Synthesis, characterization and antitumor activity of novel tetrapodal 1,4-dihydropyridines: p53 induction, cell cycle arrest and low damage effect on normal cells induced by genotoxic factor H2O2†
Magda F. Mohameda,
Ahmed F. Darweeshb,
Ahmed H. M. Elwahy*b and
Ismail A. Abdelhamid*b
aChemistry Department (Biochemistry Branch), Faculty of Science, Cairo University, Giza, Egypt
bChemistry Department, Faculty of Science, Cairo University, Giza, Egypt. E-mail: aelwahy@hotmail.com; ismail_shafy@yahoo.com
First published on 15th April 2016
Abstract
Synthesis of novel tetrakis(2,6-dimethyl-4-phenyl-1,4-dihydropyridinyl)methanes 5a–d by acid-catalyzed condensation of the tetrakis-aldehydes 6a–d with eight equivalents of 3-aminobut-2-enenitrile 2 is reported. The structures of 5a–d are confirmed by different spectral tools. In vitro, cytotoxic screening assay for novel tetrapodal 1,4-dihydropyridines (5a–d) was performed on five different human cell lines (HCT116, A549, MCF7, PC3, and HEPG2). The compounds showed higher cytotoxic activity against (A549, HCT116, and MCF7) cell lines. The loss of the cytotoxic activity was observed in the case of PC3 and HEPG2 cell lines. Compound 5b showed the highest cytotoxic activity against the three lines (A549, HCT116, and MCF7). In an attempt to know the mechanism followed by the compounds to inhibit cell proliferation, compound 5b was chosen for molecular studies. Compound 5b induced apoptotic inhibition of the proliferation of human colon adenocarcinoma HCT116 cells through induction of the tumor suppressor protein p53, BAX, and through the inhibition of anti-apoptotic proteins by decreasing BCL2 gene expression using real-time PCR. Regarding cell cycle analysis, compound 5b induced G1 arrest against the three lines (MCF7, HCT116, and A549). Compound 5b has been found to reduce apoptosis of human normal melanocytes HFB4 and normal fibroblasts BHK that has been treated with genotoxic factor H2O2. Moreover, compound 5b has a potent protective effect against DNA damage, as indicated by the in vitro studying of different concentrations of 5b against two different types of healthy DNA (calf-thymus DNA and pBR322 DNA).
Introduction
The Hantzsch reaction is classified as one of the most powerful routes for the synthesis of 1,4-dihydropyridines (1,4-DHPs) that have therapeutic and pharmacological properties.1–14 A number of 1,4-dihydropyridines such as nifedipine, nicardipine, and others have proved to be some of the most important chemical classes introduced into biological sciences during the last few decades. They have dramatically improved the therapeutic standard in the treatment of coronary heart disease and for the treatment of hypertension.9,15–23 1,4-Dihydropyridines exhibit several other pharmaceutical applications including platelet antiaggregatory activity,24 neuroprotectant,25 antitumor,13 chemosensitizers in tumor therapy,26 anticonvulsant activity,27 selective adenosine-A3 receptor antagonism28 and radioprotective activity.29 In additions, they are acting as a cerebral anti-ischemic agent in the treatment of Alzheimer's disease.30 They have also been used as drugs for the treatment of a number of other diseases.1,14,24 Moreover, enamines have also been widely used as the key-intermediates to a variety of heteroaromatics.31–36 Furthermore, over recent years, there have been an increasing number of reports regarding so-called ‘multi-armed’ molecules for their wide range of applications.37–42 This class of compounds is considered important hosts for constructing microporous networks possessing selective inclusion properties and their applications in supramolecular host–guest chemistry have been recently reported.43,44 Some related compounds have also been used for the formation of discotic mesogens,45–47 organic electronic and optoelectronic materials.48–50 They have also been used as building units for dendrimers.51–53 Moreover, Grillaud and Bianco recently reported on the interesting biological activity of some diverse multivalent scaffolds.54 In conjunction with our ongoing research work on enamines33–35,55,56 and multi-armed molecules,57–62 we report herein the synthesis of novel tetrakis(1,4-dicyanodihydropyridine) derivatives to explore their activities as potential antitumor agents. In addition, molecular studies were performed on the most active compound 5b in attempts to know the mechanism of these derivatives as anticancer agents.Results and discussion
Chemistry
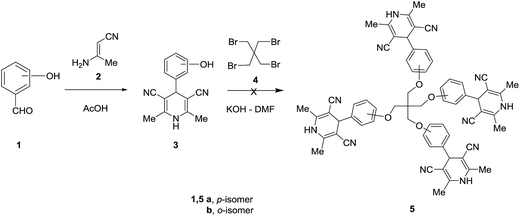
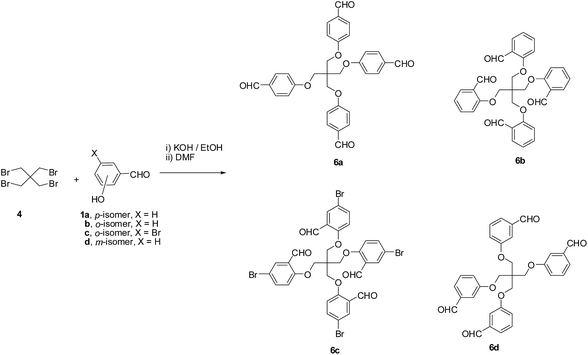
Our first attempt to synthesize the target tetrapodal dihydropyridines 5a–d includes initially the synthesis of the monopodal 4-(hydroxyphenyl)-2,6-dimethyl-1,4-dihydropyridine-3,5-dicarbonitrile 3 as precursors followed by fourfold substitution of 1,3-dibromo-2,2-bis(bromomethyl)propane 4 with four equivalents of the potassium salt of 3. Unfortunately, the reaction did not lead to the formation of 5 and gave instead a mixture of products that were difficult to separate and have not been fully characterized (Scheme 1). The synthesis of compounds 3 can be achieved by the condensation of the appropriate hydroxyaldehyde 1 with β-aminocrotonitrile 2 in refluxing acetic acid.63,64In a search for an alternative pathway to prepare the target tetrakis(2,6-dimethyl-4-phenyl-1,4-dihydropyridinylphenoxymethyl)methanes 5, our attention turned to utilize tetrakis-aldehydes 6 as precursors which could then undergo acid-catalyzed condensation with eight equivalents of β-aminocrotononitrile 2 in acetic acid at reflux. Tetrakis(4-formylphenoxymethyl)methanes 6a–d were prepared by modification of literature procedures described for the synthesis of 6a.65–67 Thus, reaction of the potassium salt of 2-, 3- or 4-hydroxybenzaldehyde (obtained upon treatment of 1a–d with ethanolic KOH) with 1,3-dibromo-2,2-bis(bromomethyl)propane 4 in DMF at reflux afforded 6a–d in good yields (Scheme 2).
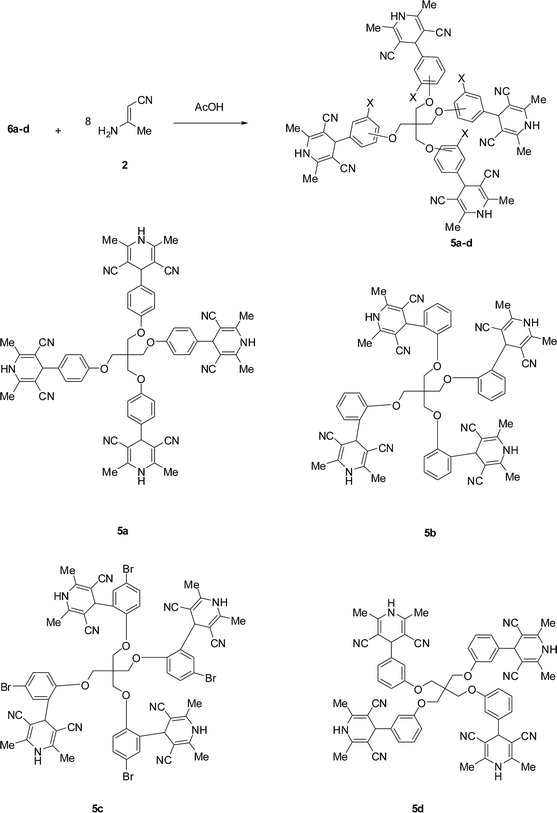
In the next step, acid-catalyzed condensation of the tetrakis-aldehydes 6a–d with eight equivalents of 3-aminobut-2-enenitrile 2 gave the corresponding tetrakis-(2,6-dimethyl-4-phenyl-1,4-dihydropyridinyl)methanes 5a–d in good to moderate yields as pale yellow crystals (Scheme 3). The structures of 5a–d were confirmed by IR, NMR, mass spectra, and elementary analysis. The symmetry of compounds 5a–d manifested as a single set of signals in the NMR spectra that were characteristic of the equivalent OCH2, CH, NH and Me groups. Thus, the 1H NMR spectrum of 5a revealed a characteristic singlet integrated by 24H at 2.01 ppm for the eight methyl groups. It also showed a singlet signal at 4.33 ppm for the pyridine-H(4). In addition, it exhibited two singlet signals characteristic for the two –OCH2 and NH groups at 4.31 and 9.43 ppm, respectively. It also featured aromatic protons as two doublets at 7.01 and 7.16 ppm, respectively. Furthermore, the 13C NMR spectrum of 5a was found to be in agreement with the proposed structure, it showed the methyl signal at 17.7 ppm and the pyridine-C(4) at 44.6 ppm. It also featured a CN signal at 119.2 ppm. All other carbon signals appeared at their expected positions.
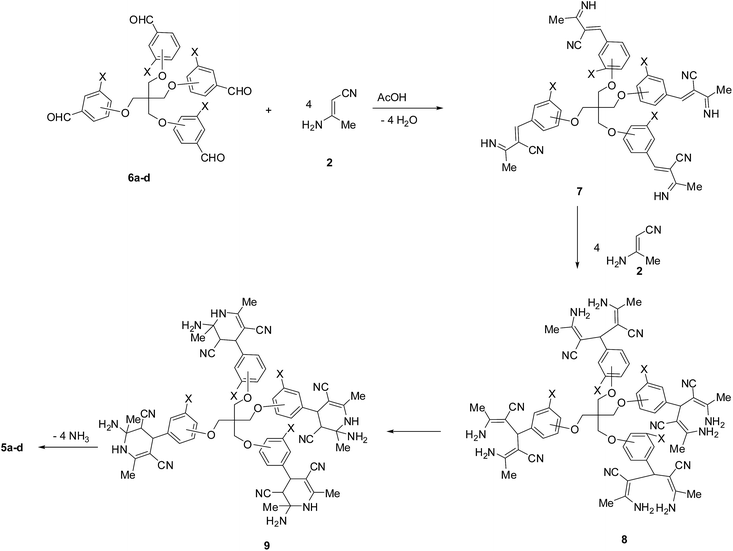
The formation of 5 may be explained by the plausible mechanism outlined in Scheme 4. The reaction involves initial condensation of the tetrakis-aldehydes 6 with four moles of the β-aminocrotonitrile 2 to yield the unstable ylidene derivative 7. The latter ylidene intermediate then reacts with another four moles of β-aminocrotonitrile 2 yielding the diamine 8, that immediately cyclizes to give the tetrahydropyridine 9. Finally, the intermediate 9 loses four moles of NH3 yielding the final isolable product 5a–d (Scheme 4).
Antitumor evaluations
An important aspect for any drug candidate to be used as a chemotherapeutic agent is that the molecule should eliminate the target tumor cells without affecting the viability of mammalian cells.68 For this purpose, in vitro anticancer screening assay for compounds 5a–d was performed on five different human cell lines (HCT116, A549, MCF7, PC3, and HEPG2) using various concentrations of tested compounds (0, 5, 15, 25 and 50 μg ml−1), and doxorubicin was used as positive standard control (Table 1). Upon exploration of the anticancer activity data (Fig. 1 and Table 1), it has been observed that the synthesized compounds are potent and promising antitumor compounds. Among the compounds tested, 5b is considered the most potent one against three cell lines (HCT116, A549, and MCF7) with the highest IC50 values (14.3, 11.2, and 12.4 μg ml−1), respectively, compared to other compounds. On the other hand, no IC50 values were recorded with PC3 and HEPG2. Morshed et al.69 assumed that some derivatives of 1,4-dihydropyridines induce non-apoptotic cell death in tumor cell lines. So, we suggest that these compounds may induce their cytotoxic effect through cell cycle arrest, p53 induction, or through reduction of angiogenic effect.| Sample | IC50 values (μg ml−1) ± SD | ||||
|---|---|---|---|---|---|
| HCT116 | A549 | MCF7 | PC3 | HEPG2 | |
| 5a | 30.2 ± 0.040 | 17.9 ± 0.049 | 11 ± 0.037 | — | — |
| 5b | 14.3 ± 0.011 | 11.2 ± 0.023 | 12.4 ± 0.029 | — | — |
| 5c | 17.8 ± 0.048 | 21.1 ± 0.033 | 24.5 ± 0.043 | — | — |
| 5d | 16.1 ± 0.042 | 18.7 ± 0.050 | 26.9 ± 0.047 | — | — |
| Doxorubicin | 8.6 ± 0.03 | 9.8 ± 0.025 | 11 ± 0.033 | 4.8 ± 0.39 | 7.7 ± 0.041 |
Real-time polymerase chain reaction (qt-PCR)
Using the real-time PCR technique and specific primers for (BAX, BCL2, and p53) genes, data on Table 2 demonstrated that compound 5b (which is the most active compound in the synthesized group) has induced apoptotic inhibition of the proliferation of human colon adenocarcinoma HCT116 cells through induction of the tumor suppressor protein p53, pro-apoptotic BAX, and finally through the inhibition of anti-apoptotic proteins by decreasing BCL2 gene expression.| Samples | BAX | BCL2 | P53 |
|---|---|---|---|
| Control (HCT116 cells) | 1.03 | 1.00 | 1.005 |
| Test (cells treated with IC50 value of compound 5b) | 2.9 | 0.48 | 4.2 |
Thus, the expression of p53, BAX (a pro-apoptotic protein) and depression of BCL2 (anti-apoptotic protein) genes relative to control (Fig. 2) promote the apoptotic death of HCT cell line. Thus, the expression of p53, BAX (a pro-apoptotic protein) and depression of BCL2 (anti-apoptotic protein) genes relative to control (Fig. 2) promote the apoptotic death of HCT cell line.
 | ||
| Fig. 2 A schematic diagram describing the relative m-RNA transcription of BAX, p53, and BCL2 genes to control. | ||
Cell cycle analysis of A549, HCT116, and MCF7 cells treated with compound 5b
Herein we are developing cell cycle-based mechanism-targeted cancer therapies that emulate the body's natural process in order to stop the growth of cancer cells using the most active compound 5b in the synthesized group. This approach can limit the side effect or the damage of the normal cells. The HCT116 and MCF7 cell lines treated with 5b at IC50 (14.3 μg ml−1 and 12.4 μg ml−1), respectively, induced G1 phase cells accumulation. G1 arrest in the two lines HCT116 and MCF7 may be related to a reduction in phosphorylation of retinoblastoma protein (pRB), regulation of G1 to S phase transition, a decrease in the levels of some of G1 specific cell cycle proteins, increased expression of p21, and inhibitory protein of CDK/cyclin complexes.70 Other studies suggested that 1,4-dihydropyridine nucleus may exert its action of G1 arrest by reducing P27 expression which is a protein that binds to cyclin and CDK blocking entry into S phase of MCF7 cell line. Recent studies also suggested that reduced levels of p27 predict a poor outcome for breast cancer patients.71 The results have shown that a reduction in S phase cells population of both HCT116 and MCF7 when treated with 5b compared to their controls, explained that the cell's nuclear chromosomes were not duplicated (Fig. 3). In addition, it was observed that there was zero cells population in M phase of MCF7 treated with 5b, and this has proved that 5b strongly prevent MCF7 cell's nucleus from the division. Unexpectedly, cell cycle analysis of A549 cell line revealed that treatment of A549 cell line with 5b (at concentration 11.2 μg ml−1, for 48 h) induced both S phase and M cells accumulation. Thus, we can suggest that 5b is a potent antiproliferative agent, as it causes G1 cell cycle arrest and growth inhibition of HCT116 and MCF7 cells.Compound 5b reduces damage effect on normal HBF4 and BHK cells treated with H2O2
The cytotoxic drugs that interfere with the mechanism of tumor cell division may produce limited harmful effects on normal tissues. In an attempt to prove that 5b reduces the probability of damage of healthy genes that are exposed to some apoptosis-associated characteristics; such as internucleosomal DNA fragmentation, necrosis, or mutagenesis, the normal cells were exposed to apoptosis by genotoxic (H2O2). The cells were then in vitro treated with compound 5b for 48 h. We have noted that the percentage of apoptosis and necrosis were reduced to a large extent compared to the controls (untreated 5b cells) as shown in (Fig. 4 and Table 3). The percentage of apoptosis was presented in the lower right quadrant and percentage of necrosis was presented in the upper right quadrant. We assumed that 5b has the ability to stimulate DNA repair, and reduce internucleosomal DNA damage of normal cells exposed to apoptosis. Our suggestion could be supported by Ryabokon et al.72| Cells | Before 5b treatment | After 5b treatment | ||
|---|---|---|---|---|
| % of apoptotic | % of necrotic cells | % of apoptotic | % of necrotic cells | |
| HFB4 | 23.9% | 19.7% | 9.2% | 16.6% |
| BHK | 24.04% | 10.9% | 7.78% | 7.7% |
Protective effect of 5b against calf-thymus DNA and plasmid pBR322 DNA damage
In vitro studies using different concentrations of 5b against two different types of DNA (CT-DNA, and plasmid pBR322 DNA), respectively, were also investigated. The degradation effect of 5b on CT-DNA and pBR322 was studied by gel electrophoresis through examination of the effect of different concentrations of 5b (10, 20, 30, and 50 mg ml−1), respectively, on both types of DNA. The results are presented in Fig. 5. Compound 5b with all tested concentrations showed no damage effect on the calf-thymus DNA. So, the direct contact of compound 5b was protective against calf-thymus DNA damage at doses ≤50 mg ml−1 (Fig. 5I). In the case of pBR322 DNA, gel electrophoresis assays showed that 5b at all tested concentrations reduced the number of DNA strand breaks. The form II (supercoiled form) of the plasmid pBR322 was not opened, which mean that the plasmid pBR322 was not degraded (Fig. 5III). Approximately, the same results were also obtained even in the presence of a strong oxidizing agent, only the nicked form of the plasmid was affected (Fig. 5II). Thus, all doses of 5b have strong protective effect against pBR322 DNA damage. Data recorded was compatible to that obtained by Morshed et al.,69 when they tested the effect of 1,4-dihydropyridines on internucleosomal DNA. This study was critically applied during radiotherapy course that induced mutation rates, where different derivative of (1,4-DHP) has been shown to reduce spontaneous damage of DNA.72Conclusion
In conclusion, we have developed a simple and efficient method for the synthesis of tetrakis-aldehydes. The synthetic utility of these compounds as building blocks for novel tetrapodal 1,4-dihydropyridines have also been investigated. These new classes of tetrapodal 1,4-dihydropyridines are interesting both in their own right as unusual molecules and for their pharmacological and biological activities. In vitro anticancer screening of compounds, 5a–d revealed higher anti-proliferative effect against human tumor cell lines (A549, HCT116, and MCF7) while no cytotoxic activity was reported against the two lines PC3 and HEPG2. Among the studied compounds, the most interesting derivative is compound 5b which is the most potent one against the human lines (HCT116, A549, and MCF7). Molecular studies proved that compound 5b is a potent antiproliferative agent, as it causes G1 cell cycle arrest and growth inhibition of HCT116 and MCF7 cells. Also, it induces the tumor suppressor proteins (p53, and BAX), and repressed anti-apoptotic protein BCL2. Moreover, it stimulates DNA repair and reduces DNA strand breaks of normal lines.Experimental
Chemistry
4-{2,2-Bis[(4-formylphenoxy)methyl]-3-(4-formylphenoxy)propoxy}benzaldehyde (6a). Pale yellow solid 77% (hexane/ethyl acetate): mp 181 °C (lit.73 mp 179–180 °C); IR ν 2742, 1691, 1600, 1578, 1508, 1467, 1427, 1392, 1312, 1248, 1214, 1158, 1048, 1029, 859, 831, 754 cm−1; 1H NMR (CDCl3) δ 4.48 (s, 8H), 7.05 (d, J = 8.6 Hz, 8H), 7.83 (d, J = 8.6 Hz, 8H), 9.89 (s, 4H) ppm; 13C NMR (CDCl3, TMS) δ 44.6, 66.4, 114.8, 130.5, 131.9, 163.1, 190.6 ppm; MS m/z 552 (M+), 307, 289, 219, 154. Anal. calcd for C33H28O8: C, 71.73; H, 5.11. Found: C, 71.82; H, 5.21.
2,2′-(2,2-Bis((2-formylphenoxy)methyl)propane-1,3-diyl)bis(oxy)dibenzaldehyde (6b). Pale yellow solid 74% (hexane/ethyl acetate), mp 152–154 °C; IR (KBr) ν 3120, 2924, 1662, 1512, 1284, 1026, 756 cm−1; 1H NMR (DMSO-d6) δ 4.32 (s, 8H, CH2), 7.08–7.13 (m, 4H), 7.27–7.36 (m, 4H), 7.63–7.72 (m, 8H), 10.41 (s, 4H, CHO); MS m/z (%) 552 (0.47, M+), 173 (21.9), 159 (27.3), 145 (40.8), 135 (100), 121 (86.9), 107 (22.5), 77 (47.5). Anal. calcd for C33H28O8: C, 71.73; H, 5.11. Found: C, 71.66; H, 5.05%.
6,6′-(2,2-Bis((4-bromo-2-formylphenoxy)methyl)propane-1,3-diyl)bis(oxy)bis(3 bromobenzaldehyde) (6c). Pale yellow solid 72% (dioxane), mp 212–214 °C; IR (KBr) ν 2954, 2850, 1681, 1589, 1392, 1234, 1180, 1122, 817 cm−1; 1H NMR (DMSO-d6) δ 4.63 (s, 8H, CH2), 7.31 (d, 4H, J = 8.7 Hz), 7.69 (s, 4H), 7.77 (d, 4H, J = 8.7 Hz), 10.28 (s, 4H, CHO); MS m/z (%) 868 (1.5, M+), 450 (5.1), 314 (5.8), 265 (28.0), 212 (100), 158 (69.5), 145 (71.7), 115 (29.3), 76 (27.5). Anal. calcd for C33H24Br4O8: C, 45.65; H, 2.79. Found: C, 45.45; H, 2.61%.
3,3′-(2,2-Bis((3-formylphenoxy)methyl)propane-1,3-diyl)bis(oxy)dibenzaldehyde (6d). Pale yellow powder (hexane/benzene); mp 151–153 °C; IR (KBr) ν 3062, 2943, 1693, 1597, 1261 cm−1; 1H NMR (DMSO-d6) δ 4.44 (s, 8H, CH2), 7.30–7.49 (m, 16H), 9.93 (s, 4H, CHO); MS m/z (%) 552 (7.3, M+), 370 (15.9), 235 (100), 135 (7.4), 107 (10.2), 43 (8.4). Anal. calcd for C33H28O8: C, 71.73; H, 5.11. Found: C, 71.57; H, 5.34%.
4,4′-(4,4′-(2,2-Bis((4-(3,5-dicyano-2,6-dimethyl-1,4-dihydropyridin-4-yl)phenoxy)methyl)-propane-1,3-diyl)bis(oxy)bis(4,1-phenylene))bis(2,6-dimethyl-1,4-dihydropyridine-3,5-dicarbonitrile) (5a). Pale yellow powder; mp >300 °C; IR (KBr) ν 3313, 2927, 2198, 1508, 1230, 1029, 852, 621 cm−1; 1H NMR (DMSO-d6) δ 2.01 (s, 24H, CH3), 4.31 (s, 8H, CH2), 4.33 (s, 4H, pyridine-4H), 7.01 (d, 8H, J = 7.5 Hz), 7.16 (d, 8H, J = 7.8 Hz), 9.43 (s, 4H, NH); 13C NMR (DMSO-d6) δ 17.7, 40.2, 44.6, 66.7, 82.9, 114.9, 119.2, 128.7, 136.7, 146.3, 158.0; MS m/z (%) 1069 (2.28, M+), 724 (7.9), 476 (11.9), 314 (53.3), 262 (92.1), 95 (29.2), 64 (100). Anal. calcd for C65H56N12O4: C, 73.02; H, 5.28; N, 15.72. Found: C, 73.18; H, 5.15; N, 15.84%.
4,4′-(2,2′-(2,2-Bis((2-(3,5-dicyano-2,6-dimethyl-1,4-dihydropyridin-4-yl)phenoxy)methyl)-propane-1,3-diyl)bis(oxy)bis(2,1-phenylene))bis(2,6-dimethyl-1,4-dihydropyridine-3,5-dicarbonitrile) (5b). Pale yellow powder; mp 284–286 °C; IR (KBr) ν 3456, 2927, 2198, 1508, 1288, 1234, 1029, 756, 632 cm−1; 1H NMR (DMSO-d6) δ 2.03 (s, 24H, CH3), 4.20 (s, 8H, CH2), 4.94 (s, 4H, pyridine-4H), 7.02–7.10 (m, 8H), 7.23–7.30 (m, 8H), 9.42 (s, 4H, NH); 13C NMR (DMSO-d6) δ 17.7, 40.8, 44.3, 66.7, 82.6, 113.8, 114.3, 119.2, 120.5, 129.9, 145.6, 146.7, 158.8; MS m/z (%) 1069 (0.79, M+), 726 (18.2), 475 (10.6), 314 (40.0), 250 (100), 158 (38.3), 80 (56.8). Anal. calcd for C65H56N12O4: C, 73.02; H, 5.28; N, 15.72. Found: C, 72.89; H, 5.40; N, 15.64%.
4,4′-(6,6′-(2,2-Bis((4-bromo-2-(3,5-dicyano-2,6-dimethyl-1,4-dihydropyridin-4-yl)phenoxy)-methyl)propane-1,3-diyl)bis(oxy)bis(3-bromo-6,1-phenylene))bis(2,6-dimethyl-1,4-dihydropyridine-3,5-dicarbonitrile) (5c). Pale yellow powder; mp 268–270 °C; IR (KBr) ν 3464, 2924, 2202, 1512, 1280, 1107, 1029, 813, 601 cm−1; 1H NMR (DMSO-d6) δ 2.01 (s, 24H, CH3), 4.48 (s, 8H, CH2), 5.01 (s, 4H, pyridine-4H), 7.09 (d, 4H, J = 8.7 Hz), 7.31–7.39 (m, 8H), 9.46 (s, 4H, NH); 13C NMR (DMSO-d6) δ 17.8, 21.1, 44.6, 68.4, 81.9, 113.0, 115.2, 119.1, 131.6, 132.2, 134.5, 147.2, 154.4. Anal. calcd for C65H52Br4N12O4: C, 56.38; H, 3.78; N, 12.14. Found: C, 56.16; H, 3.58; N, 11.86%.
4,4′-(3,3′-(2,2-Bis((3-(3,5-dicyano-2,6-dimethyl-1,4-dihydropyridin-4-yl)phenoxy)methyl)-propane-1,3-diyl)bis(oxy)bis(3,1-phenylene))bis(2,6-dimethyl-1,4-dihydropyridine-3,5-dicarbonitrile) (5d). Pale yellow powder; mp 214–216 °C; IR (KBr) ν 3464, 2927, 2202, 1508, 1276 cm−1; 1H NMR (DMSO-d6) δ 2.04 (s, 24H, CH3), 4.37 (s, 8H, CH2), 4.39 (s, 4H, pyridine-4H), 6.85–6.98 (m, 12H), 7.27–7.32 (m, 4H), 9.46 (s, 4H, NH); 13C NMR (DMSO-d6) δ 17.7, 40.8, 44.3, 66.7, 82.6, 113.8, 114.3, 119.2, 120.5, 129.9, 145.6, 146.7, 158.8; MS m/z (%) 1069 (10.6, M+), 992 (73.0), 611 (10.5), 481 (21.9), 249 (100), 57 (11.8). Anal. calcd for C65H56N12O4: C, 73.02; H, 5.28; N, 15.72. Found: C, 73.32; H, 5.01; N, 15.55%.
Biology
| Genes | Primer sequence |
|---|---|
| BAX | Forward: 5′-GGCCCACCAGCTCTGAGCAGA-3′ |
| Reverse: 5′-GCCACGTGGGCGTCCCAAAGT-3′ | |
| BCL2 | Forward: 5′-ACAACATCG CCCTGTGGATGAC-3′ |
| Reverse: 5′-ATAGCTGATTCGACGTTTTGCC-3′ | |
| P53 | Forward: 5′-GTTCCGAGAGCTGAATGAGG-3′ |
| Reverse: 5′-TTATGGCGGGAGGTCGACTG-3′ |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 nuclei. Cell cycle analysis was performed with the Epics XL coulter software package (Beckman Coulter Crop., U.S.A.).
000 nuclei. Cell cycle analysis was performed with the Epics XL coulter software package (Beckman Coulter Crop., U.S.A.).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10)) was added to samples and process by flow cytometry within 1 h for maximal signal, where cells were washed twice with phosphate-buffered saline (PBS) and analysed under a fluorescence microscope. Cells exhibiting green fluorescence on the plasma-membranes were counted and estimated as a percentage of total cells.
10)) was added to samples and process by flow cytometry within 1 h for maximal signal, where cells were washed twice with phosphate-buffered saline (PBS) and analysed under a fluorescence microscope. Cells exhibiting green fluorescence on the plasma-membranes were counted and estimated as a percentage of total cells.References
- M. E. Letelier, P. Entrala, C. López-Alarcón, V. González-Lira, A. Molina-Berríos, J. Cortés-Troncoso, J. Jara-Sandoval, P. Santander and L. Núñez-Vergara, Toxicol. In Vitro, 2007, 21, 1610–1618 CrossRef CAS PubMed.
- T. Shamim, M. Gupta and S. Paul, J. Mol. Catal. A: Chem., 2009, 302, 15–19 CrossRef CAS.
- G. Swarnalatha and G. Prasanthi, Int. J. ChemTech Res., 2011, 3, 75–89 CAS.
- R. Sridhar and P. T. Perumal, Tetrahedron, 2005, 61, 2465–2470 CrossRef CAS.
- S. Paul, S. Sharma, M. Gupta, D. Choudhary and R. Gupta, Bull. Korean Chem. Soc., 2007, 28, 336–338 CrossRef CAS.
- F. Saikh, R. De and S. Ghosh, Tetrahedron Lett., 2014, 55, 6171–6174 CrossRef CAS.
- C. Zapata-Urzúa, M. Pérez-Ortiz, G. A. Acosta, J. Mendoza, L. Yedra, S. Estradé, A. Álvarez-Lueje, L. J. Núñez-Vergara, F. Albericio, R. Lavilla and M. J. Kogan, J. Colloid Interface Sci., 2015, 453, 260–269 CrossRef PubMed.
- V. Sridharan, P. T. Perumal, C. Avendaño and J. C. Menéndez, Tetrahedron, 2007, 63, 4407–4413 CrossRef CAS.
- L. Navidpour, H. Shafaroodi, R. Miri, A. R. Dehpour and A. Shafiee, Farmaco, 2004, 59, 261–269 CrossRef CAS PubMed.
- A. Debache, W. Ghalem, R. Boulcina, A. Belfaitah, S. Rhouati and B. Carboni, Tetrahedron Lett., 2009, 50, 5248–5250 CrossRef CAS.
- M. Filipan-Litvić, M. Litvić and V. Vinković, Tetrahedron, 2008, 64, 5649–5656 CrossRef.
- R. Miri, K. Javidnia, B. Hemmateenejad, M. Tabarzad and M. Jafarpour, Chem. Biol. Drug Des., 2009, 73, 225–235 CAS.
- H.-A. S. Abbas, W. A. El Sayed and N. M. Fathy, Eur. J. Med. Chem., 2010, 45, 973–982 CrossRef CAS PubMed.
- J. Marco-Contelles, R. León, C. de Los Ríos, A. Guglietta, J. Terencio, M. G. López, A. G. García and M. Villarroya, J. Med. Chem., 2006, 49, 7607–7610 CrossRef CAS PubMed.
- C. O. Kappe, Eur. J. Med. Chem., 2000, 35, 1043–1052 CrossRef CAS PubMed.
- D. Rampe and J. Michael Kane, Drug Dev. Res., 1994, 33, 344–363 CrossRef CAS.
- B. Loev, M. M. Goodman, K. M. Snader, R. Tedeschi and E. Macko, J. Med. Chem., 1974, 17, 956–965 CrossRef CAS PubMed.
- F. Bossert, H. Meyer and E. Wehinger, Angew. Chem., Int. Ed. Engl., 1981, 20, 762–769 CrossRef.
- T. Iijima, T. Yanagisawa and N. Taira, J. Mol. Cell. Cardiol., 1984, 16, 1173–1177 CrossRef CAS PubMed.
- S. Bohlooli, M. Mahmoudian, G. G. Skellern, M. H. Grant and J. N. A. Tettey, J. Pharm. Pharmacol., 2004, 56, 1469–1475 CrossRef CAS PubMed.
- S. Subramani, C. Vijayanand and E. Tharion, Br. J. Pharmacol., 2002, 137, 756–760 CrossRef CAS PubMed.
- J. L. Reid, P. A. Meredith and F. Pasanisi, J. Cardiovasc. Pharmacol., 1985, 7(suppl. 4), S18–S20 CrossRef PubMed.
- J. D. Wallin, M. E. Cook, L. Blanski, G. S. Bienvenu, G. G. Clifton, H. Langford, P. Turlapaty and A. Laddu, Am. J. Med., 1988, 85, 331–338 CrossRef CAS PubMed.
- R. G. Bretzel, C. C. Bollen, E. Maeser and K. F. Federlin, Am. J. Kidney Dis., 1993, 21, S53–S64 CrossRef.
- V. Klusa, Drugs Future, 1995, 20, 135–138 CrossRef.
- R. Boer and V. Gekeler, Drugs Future, 1995, 20, 499 Search PubMed.
- J. M. Tusell, S. Barrón and J. Serratosa, Brain Res., 1993, 622, 99–104 CrossRef CAS PubMed.
- J. L. Jiang, A. M. van Rhee, N. Melman, X. D. Ji and K. A. Jacobson, J. Med. Chem., 1996, 39, 4667–4675 CrossRef CAS PubMed.
- I. Donkor, Bioorg. Med. Chem., 1998, 6, 563–568 CrossRef CAS PubMed.
- R. Bretzel, C. Bollen, E. Maeser and K. Federlin, Drugs Future, 1992, 17, 465 CrossRef.
- H. M. E. Hassaneen, H. M. Hassaneen and M. H. Elnagdi, Z. Naturforsch., B: J. Chem. Sci., 2004, 59, 1132–1136 CAS.
- S. A. S. Ghozlan, I. A. Abdelhamid, H. Gaber and M. H. Elnagdi, J. Chem. Res., 2004, 789–793 CrossRef CAS.
- S. A. S. Ghozlan, I. A. Abdelhamid, H. M. Hassaneen and M. H. Elnagdi, J. Heterocycl. Chem., 2007, 44, 105–108 CrossRef CAS.
- S. A. S. Ghozlan, I. A. Abdelhamid, H. M. Hassaneen and M. H. Elnagdi, J. Heterocycl. Chem., 2007, 44, 105–108 CrossRef CAS.
- S. M. Riyadh, I. A. Abdelhamid, H. M. Al-Matar, N. M. Hilmy and M. H. Elnagdi, Heterocycles, 2008, 75, 1849–1905 CrossRef CAS.
- S. A. S. Ghozlan, A. G. Ahmed and I. A. Abdelhamid, J. Heterocycl. Chem., 2015 DOI:10.1002/jhet.2341.
- F. M. Menger and V. A. Azov, J. Am. Chem. Soc., 2002, 124, 11159–11166 CrossRef CAS PubMed.
- M. Gingras, A. Pinchart and C. Dallaire, Angew. Chem., Int. Ed., 1998, 37, 3149–3151 CrossRef CAS.
- M. AL-Smadi and S. Mohammad, J. Heterocycl. Chem., 2009, 46, 201–206 CrossRef CAS.
- S. Achelle, N. Plé, D. Kreher, A.-J. Attias, I. Arfaoui and F. Charra, Tetrahedron Lett., 2009, 50, 7055–7058 CrossRef CAS.
- A. H. M. Elwahy, Tetrahedron Lett., 2001, 42, 5123–5126 CrossRef CAS.
- H.-J. van Manen, R. H. Fokkens, F. C. J. M. van Veggel and D. N. Reinhoudt, Eur. J. Org. Chem., 2002, 2002, 3189–3197 CrossRef.
- J. W. Steed, D. R. Turner and K. Wallace, Core Concepts in Supramolecular Chemistry and Nanochemistry, John Wiley & Sons, 2007 Search PubMed.
- J. W. Steed and J. L. Atwood, Supramolecular Chemistry, John Wiley & Sons, 2013 Search PubMed.
- E. M. García-Frutos, A. Omenat, J. Barberá, J. L. Serrano and B. Gómez-Lor, J. Mater. Chem., 2011, 21, 6831 RSC.
- B. Gómez-Lor, B. Alonso, A. Omenat and J. L. Serrano, Chem. Commun., 2006, 5012–5014 RSC.
- M. Talarico, R. Termine, E. M. García-Frutos, A. Omenat, J. L. Serrano, B. Gómez-Lor and A. Golemme, Chem. Mater., 2008, 20, 6589–6591 CrossRef CAS.
- B. Traber, J. J. Wolff, F. Rominger, T. Oeser, R. Gleiter, M. Goebel and R. Wortmann, Chemistry, 2004, 10, 1227–1238 CrossRef CAS PubMed.
- T. Shoji, J. Higashi, S. Ito, T. Okujima, M. Yasunami and N. Morita, Chemistry, 2011, 17, 5116–5129 CrossRef CAS PubMed.
- F.-K. Bruder, R. Hagen, T. Rölle, M.-S. Weiser and T. Fäcke, Angew. Chem., Int. Ed. Engl., 2011, 50, 4552–4573 CrossRef CAS PubMed.
- D. Astruc, E. Boisselier and C. Ornelas, Chem. Rev., 2010, 110, 1857–1959 CrossRef CAS PubMed.
- B. K. Nanjwade, H. M. Bechra, G. K. Derkar, F. V Manvi and V. K. Nanjwade, Eur. J. Pharm. Sci., 2009, 38, 185–196 CrossRef CAS PubMed.
- P. Rajakumar and S. Raja, Tetrahedron Lett., 2008, 49, 6539–6542 CrossRef CAS.
- M. Grillaud and A. Bianco, J. Pept. Sci., 2015, 21, 330–345 CrossRef CAS PubMed.
- I. A. Abdelhamid, S. A. S. Ghozlan, H. Kolshorn, H. Meier and M. H. Elnagdi, Heterocycles, 2007, 71, 2627–2637 CrossRef CAS.
- S. A. S. Ghozlan, I. A. Abdelhamid, M. H. Elnagdi and H. M. Gaber, J. Heterocycl. Chem., 2005, 42, 1185–1189 CrossRef CAS.
- M. R. Shaaban and A. H. M. Elwahy, J. Heterocycl. Chem., 2012, 49, 640–645 CrossRef CAS.
- A. H. M. Elwahy and M. R. Shaaban, Curr. Org. Synth., 2015, 10, 425–466 Search PubMed.
- A. H. M. Elwahy, A. A. Abbas and Y. A. Ibrahim, J. Chem. Res., 1998, 184–185 RSC , (M) 901–914.
- O. M. Sayed, A. E. M. Mekky, A. M. Farag and A. H. M. Elwahy, J. Heterocycl. Chem., 2015 DOI:10.1002/jhet.2373.
- O. M. Sayed, A. E. M. Mekky, A. M. Farag and A. H. M. Elwahy, J. Sulfur Chem., 2014, 36, 124–134 CrossRef.
- I. A. Abdelhamid, A. F. Darweesh and A. H. M. Elwahy, Tetrahedron Lett., 2015, 56, 7085–7088 CrossRef CAS.
- J. Kuthan, A. Kurfürst, Z. Prošek and J. Paleček, Collect. Czech. Chem. Commun., 1978, 43, 1068–1079 CrossRef CAS.
- E. Mohr, J. Prakt. Chem., 1897, 56, 124–142 CrossRef CAS.
- R. Banerjee, S. Maiti and D. Dhara, Green Chem., 2014, 16, 1365–1373 RSC.
- D. Laliberté, T. Maris and J. D. Wuest, J. Org. Chem., 2004, 69, 1776–1787 CrossRef PubMed.
- V. Kungurtsev, J. Laakkonen, A. G. Molina and P. Virta, Eur. J. Org. Chem., 2013, 2013, 6687–6693 CrossRef CAS.
- A. Sonawnae, Int. J. Nanomed., 2012, 7, 1805 CrossRef PubMed.
- S. R. M. D. Morshed, K. Hashimoto, Y. Murotani, M. Kawase, A. Shah, K. Satoh, H. Kikuchi, H. Nishikawa, J. Maki and H. Sakagami, Anticancer Res., 2005, 25, 2033–2038 CAS.
- J. Yoshida, T. Ishibashi and M. Nishio, Biochem. Pharmacol., 2007, 73, 943–953 CrossRef CAS PubMed.
- P. S. Steeg and J. S. Abrams, Nat. Med., 1997, 3, 152–154 CrossRef CAS PubMed.
- N. I. Ryabokon, R. I. Goncharova, G. Duburs and J. Rzeszowska-Wolny, Mutat. Res., 2005, 587, 52–58 CAS.
- F.-T. Luo and C.-H. Chen, Heterocycles, 2001, 55, 1663–1678 CrossRef CAS.
- S. A. S. Ghozlan, M. F. Mohamed, A. G. Ahmed, S. A. Shouman, Y. M. Attia and I. A. Abdelhamid, Arch. Pharm., 2015, 348, 113–124 CrossRef CAS PubMed.
- M. W. Pfaffl, Nucleic Acids Res., 2001, 29, 45e–45 CrossRef.
- T.-M. Cao, F.-Y. Hua, C.-M. Xu, B.-S. Han, H. Dong, L. Zuo, X. Wang, Y. Yang, H.-Z. Pan and Z.-N. Zhang, Ann. Hematol., 2006, 85, 434–442 CrossRef CAS PubMed.
- M. F. Mohamed, M. S. Mohamed, M. M. Fathi, S. A. Shouman and I. A. Abdelhamid, Anti-Cancer Agents Med. Chem., 2014, 14, 1282–1292 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6ra04974e |
| This journal is © The Royal Society of Chemistry 2016 |