The Netherlands. E-mail: chen@cml.leidenuniv.nl; Fax: +31 71 527 7434; Tel: +31 71 527 7463
bNational Institute of Public Health and the Environment-RIVM, Bilthoven, The Netherlands
cDepartment of Medical and Biological Research, Institute of Bioorganic Chemistry & Petrochemistry, 1 Murmanska Street, Kyiv 02660, Ukraine
First published on 20th May 2016
Abstract
Categorization of the environmental hazards associated with engineered nanomaterials (ENMs) is important for evaluating the potential risks brought by commercialized ENMs. Such a task is so far severely hindered because of an insufficient amount of available toxicity data. As biological assays are costly and time-consuming and also face the ethical issue of animal use, computational modeling such as (quantitative) nanostructure–activity relationships (nano-(Q)SARs) is valued as a potential tool to fill in the data gaps. With this in mind, nano-SARs classifying the ecotoxicity of ENMs were developed in this study with the aims: (i) to examine the availability of nanoecotoxicity data in developing nano-SARs; and (ii) to build nano-SARs that assist the hazard categorization of ENMs for regulatory purposes. Multi-source ecotoxicity data were retrieved, on the basis of which descriptors quantifying the ENM structures were calculated. By employing four extensively used tree algorithms, global nano-SARs across species and species-specific models were derived with significant predictive power. For the LC50 global models, the functional tree, C4.5 decision tree and random tree models all correctly classified more than 70.0% of the samples on training (320 ENMs) and test sets (80 ENMs). The functional tree predicting the toxicity of metallic ENMs to Danio rerio showed accuracies of 93.4% and 100% on, respectively, training (76 ENMs) and test sets (18 ENMs). Descriptors present in the species-specific models were analyzed to discuss the key factors affecting nanotoxicity. With easily obtained descriptors and transparent predictive rules, we believe the developed nano-SARs could assist the expedited review of ENMs' hazards and facilitate better-informed regulatory decisions of ENMs.
Introduction
Assessing the potential environmental risks posed by engineered nanomaterials (ENMs) is essential to ensure that the marketed ENMs are used as safely as possible. It is believed that, a preliminary categorization of ENMs will benefit the early stages of qualitative risk analysis either by manufacturers or by regulators, to target the ENMs of high risk concerns and so as to prioritize more detailed testing of ENMs.1 The European Chemicals Agency, for instance, has released reports and documents alike to address the usefulness of ENM grouping serving to the streamline testing for the regulatory purposes.1 The U.S.-Canada Regulatory Cooperation Council also reported development of the classification scheme for ENMs in order to identify the ‘ENMs of concern’ that are likely to behave differently compared to their bulk scale counterparts.2 Generally, one of the commonly used strategies of ENM categorization is to group ENMs based on different measures of biological activities. An example of this can be found in the CLP-Regulation (EC) No. 1272/2008, which suggests that chemicals can be classified as acutely toxic or as chronically toxic at multiple levels according to the outcomes of standardized toxicity tests.3,4 Another example is the EU Directive 93/67/EEC that recommends to rank the chemical hazard to aquatic species into four hierarchies, i.e. very toxic, toxic, harmful and not classified, on the basis of at least three standard test species algae, crustacea and fish.5 Unsurprisingly, however, those risk potential-based material categorizations require an enormous amount of hazard information of ENMs intended for adequately evaluating the safety of the materials. Given the substantial number of existing, non-tested ENMs and the rapid growth of ENMs innovation, it is, consequently, expected that alternatives of testing assays such as (quantitative) nanostructure–activity relationships (termed as nano-(Q)SARs) could be effectively used to fill in data gaps while with the minimum of financial cost and time consumption. The application of (Q)SARs in ENM categorization is seen to be quite advanced as it is capable of promoting the safe use of ENMs.6 Meanwhile, employing (Q)SARs in ENMs' risk assessment also meets the 3R's principle (refine, reduce or replace) of animal use in toxicity testing.7Previously, a few nano-(Q)SAR models have been established by linking ENMs' biological responses to the experimental and/or computational characterization of ENMs.8 One of the issues so far in developing nano-(Q)SARs is that a relatively small number of datasets were repeatedly used by different studies.9 This may be because of one of the obstacles of using multi-source data in developing nano-(Q)SARs being the lack of data consistency between diverse researches. This lack of data leads to the difficulty of comprehensively characterizing the structures of ENMs in an entire dataset especially for fully quantifying the information on surface coatings and functional groups of ENMs. However, given the constantly increasing amount of scientific resources from numerous scientific programs on nanomaterial safety, and given the urgent need of further development in computational nanotoxicology to assist the risk assessment of nanomaterials, nano-(Q)SARs based on the integration and maximization of the use of existing nanotoxicity data also seems to be of particular importance. We hence aimed to derive classification nano-SARs by using the currently available and accessible nanotoxicity data on environmental species shared in various publications and scientific resources. Feasible strategy of computationally characterizing the structures of ENMs was chosen. The purposes of this study are summarized as, firstly, to examine the availability of existing nanoecotoxicity data in developing nano-SARs; and secondly, to build classification models for ENMs assisting the nanomaterial hazard categorization for estimating the risks of metal-based nanomaterials.
To begin with, three datasets were obtained from various publications and scientific resources, and considered for the use of modeling. The structural descriptors were calculated using a web-based platform Online Chemical Modeling Environment (OCHEM) which characterize the information of the core of metal-based ENMs.10 To acquire transparent and easily applicable classification models, four extensively employed tree algorithms embedded in the Weka (version 3.6) were considered for modeling, namely functional tree, C4.5 decision tree, random tree and simple CART.11 Based on the descriptors and algorithms, global nano-SARs across species as well as species-specific models were developed with significant predictability. The global models are favorable for ranking the general biological effects of ENMs regardless of targeted species, while species-specific models are able to offer in depth knowledge of nanotoxicity and may also be more applicable when the estimation of nanotoxicity is based on certain species (e.g. categorize ENMs based on EU Directive 93/67/EEC). Descriptors appearing in the species-specific nano-SARs were analyzed in light of a mechanistic interpretation of the toxicity triggered by metallic ENMs. The present study examined the availability of published nanoecotoxicity data in deriving nano-(Q)SARs and demonstrated the possibility of building nano-SARs using multi-source datasets.
Methods
Datasets
We previously established a database summarizing and describing the toxicity of metal-based ENMs to selected organisms in light of the development of nano-(Q)SARs.8 Records of the commonly used toxicity endpoints in this database, including EC50 (the effective concentration that causes 50% response), EC20, LC50 (the concentration which leads to 50% mortality), LC20, MIC (minimum inhibitory concentration) and NOEC (no observed effect concentration) were manually uploaded to the web-based platform on 18th August, 2015.10 Using the OCHEM platform, an analysis of the available ecotoxicity data of metal-based ENMs was performed on 28th August, 2015, which provided us with three datasets containing the toxicity of various metal-based ENMs to different hierarchies of species: (I) 400 ENMs from 90 publications or reports provided with experimental data on LC50; (II) 450 ENM records from 79 publications or reports with quantitative information on EC50 values; and (III) 166 ENMs obtained from 13 publications with experimental values of the MIC. MIC characterizes the antimicrobial properties of ENMs and is therefore a common experimental endpoint in antimicrobial assays. Even though the use of MIC does not currently fit into the scheme of evaluating ENMs' risks based on different species, we still included this case study so as to further examine the feasibility of building nano-SARs for different hierarchies of species. Units of the toxicity values were unified into mg L−1 in the datasets. For building global nano-SARs across species, the three datasets I, II and III were used as three case studies. As for constructing models for single species, from each of the datasets two species with the most toxicity endpoint records were chosen. As a result, the selected species were Danio rerio (94 records including embryo, LC50), Daphnia magna (102, LC50), Pseudokirchneriella subcapitata (66, EC50), Daphnia magna (105, EC50), Escherichia coli (41, MIC), and Staphylococcus aureus (39, MIC).As it is acknowledged, thresholds that discretize the numeric values are of significant importance for building classification models, which thus should be carefully discussed and selected on the basis of different strategies and application requirements.12 In this study, we initially examined the tendency of model predictability with the shift of threshold values. And afterwards thresholds that lead to the most balanced predictive performances were conditionally considered. Referring to the regulations and directives nowadays in force, consideration of the thresholds for global models was restricted to the values of 0.1, 1.0, 10.0 and 100.0 mg L−1, which are, for instance, used by both the aforementioned CLP-Regulation (EC) No. 1272/2008 and the EU Directive 93/67/EEC. For the species-specific nano-SARs, thresholds of 1.0, 10.0, 100.0 mg L−1 were taken (for Escherichia coli and Staphylococcus aureus only 10.0 and 100.0 mg L−1 because of narrower variation of toxicity values). Within each dataset the records were ranked based on the values of the toxicity endpoints. ENMs with toxicity values less than pre-specified threshold value were assigned to the ‘active’ class, and the rest of ENMs were labeled as ‘inactive’. When building models, 20% of the dataset was exclusively utilized for external validation.
Descriptor calculation
Obtaining the structural descriptors of ENMs is essential to characterize the structures of ENMs besides the experimental measures. Using the ‘Calculate descriptors’ function implemented in OCHEM, three types of descriptors were calculated and acquired, the E-state, ALogPS, and Chemaxon descriptors. For the E-state, both atom and bond types were considered for the indices and counts descriptors during calculation. The selected subgroups of Chemaxon descriptors are elemental analysis, charge, geometry, partitioning, protonation and isomers that are generated at the specified pH value 7.4. For deriving global nano-SARs, all the three types of descriptors were considered. And as for species-specific models the selection of descriptors was narrowed down to the ALogPS and Chemaxon descriptors in order to allow for easier and better understanding of the underlying toxicity mechanisms with the assistance of the descriptors.Modeling algorithms
In order to build transparent rule-based nano-SARs that are easy to interpret and are capable of revealing information insight into the roles of structural descriptors, tree algorithms in Weka (version 3.6) were considered in the study. To avoid coincidence and also compare model performance, four extensively employed tree methods were used including functional tree, C4.5 decision tree, random tree and simple CART.11In a functional tree model, both decision nodes and leaf nodes could contain tests based on either original input descriptors or the logistic regressions of descriptors.13 For binary classifications, prediction in the leaf nodes using logistic regressions of descriptors could be explained as in Fig. 1, where Pactive and Pinactive are categorical possibilities needed to be compared; factive and finactive are the regressions of descriptors generated by the algorithm; inactive and active are the class labels to be returned for an observation.
The C4.5 decision tree is an extension of the earlier ID3 algorithm.14 It generates decision-based tree models in which each inner node contains a test only on the original input descriptors.15 For each test, a splitting cut-off value is provided and used for value comparison. The classification of ENM toxicity is accomplished by traversing a tree model from the root node to leaf nodes. Upon reaching the leaf nodes, labels (active or inactive) stored in the nodes will be returned as predictions.
The random tree algorithm constructs a tree randomly from a set of possible trees in which each tree has an equal chance of being sampled.16 A random tree is grown (without pruning) from data that has k randomly selected attributes at each node.17 The decision nodes contain queries only employing input descriptors and splitting thresholds, and leaf nodes comprise the category labels that an observation will be classified as. In the study, the k-value was set at 0 by default and the number of randomly chosen attributes was determined as log2(number of attributes) + 1. No depth restriction was set as the ‘maxDepth’ was 0 by default.
As a decision tree learner for classification, the simple CART (classification and regression tree) employs the minimal cost-complexity pruning of the CART algorithm when constructing predictive trees.18 It finds cost-complexity, a measure of average error reduced per leaf, and calculates the number of errors for each node when the subtrees are replaced by leaves.19 The simple CART generates binary decision tree models for categorization issues. It handles the missing data by ignoring that record.20
Model performance evaluation
To estimate the predictive power of generated models, each dataset was randomly split into a training set (80%) and a test set (20%) before model construction. The learning process on the training set was executed in 10-fold cross validation to ensure the model stability. Predictive accuracy was characterized by four statistical parameters, defined as sensitivity (SE = TP/AP), specificity (SP = TN/AN), accuracy (Q = (TP + TN)/(AP + AN)), and correct classification rate (CCR = 0.5(sensitivity + specificity)). Thereinto, TP represents the predicted number of true positives (or active class), TN stands for the predicted number of true negatives (or inactive class). AP and AN are numbers of actual positives and negatives observed, respectively. Reportedly, classification accuracy higher than 70% is considered as high predictive performance.21 And classification models with CCR of both training and test sets higher than 60.0% would be considered acceptable.22 Model complexity was characterized by the size of the tree (number of nodes). Additionally, the significance of test sets was also verified by randomly permuting class labels of the test sets for global nano-SARs. The predictive results on these disjoint datasets should be approximately 50% (close to the no-information rate) for binary classifications with balanced datasets.23Results and discussion
Global nano-SARs across species
The influence of cut-off thresholds on model performances was primarily studied using the datasets I II and III. As can be seen in Fig. S1,† both high (0.1 mg L−1) and low (100.0 mg L−1) threshold values were evidenced to result in biased predictions. The thresholds selected for dataset I (LC50), II (EC50) and III (MIC) are respective 1.0, 10.0 and 10.0 mg L−1 to discretize numeric values for the case studies. After data discretization, dataset I was found to contain 175 ENMs of the active class and 225 of the inactive class; dataset II consisted of 246 ENMs labeled as active and 204 labeled as inactive; and dataset III has 87 ENMs from the active group and 79 from the inactive group. Using the OCHEM platform, 107, 95 and 122 computational descriptors were obtained for the datasets I, II and III, respectively. Different nano-SARs were derived based on the descriptors which were linked to the nanotoxicity by the functional tree, C4.5 decision tree, random tree and simple CART algorithms. An overview of the generated classification models is given in Table 1, in terms of modeling method, size of tree, sub-dataset, sensitivity, specificity, accuracy, and CCR. More details of the developed nano-SARs can be found in the ESI.†| Method | Size of tree | Dataset | Sensitivity | Specificity | Accuracy | CCR |
|---|---|---|---|---|---|---|
| Case study I – LC50 (ntraining = 320, ntest = 80), threshold value 1.0 mg L−1 | ||||||
| FT | 1 | Training set | 0.750 | 0.678 | 0.709 | 0.714 |
| Test set | 0.686 | 0.733 | 0.713 | 0.710 | ||
| C4.5 | 5 | Training set | 0.671 | 0.750 | 0.716 | 0.711 |
| Test set | 0.686 | 0.733 | 0.713 | 0.710 | ||
| RT | 55 | Training set | 0.679 | 0.728 | 0.706 | 0.704 |
| Test set | 0.629 | 0.778 | 0.713 | 0.704 | ||
| Simple CART | 11 | Training set | 0.707 | 0.678 | 0.691 | 0.693 |
| Test set | 0.686 | 0.689 | 0.688 | 0.688 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Case study II – EC50 (ntraining = 360, ntest = 90), threshold value 10.0 mg L−1 | ||||||
| FT | 1 | Training set | 0.741 | 0.503 | 0.633 | 0.622 |
| Test set | 0.796 | 0.415 | 0.622 | 0.606 | ||
| C4.5 | 9 | Training set | 0.695 | 0.546 | 0.628 | 0.621 |
| Test set | 0.816 | 0.415 | 0.633 | 0.616 | ||
| RT | 39 | Training set | 0.741 | 0.479 | 0.622 | 0.610 |
| Test set | 0.816 | 0.439 | 0.644 | 0.628 | ||
| Simple CART | 17 | Training set | 0.650 | 0.564 | 0.611 | 0.607 |
| Test set | 0.796 | 0.439 | 0.633 | 0.618 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Case study III – MIC (ntraining = 133, ntest = 33), threshold value 10.0 mg L−1 | ||||||
| FT | 3 | Training set | 0.743 | 0.762 | 0.752 | 0.753 |
| Test set | 0.706 | 0.688 | 0.697 | 0.697 | ||
| C4.5 | 3 | Training set | 0.743 | 0.778 | 0.759 | 0.761 |
| Test set | 0.706 | 0.688 | 0.697 | 0.697 | ||
| RT | 13 | Training set | 0.814 | 0.587 | 0.707 | 0.701 |
| Test set | 0.706 | 0.688 | 0.697 | 0.697 | ||
| Simple CART | 3 | Training set | 0.743 | 0.778 | 0.759 | 0.761 |
| Test set | 0.706 | 0.688 | 0.697 | 0.697 | ||
For case study I, the learning process was executed on the basis of 320 ENMs in the training set, while models were validated on the test set comprising 80 ENMs. A cut-off value of 1.0 mg L−1 was applied to enable the derivation of nano-SARs. By comparison, functional tree, C4.5 decision tree and simple CART generated tree models with relatively low complexity (size of tree are respective 1, 5 and 11). As shown in Table 1, the random tree model was observed to be larger with a tree size of 55. These nano-SARs applied to the training set yielded accuracies of 70.9% (functional tree), 71.6% (C4.5 decision tree), 70.6% (random tree) and 69.1% (simple CART). Except for the simple CART model which correctly predicted 68.8% of the observations from the test set, accuracies of the LC50-related nano-SARs on the test set were all found to exceed 70.0%. The CCR values calculated on sensitivity and specificity are higher than 60.0% for all the four models. Specifically, the C4.5 decision tree model merely contains two structural descriptors maximalprojectionsize and molecularpolarizability which belong to the Chemaxon descriptors. The descriptor maximalprojectionsize relates to the size of the molecule perpendicular to the minimal projection area surface (based on the van der Waals radius). And molecularpolarizability associates with the polarizability of the molecule. This means that the influence of both size and polarizability of the core element of ENMs was indicated. The simple CART model consists of five descriptors correlated with the geometrical size (minimalprojectionsize, maximalprojectionarea, minimalprojectionradius), molecular polarizability (averagemolecularpolarizability), and accessible surface areas of all atoms with negative partial charge (asa_ASA−). Owing the higher model complexity, however, the simple CART model was found to yield no higher predictive performance compared to the C4.5 decision tree. The functional tree has a relatively simpler tree structure with only one node but used more input descriptors in the logistic regressions.
With respect to the case study II, the 450 ENMs were randomly distributed to a training set of 360 ENMs and a test set of 90 ENMs. Numeric values of EC50 were discretized by a threshold of 10.0 mg L−1. ENMs with EC50 values less than 10.0 mg L−1 were labeled as active, and the rest of ENMs were considered inactive. From the results shown in Table 1, accuracies of all the models are between 60.0% and 65.0% for both training sets and test sets. This resulted from the low specificity of the nano-SARs while the models' sensitivities were considered reasonable. Thus the constructed EC50 models possess relatively low predictability for the inactive class. The unbalanced performances on both classes also resulted in the low CCRs between 60.0% and 65.0%.
Moreover, SAR-like models were also developed to predict the MICs of ENMs to various bacteria. In case study III, 133 ENMs were used to train the models and 33 ENMs were left out for the external validation. A threshold of 10.0 mg L−1 categorizes the ENMs into the active class (MIC < 10.0 mg L−1) or the inactive class (MIC ≥ 10.0 mg L−1). The results depicted in Table 1 show that the C4.5 decision tree and the simple CART models exhibited the best predictability on the training set (both 75.9%), followed by the functional tree (75.2%) and the random tree models (70.7%). Predictive performances of the four nano-SARs on the test set gave the same results of 69.7% accuracy. CCRs of the training set are higher than 70.0% and those of the four test sets are all 69.7%. Except the most complex random tree model, the functional tree, C4.5 decision tree and simple CART models have the same tree size of 3. Meanwhile, for both the C4.5 decision tree and the simple CART only the structural descriptor ALogPS_logS appeared in the built nano-SARs which is associated with water solubility. The functional tree constructed the models using ten descriptors in its logistic regressions as can be seen in the ESI.†
The LC50 related functional tree, C4.5 decision tree and random tree models showed reasonable predictability with accuracy (on training and test sets) higher than 70.0% and CCR higher than 60.0%, and with balanced performances on both categories. Based on a training set of 320 ENMs and test set of 80 ENMs, the C4.5 decision tree model is seen as relatively more concise as it only contains 5 nodes in the tree and uses two structural descriptors (maximalprojectionsize and molecularpolarizability), as shown in Fig. 2. Models presented in case study III were also considered acceptable based on the sensitivity, specificity, accuracy, CCR and also tree complexity. As the developed nano-SARs exhibited similar predictive results on test sets, the significance of the test sets used in external validation was subsequently examined. We permuted the class labels in each test set for five times and validated the models with these randomized datasets afterwards. The results are depicted in Fig. 3. As to case study I, the predictive accuracies on permuted test sets are between 46.3% and 58.8%. For case study II and III, it is 42.2–55.6% and 39.4–57.6%, respectively. Thus for all three cases, performances of the developed nano-SARs on the disjoint datasets are approximately 50% which is close to the no-information rate for binary classifications.23 It is therefore concluded that the original test sets are significant for model validation in the case studies I, II and III.
 | ||
| Fig. 2 Developed C4.5 decision tree for the LC50 of metal-based ENMs. If LC50 < 1.0 mg L−1 the ENM is judged as active, and if LC50 ≥ 1.0 mg L−1 the ENM is inactive. | ||
Species-specific nano-SARs
Besides global models, species-specific nano-SARs were also built using the retrieved experimental data. This is in accordance with the recommendation of EU Directive 93/67/EEC ranking the hazards of ENMs to aquatic species. To begin with, from each dataset two species with the most data records were chosen for model development, which are Danio rerio (94 records) and Daphnia magna (102 records) from dataset I, Daphnia magna (105 records) and Pseudokirchneriella subcapitata (66 records) from dataset II, and Escherichia coli (41 records) and Staphylococcus aureus (39 records) from dataset III. For building models, two typical tree algorithms among the four selected methods, the functional tree and C4.5 decision tree algorithms were employed along with the ALogPS and Chemaxon descriptors. Cut-off thresholds investigated are respective 1.0, 10.0 and 100.0 mg L−1. Performances of the derived nano-SARs are summarized in Tables S1–S3 in the ESI.† Models that exhibited significant predictive power are summarized and described in Table 2 and Fig. 4. Nano-SARs were obtained for different hierarchies of species, i.e. Danio rerio (fish), Daphnia magna (crustacean), Pseudokirchneriella subcapitata (algae), and Staphylococcus aureus (bacteria). Details of these nano-SARs are presented in Table 3, including the number of ENMs, size of the developed tree model, number of descriptors and the names of descriptors involved.| Threshold (mg L−1) | Dataset | Sensitivity | Specificity | Accuracy | CCR | |
|---|---|---|---|---|---|---|
| Danio rerio, ntraining = 76, ntest = 18, LC50 | ||||||
| FT | 100.0 | Training set | 0.943 | 0.913 | 0.934 | 0.928 |
| Test set | 1.000 | 1.000 | 1.000 | 1.000 | ||
| C4.5 | Training set | 0.906 | 0.913 | 0.908 | 0.910 | |
| Test set | 1.000 | 1.000 | 1.000 | 1.000 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Daphnia magna, ntraining = 82, ntest = 20, LC50 | ||||||
| FT | 1.0 | Training set | 0.843 | 0.968 | 0.890 | 0.906 |
| Test set | 0.750 | 1.000 | 0.850 | 0.875 | ||
| C4.5 | Training set | 0.843 | 0.968 | 0.890 | 0.906 | |
| Test set | 0.750 | 1.000 | 0.850 | 0.875 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Pseudokirchneriella subcapitata, ntraining = 53, ntest = 13, EC50 | ||||||
| FT | 1.0 | Training set | 0.944 | 0.914 | 0.925 | 0.929 |
| Test set | 0.750 | 1.000 | 0.923 | 0.875 | ||
| C4.5 | Training set | 0.944 | 0.914 | 0.925 | 0.929 | |
| Test set | 0.750 | 1.000 | 0.923 | 0.875 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||
| Staphylococcus aureus, ntraining = 32, ntest = 7, MIC | ||||||
| C4.5 | 100.0 | Training set | 0.833 | 0.875 | 0.844 | 0.854 |
| Test set | 0.800 | 1.000 | 0.857 | 0.900 | ||
| Nano-SAR | Method | ENMs number | Tree size | Descriptor number | List of descriptors |
|---|---|---|---|---|---|
| Danio rerio LC50 values | FT | 94 | 3 | 7 | Averagemolecularpolarizability, molecularpolarizability, mass, volume, plattindex, apKb1, ALogPS_logS |
| C4.5 | 94 | 5 | 2 | Exactmass, asa_ASA | |
| Daphnia magna LC50 values | FT | 102 | 1 | 8 | Molecularpolarizability, tholepolarizability_a_xx, tholepolarizability_a_zz, exactmass, volume, logp, asa_ASA+, asa_ASA_P |
| C4.5 | 102 | 3 | 1 | asa_ASA− | |
| Pseudokirchneriella subcapitata EC50 values | FT | 66 | 1 | 8 | Molecularpolarizability, tholepolarizability_a_yy, mass, minimalprojectionarea, volume, dreidingenergy, hyperwienerindex, ALogPS_logS |
| C4.5 | 66 | 3 | 1 | Minimalprojectionarea | |
| Staphylococcus aureus MIC values | C4.5 | 39 | 3 | 1 | ALogPS_logS |
The nano-SARs categorizing nanotoxicity to Danio rerio gave accuracies of 93.4% (functional tree) and 90.8% (C4.5 decision tree) on corresponding training sets (76 ENMs), and 100% accuracy on the two test sets (18 ENMs). Sensitivity and specificity of the two models are all above 90.0% on the training and test sets (Table 2). This demonstrates the high predictability of the developed models. Model stability was ensured by executing 10-fold cross validation. Size of the corresponding functional tree model is 3 which means the nano-SAR only consists of one inner node and two decision nodes. As to Daphnia magna, the training set has 82 ENMs as samples for the learning process and the test set is comprised by 20 ENMs for validation. Accuracies of both the functional tree and the C4.5 decision tree models were shown to be 89.0% (training set) and 85.0% (test set) that are statistically significant. The CCRs of the model exceeded 85.0%. As shown in Table 3, the sizes of the functional tree and the C4.5 decision tree are respectively 1 and 3. With regards to Pseudokirchneriella subcapitata, functional tree and C4.5 decision tree models were built on the basis of 53 ENMs and validated by 13 ENMs. Predictive accuracies are as high as 92.5% on training set and 92.3% on test set with regard to both the functional tree and C4.5 decision tree with high CCR values. Moreover, built on a training set of 32 ENMs, the C4.5 decision tree model predicting the MIC to Staphylococcus aureus also exhibited significant predictability of 84.4% and 85.7% for the training and test set, respectively.
Notably, even though mechanisms of the toxicity induced by metal-based ENMs to various hierarchies of species may vary, some descriptors in the models characterizing similar factors of ENMs were commonly observed and identified. As shown in Table 3 and Fig. 4, descriptors representing molecular polarizability frequently appeared in the functional tree models. Those descriptors include the averagemolecularpolarizability, molecularpolarizability, tholepolarizability_a_xx, tholepolarizability_a_zz and tholepolarizability_a_yy, which characterize different aspects of the electronic polarizability's contribution to nanotoxicity. Molecular polarizability measures the ability of the outer shell electrons in a molecule to move easily toward an external perturbation.24 Higher polarizability of the electrons in a molecule results in easier movement of electrons induced by an external electric field, which may trigger a series of biological reactions and lead to the toxicity of the materials.25 For instance, detachment of an electron activated by solar radiation could stimulate the generation of hydroxyl radical OH˙ as described in the study of Kar et al.:26
| e− + O2 → O2˙− |
| O2˙− + 2H+ + e− → H2O2 |
| O2˙− + H2O2 → OH˙ + OH− + O2 |
| H+ + H2O ⇒ OH˙ + H+ |
Another discriminating factor is the accessible surface area of ENM cores that is quantified by asa_ASA (solvent accessible surface area), asa_ASA+ (solvent accessible surface area of all atoms with positive partial charge), asa_ASA_P (solvent accessible surface area of all polar atoms), and asa_ASA− (solvent accessible surface area of all atoms with negative partial charge) in the nano-SARs. The accessible surface area is defined as the accessible surface of molecules to a solvent.27 For uncoated ENMs, the exposed surface area to the surroundings reflects the amount of atoms to be displayed on the surface and the potential of molecules to interact with the subcellular structures of species. As acknowledged, one of the outstanding properties of ENMs is the higher surface/volume ratio compared to that of their bulk counterparts which provides them increased surface reactivity and therefore possibly high toxicity.28 As surface coatings are able to influence the toxicity of ENMs to species, surface area of ENM core still seems to play a role in nanotoxicity for the ENMs with modified surface. Moreover, descriptors quantifying the solubility were also observed such as apKb1 (dissociation constant) and ALogPS_logS (solubility in water) generated by OCHEM. Previous studies have shown that ENMs with less hardness and high solubility tend to exhibit stronger hazard effects.29 This may be because the metal-ion leaching from ENM surface could act as one of the key factors inducing nanotoxicity.30,31 Take Cu ENMs as an example, the release of Cu2+ from Cu-based nanoparticles could cause the generation of OH˙ as follows:32
| O2˙− + Cu2+ → O2 + Cu+ |
| Cu+ + H2O2 → Cu2+ + OH− + OH˙ |
The toxicity of ENMs may occur when the derived reactive oxygen species and the ions per se jointly or independently interact with the subcellular structures of species. Meanwhile, the geometrical descriptors minimalprojectionarea and minimalprojectionarea were also utilized in the model which indicate the spatial arrangement of the atoms forming a molecule. These descriptors are associated with the molecular surface information obtained from atomic van der Waals areas and their overlap.25 The descriptors relate to mass (mass, exactmass) and complexity (plattindex) were used in the nano-SARs as well. The platt index is the sum of the degrees of all edges in the molecular graph, and is a considerably better measure of molecular complexity than merely the number of edges.33,34
Implications to the risk assessment of ENMs
On the basis of the computational descriptors offered by OCHEM and the assembled ecotoxicity data of metal-based nanomaterials, the developed LC50- and MIC-related global models and the species-specific nano-SARs showed reasonable predictive power. This demonstrates that it is indeed feasible to build nano-SARs using multi-source datasets if the structures of ENMs are appropriately characterized. It also again confirms that the nano-(Q)SARs ought to be viewed as a potentially helpful tool in assisting the expedited review of ENM hazard categorization for the risk assessment of nanomaterials. With the experimental data retrieved from different scientific resources inconsistently characterizing the structures of ENMs, we managed to build nano-SARs classifying the nanoecotoxicity using descriptors solely representing the ENM cores. Such modeling tasks employing large datasets critically rely on the availability and quality of the datasets, and also on the comprehensive representation of ENM structures based on provided information. To accelerate the development of (Q)SAR-like models for nanomaterials much needs to be improved. Agreement on better data quality and availability are essential for nano-(Q)SARs with respect to both the toxicological and the componential aspects of the studied ENMs.9 That is, the problem so far of the successful application of computational nanotoxicology is rather experimental, together with inadequate computational quantifications of ENM structures, than mathematical or statistical.9 Unlike individual chemicals that are structurally unambiguous and possibly less complex, nanomaterials often exist as populations of materials varying in sizes, shapes, composites and functional groups, etc. which can all significantly influence their biological interactions with environmental species.8 The structural uncertainty of the materials brings difficulty to experimentalists to offer complete and precise characterization of ENM structures, which subsequently hinders the calculation of representative descriptors for ENMs even when the compositions may have been properly provided.35 The lack of data consistency especially in characterizing the structure of ENMs prevents the use of experimental data in developing nano-(Q)SARs, and may be one of the driving reasons why only a few datasets have been repeatedly used by the state-of-art of nano-(Q)SARs.Conclusions
In this study, global nano-SARs across species and species-specific models classifying the ecotoxicity of metal-based ENMs were proposed. The models are intended to assist the nanomaterial hazard categorization and facilitate the ENM-related risk assessment and regulatory decision-making. To test the availability of existing nanotoxicity data in developing nano-(Q)SARs, datasets containing ecotoxicity information of ENMs from various publications or scientific resources were used including the LC50 (400 ENMs), EC50 (450 ENMs) and MIC (166 ENMs) related datasets. Due to the limited information characterizing the coating and functional groups of ENMs, descriptors were generated by the OCHEM to represent the core of the metal-based ENMs. Using the tree algorithms selected, easily interpretable and applicable classification nano-SARs were derived with significant predictability. The LC50 and MIC related global nano-SARs exhibited up to more than 70% accuracy of classification. The species-specific models were also developed to categorize the toxicity of metal-based ENMs to Danio rerio, Daphnia magna, Pseudokirchneriella subcapitata, and Staphylococcus aureus. Descriptor analysis indicated the role of molecular polarizability, accessible surface area and metal-ion leaching in affecting the ecotoxicity of ENMs.Acknowledgements
We thank the WEKA Machine Learning Project for the open-source software, and the OCHEM Team for the free access of the OCHEM platform. Guangchao Chen greatly thanks the funding support by the Chinese Scholarship Council (201306060076). This research is funded by the NATO project number SFPP 984401. Martina Vijver is funded by NWO-VIDI project number 864.13.010.References
- H. Godwin, C. Nameth, D. Avery, L. L. Bergeson, D. Bernard, E. Beryt, W. Boyes, S. Brown, A. J. Clippinger, Y. Cohen, M. Doa, C. O. Hendren, P. Holden, K. Houck, A. B. Kane, F. Klaessig, T. Kodas, R. Landsiedel, I. Lynch, T. Malloy, M. B. Miller, J. Muller, G. Oberdorster, E. J. Petersen, R. C. Pleus, P. Sayre, V. Stone, K. M. Sullivan, J. Tentschert, P. Wallis and A. E. Nel, ACS Nano, 2015, 9, 3409–3417 CrossRef CAS PubMed.
- Regulatory Cooperation Council – Nanotechnology Initiative (RCC-NI), Work element 2. Development of a classification scheme for nanomaterials regulated under the new substances programs of Canada and the United States, 2013, p. 17 Search PubMed.
- Regulation (EC) No. 1272/2008 of the European Parliament and of the Council on classification, labelling and packaging of substances and mixtures, Official Journal of the European Union, L353, 2008, pp. 1–1355.
- K. Juganson, A. Ivask, I. Blinova, M. Mortimer and A. Kahru, Beilstein J. Nanotechnol., 2015, 6, 1788–1804 CrossRef CAS PubMed.
- Commission of the European Communities (CEC), Technical Guidance Document in Support of Commission Directive 93/67/EEC on Risk Assessment for New Notified Substances. Part II, Environmental Risk Assessment, Office for Official Publications of the European Communities, Luxembourg, Luxembourg, 1996 Search PubMed.
- R. Tantra, C. Oksel, T. Puzyn, J. Wang, K. N. Robinson, X. Z. Wang, C. Y. Ma and T. Wilkins, Nanotoxicology, 2015, 9, 636–642 CrossRef CAS PubMed.
- W. M. S. Russell and R. L. Burch, The Principles of Humane Experimental Technique, Methuen, London, 1959 Search PubMed.
- G. Chen, M. G. Vijver and W. J. G. M. Peijnenburg, Altern. Lab. Anim., 2015, 43, 221–240 Search PubMed.
- D. Winkler, Toxicol. Appl. Pharmacol., 2016, 299, 96–100 CrossRef CAS PubMed.
- I. Sushko, S. Novotarskyi, R. Körner, A. K. Pandey, M. Rupp, W. Teetz, S. Brandmaier, A. Abdelaziz, V. V. Prokopenko, V. Y. Tanchuk, R. Todeschini, A. Varnek, G. Marcou, P. Ertl, V. Potemkin, M. Grishina, J. Gasteiger, C. Schwab, I. I. Baskin, V. A. Palyulin, E. V. Radchenko, W. J. Welsh, V. Kholodovych, D. Chekmarev, A. Cherkasov, J. Aires-de-Sousa, Q. Y. Zhang, A. Bender, F. Nigsch, L. Patiny, A. Williams, V. Tkachenko and I. V. Tetko, J. Comput.–Aided Mol. Des., 2011, 25, 533–554 CrossRef CAS PubMed.
- M. Hall, E. Frank, G. Holmes, B. Pfahringer, P. Reutemann and I. H. Witten, SIGKDD Explor., 2009, vol. 11, pp. 10–18 Search PubMed.
- R. Liu, H. Y. Zhang, Z. X. Ji, R. Rallo, T. Xia, C. H. Chang, A. Nel and Y. Cohen, Nanoscale, 2013, 5, 5644–5653 RSC.
- J. Gama, Mach. Learn., 2004, 55, 219–250 CrossRef.
- J. R. Quinlan, Mach. Learn., 1986, 1, 81–106 Search PubMed.
- J. R. Quinlan, C4.5: Programs for Machine Learning, Morgan Kaufmann, San Francisco, USA, 1993 Search PubMed.
- Y. Zhao and Y. Zhang, Adv. Space Res., 2008, 41, 1955–1959 CrossRef.
- M. Kukreja, S. A. Johnston and P. Stafford, BMC Bioinf., 2012, 13, 139 CrossRef PubMed.
- I. Witten, E. Frank and M. Hall, Data Mining: Practical Machine Learning Tools and Techniques, Morgan Kaufmann, San Mateo, CA, 3rd edn, 2011 Search PubMed.
- S. Rajput and A. Arora, Int. J. Comput. Appl. Tech., 2013, 75, 6–12 Search PubMed.
- S. Kalmegh, Int. J. Innov. Sci. Eng. Technol., 2015, 2, 438–446 Search PubMed.
- V. V. Kleandrova, F. Luan, H. González-Díaz, J. M. Ruso, A. Melo, A. Speck-Planche and M. N. Cordeiro, Environ. Int., 2014, 73, 288–294 CrossRef CAS PubMed.
- D. Fourches, D. Pu, C. Tassa, R. Weissleder, S. Y. Shaw, R. J. Mumper and A. Tropsha, ACS Nano, 2010, 4, 5703–5712 CrossRef CAS PubMed.
- C. Furlanello, M. Serafini, S. Merler and G. Jurman, BMC Bioinf., 2003, 4, 54 CrossRef PubMed.
- A. R. Katritzky, L. Pacureanu, D. Dobchev and M. Karelson, J. Mol. Model., 2007, 13, 951–963 CrossRef CAS PubMed.
- K. P. Singh and S. Gupta, RSC Adv., 2014, 4, 13215–13230 RSC.
- S. Kar, A. Gajewicz, T. Puzyn, K. Roy and J. Leszczynski, Ecotoxicol. Environ. Saf., 2014, 107, 162–169 CrossRef CAS PubMed.
- J. Zhang, X. Gao, J. Xu and M. Li, Rapid and Accurate Protein Side Chain Prediction with Local Backbone Information, in Research in Computational Molecular Biology, ed. M. Vingron and L. Wong, Springer, Berlin Heidelberg, 2008, pp. 285–299 Search PubMed.
- S. Q. Li, R. R. Zhu, H. Zhu, M. Xue, X. Y. Sun, S. D. Yao and S. L. Wang, Food Chem. Toxicol., 2008, 46, 3626–3631 CrossRef CAS PubMed.
- A. Gajewicz, N. Schaeublin, B. Rasulev, S. Hussain, D. Leszczynska, T. Puzyn and J. Leszczynski, Nanotoxicology, 2015, 9, 313–325 CrossRef CAS PubMed.
- J. Hua, M. G. Vijver, M. K. Richardson, F. Ahmad and W. J. G. M. Peijnenburg, Environ. Toxicol. Chem., 2014, 33, 2859–2868 CrossRef CAS PubMed.
- Y. Xiao, M. G. Vijver, G. Chen and W. J. G. M. Peijnenburg, Environ. Sci. Technol., 2015, 49, 4657–4664 CrossRef CAS PubMed.
- S. J. Stohs and D. Bagchi, Free Radical Biol. Med., 1995, 18, 321–336 CrossRef CAS PubMed.
- A. T. Balaban, I. Motoc, D. Bonchev and O. Mekenyan, Topological indices for structure–activity correlations, in Steric Effects in Drug Design, ed. M. Charton and I. Motoc, Springer, Berlin Verlag, 1983, pp. 21–55 Search PubMed.
- L. Saitta and J. D. Zucker, Abstraction in Artificial Intelligence and Complex Systems, Springer, New York, 2013 Search PubMed.
- D. Fourches, D. Pu and A. Tropsha, Comb. Chem. High Throughput Screening, 2011, 14, 217–225 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6ra06159a |
| This journal is © The Royal Society of Chemistry 2016 |