DNAzyme catalytic beacons-based a label-free biosensor for copper using electrochemical impedance spectroscopy†
Wenyong Huab,
Xiaobo Min*ab,
Xinyu Liab,
Shengxiang Yangab,
Langbo Yiab and
Liyuan Chaiab
aSchool of Metallurgy and Environment, Central South University, Changsha, 410083, China. E-mail: huwenyong07@126.com; mxb@csu.edu.cn; Fax: +86 0731 88710171; Tel: +86 13762157065
bNational Engineering Research Center for Pollution Control of Heavy Metals, Changsha 410083, Hunan, PR China
First published on 22nd December 2015
Abstract
In this study, we developed a novel selective method for copper quantification based on gold nanoclusters (GNCs) and DNAzyme. The GNCs were used as the sensing interface to immobilize with the DNAzyme capturing Cu2+ ions. The DNAzyme could be activated to cleave the substrate strand into two DNA fragments in the presence of Cu2+, and produce changes in the interfacial properties of the electrode. The difference in the interfacial electron-transfer resistance was probed in the presence of the reversible redox couple, Fe(CN)63−/4−, as a marker using electrochemical impedance spectroscopy (EIS). A Randles equivalent circuit was employed to evaluate the EIS results. The charge transfer resistance (RCT) of the Fe(CN)63−/4− redox indicator decreased remarkably after hybridization with Cu2+. The difference in RCT values before and after hybridization with Cu2+ showed a linear relationship with the concentration of Cu2+ in a range of 0.1–400 nM, with a detection limit of 0.0725 nM (S/N = 3). Furthermore, with the application of Cu2+ dependent DNAzyme, the proposed sensing system exhibited high selectivity. This biosensor demonstrated promising potential for Cu2+ detection in real samples.
1. Introduction
Copper is used widely but can leak into the natural environment through various routes. At low concentrations, copper is an essential nutrient. However, exposure to high levels of copper, even for a short period of time, can cause gastrointestinal disturbances, whereas long term exposure causes liver or kidney damage.1,2 Therefore, the development of a sensitive, selective and comparatively inexpensive method for Cu2+ ion detection is of great significance. Traditional quantitative methods, for example, atomic absorption spectrometry (AAS)3 and inductively coupled plasma mass spectroscopy (ICP/MS),4 provide reliable and accurate results, but require expensive and sophisticated instrumentation and complicated sample pretreatment. Many electrochemical sensors, such as voltammetric5,6 and electrochemiluminescence sensors,7,8 are simple and relatively inexpensive; however, they suffer from low reproducibility and low selectivity. Therefore, metal–biochemical and biophysical studies have attracted intense attention over the previous decades. Important insights have been gained regarding metal–DNAzymes sensors, the reason may be that DNAzyme-based biosensors have higher detection stability, sensitivity, and selectivity for a given metal reaction. On the contrary, they are insensitive to other metal ions even in complex environments, thus improving the potential applications in a real environment.9,10Furthermore, obtaining an effective immobilization platform for thiolated probe DNA and DNAzymes on the modified electrode is a key technique for improving DNA sensing efficiencies. In recent years, various nanomaterials were employed as DNA immobilization substrates and recognition elements in biosensors. For example, Zhong and co-workers fabricated a simple but sensitive turn-off assay for Pb2+ detection using the 8–17 DNAzyme based on single-walled carbon nanotubes (SWCNT).11 Tang et al.9 used ordered mesoporous carbon–gold nanoparticles (OMC–GNPs) as the platform for electrochemical biosensors for the detection of Pb2+ by electrochemical impedance spectroscopy (EIS) with Fe(CN)64−/3− as the redox couple. These biosensors could increase the sensitivity and lower the detection limit for metal ions detection. In this study, gold nanoclusters (GNCs) were used as the sensing interface to immobilize the DNA. In addition to its higher conductivity, excellent structural continuity and general biocompatibility,12 GNCs also provide a natural platform for stable DNA immobilization because of the strong gold–sulfur (Au–S) covalent-type interactions, which might extend the life and stability of the biosensor, and make the sensor assembly process easier. Although gold nanoclusters have been used in biosensors,12,13 little attention has been paid to copper ion sensors based on DNAzyme-based biosensor.
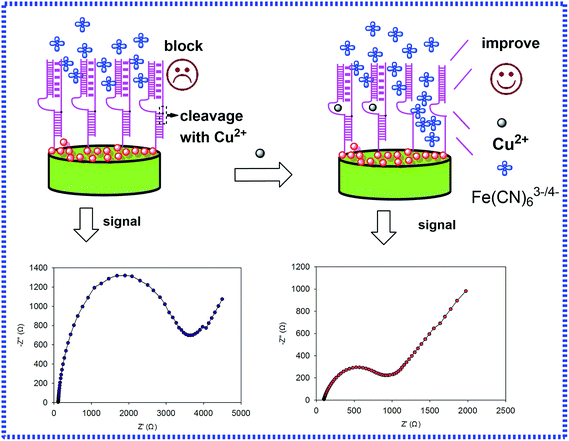
Herein, a label-free biosensor with GNCs as a transducer platform with DNAzyme was developed for the detection of copper using EIS with selectivity and sensitivity. The interfacial properties of the electrode, such as electron transfer resistance and capacitance, were investigated in the presence of a redox probe of Fe(CN)63−/4−. In the presence of Cu2+, the trans-acting catalytic beacon cleaves the sessile of the substrate into two fragments (Fig. 1), resulting in a remarkable decrease of interfacial charge-transfer resistance (RCT) for the negatively charged redox probe at the electrochemical biosensor due to the enhanced electron transfer by GNCs. Taking advantage of the RCT change, Cu2+, can be detected at concentrations as low as 0.0725 nM.
2. Experimental
2.1. Chemicals and apparatus
Copper(II) chloride was purchased from Sigma-Aldrich (USA). K3Fe(CN)6, K4Fe(CN)6, sodium ascorbate and all other chemicals were of analytical grade and used as received. All aqueous solutions were prepared using ultra-pure water (18 MΩ cm, Milli-Q, Millipore). 50 mM tris–acetate buffer (pH 7.4), containing 0.2 M NaCl and phosphate buffer saline (PBS, 0.1 M KH2PO4 and 0.1 M Na2HPO4), were used in this study. The DNA target–specific probes used for hybridization in the experiment were synthesized by Sangon (Shanghai, China) and purified by high-performance liquid chromatography. The sequences of the oligonucleotides include the following sequences:5′-HS-(CH2)6-GGTAAGCCTGGGCCTCTTTCTTTTTAAGAAAGAAC-3′ (DNA S1)
5′-AGCTTCTTTCTAATACGGCTTACC-3′ (DNA S2, a complementary substrate oligonucleotide of the DNAzyme)
CHI1230B A14535 electrochemical workstation (Chenhua Instrument, Shanghai, China) was used for electrochemical impedance spectroscopy (EIS) and cyclic voltammetry (CV) measurements. In addition, in this study, the three-electrode system includes a saturated calomel electrode (SCE) as the reference electrode, a Pt foil auxiliary electrode and a modified electrode as the working electrode. Using Agilent, 7700×, all the study was conducted at room temperature (25 °C) unless mentioned otherwise.
2.2. Sensor fabrication
Probe was activated by 2 mM TCEP, which is included to reduce disulfide bonded oligomer, and diluted by 50 mM tris–acetate buffer (pH 7.4). The bare glass carbon electrode (GCE) was polished first in alumina slurry, and then rinsed with deionized water. Finally, the electrode surface was treated using H2SO4 (0.5 M) with a cyclic voltammetry scan (between 0 and 1.2 V at a scan rate of 50 mV s−1) until a reproducible scan was obtained. After drying, Au nanoprickle clusters were electrodeposited onto the GCE.14 Electrodeposition was performed using chronoamperometry in a 1% (w/w) HAuCl4 solution containing perchloric acid at a potential of 0.18 V for 120 s. After immersion in chloroform to dissolve the polycarbonate template, the obtained electrode was rinsed thoroughly with water.Subsequently, the solution (7 μL DNA S1, 50 mM tris–acetate buffer (pH 7.4)) was dropped onto the electrode surface for self-assembling through Au–S bonding for 12 h at 4 °C. The probes for this biosensor were hybridized as follows: 6-mercapto-1-hexanol (MCH) solution (400 μL) was used to immerse the modified electrode with DNA S1 probes for 30 min to improve the stability and quality, to reduce nonspecific adsorption of DNA and to obtain a well aligned DNA monolayer. Subsequently, the modified electrode was soaked in the solution with DNA S2, and incubated at room temperature for 70 min to yield the final copper–DNAzyme assembly on the surface. The electrode was immersed in buffer (50 mM tris–acetate, 0.2 M NaCl, pH 7.4) for 10 min to reduce the nonspecific adsorption of DNA S2. Various concentrations of target Cu2+ in the buffer (50 mM tris–acetate, 0.2 M sodium ascorbate, pH 7.4) was then allowed to react with the DNA surface (50 min in a 40 °C water bath) to obtain the maximum cleavage of the substrate strand on the modified electrode. The electrode was then removed from the buffer, and allowed to cool to room temperature within 1 hour.
2.3. Impedimetric detection
A CHI1230B A14535 electrochemical workstation was used, and all the measurements were carried out at room temperature using conventional three-electrode system. The modified electrode was treated with various concentrations of Cu2+ in buffers (50 mM tris–acetate, 0.2 M NaCl, pH 7.4) for 2 h. Subsequently, it was washed with tris–acetate buffer (pH = 7.4). A conventional three-electrode system was used. Cyclic voltammetry (CV) was performed in 0.1 M PBS (pH 7.0) containing 10 mM KCl and 5 mM Fe(CN)63−/4− (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Moreover, EIS was performed in 0.1 M PBS (pH 7.4) containing 5 mM Fe(CN)63−/4− (1
1). Moreover, EIS was performed in 0.1 M PBS (pH 7.4) containing 5 mM Fe(CN)63−/4− (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and 10 mM KCl in the frequency range, 0.1 Hz to 100 kHz, with 5 mV as the amplitude at a polarization potential of 0.18 V. The data for the condition optimization and the calibration curve was the average of three measurements.
1) and 10 mM KCl in the frequency range, 0.1 Hz to 100 kHz, with 5 mV as the amplitude at a polarization potential of 0.18 V. The data for the condition optimization and the calibration curve was the average of three measurements.
The impedance spectra were plotted in the form of a complex plane diagram (−Z′′vs. Z′), and fitted to a theoretical curve corresponding to the equivalent circuit by software of EIS Spectrum Analyser. The interfacial resistance (RCT) was obtained. This could presumably be due to changes in the film thickness of the films of DNA S1 and DNA S2, which would increase the distance for electron transfer through the film and hence increase RCT. Importantly, the trans-acting catalytic strand cleaved the sessile of the substrate into two fragments in the presence of Cu2+, and the charge-transfer resistance RCT was decreased significantly. The reason may be that the DNA S2 would introduce significant disorder into the solution, and the redox probe may penetrate the film to a larger extent, giving rise to a lower RCT. To compare the results obtained from the electrodes used with or without Cu2+ and to obtain the relative signals. The ΔRCT value was defined according to the following equations:
| ΔRCT = RCT(DNAzyme) − RCT(DNAzyme+Cu) |
2.4. Analysis of environmental samples
As a further step, we attempted to provide the general applicability of this biosensor to practical samples. Four water samples were collected from Xiangjiang River, Hunan province. After filtering through a 0.2 mM membrane to remove the oils and other organic impurities, the samples were spiked with standard solutions of Cu2+ prior to the measurement using the proposed method and ICP/MS (Agilent 7700×, Agilent, USA).3. Results and discussion
3.1. DNAzyme-based electrochemical sensor
As presented in Fig. 1, the GNCs film was utilized as the platform for the immobilization of DNA and enhancing the transfer of electronics. Once in the presence of Cu2+, the trans-acting catalytic strand cleaved the sessile phosphodiester of the substrate into two fragments, leading to a change in the interfacial charge-transfer resistance of the electrodes towards the Fe(CN)64−/3− redox couple. As shown in Fig. 1, after hybridization with the probes, the impedance was relatively high, and the impedance shifted back in the presence of Cu2+. The reason might be the easier electron transfer between the GCE surface and Fe(CN)63−/4− with Cu2+. Thus, the electron transfer of the whole system could be accelerated. In fact, the changes in the charge transfer resistance (ΔRCT) of DNA films in the presence and absence of the metal ion were different and dependent on the concentration of the given metal ion. The result also coincided with the presumptive mechanism of the Cu2+ induced conformational change of the hybridization probes, which enhanced the electron transfer of the Fe(CN)64−/3− redox couple. The various conformational characteristics and charge of DNA on the surface resulted in different charge-transfer resistances for the redox indicator ions. Based on this principle, the interaction between DNA and Cu2+ led to a decrease in RCT, and ΔRCT was related to the concentration of Cu2+.3.2. Characterization of the electrode and optimization of the variables of the experimental conditions
As observed in Fig. S-1A,† the gold nanoparticles were electrodeposited onto the surface of GCE, and protuberant clusters grew along the pores of the polycarbonate template. The mean diameter of the Au nanoclusters was around 100 nm. In addition, to test the performance of the modified electrode, CV was carried out in phosphate buffer (containing 5 mM Fe(CN)63−/4− (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and 10 mM KCl, pH 7.4). As shown in Fig. S-1B,† the peak current of the redox probe was increased significantly after the immobilization of GNCs on the GCE. These cyclic voltammograms also proved that the electrode had a good current response capability. Correspondingly, EIS revealed the impedance of the GNCs/GCE and bare GCE in phosphate buffer. An almost straight line was observed with the assembled GNCs; however, an obvious increase in the interfacial resistance was observed from the GCE (Fig. S-1C†), which indicated that the introduction of GNCs could enhance the electron transfer kinetics to a large extent. In addition, the electron transfer ability of the modified electrode reflected by EIS was in accordance with the current density response reflected by CV.
1) and 10 mM KCl, pH 7.4). As shown in Fig. S-1B,† the peak current of the redox probe was increased significantly after the immobilization of GNCs on the GCE. These cyclic voltammograms also proved that the electrode had a good current response capability. Correspondingly, EIS revealed the impedance of the GNCs/GCE and bare GCE in phosphate buffer. An almost straight line was observed with the assembled GNCs; however, an obvious increase in the interfacial resistance was observed from the GCE (Fig. S-1C†), which indicated that the introduction of GNCs could enhance the electron transfer kinetics to a large extent. In addition, the electron transfer ability of the modified electrode reflected by EIS was in accordance with the current density response reflected by CV.
The experimental conditions were optimized before the quantitative analysis of Cu2+. Fig. 2A shows the effects of self-assembly time of the capture probe (DNA S1) on the modified electrode surface. With the self-assembly time, the ΔRCT also increased, and reached a plateau at 10 h. Therefore, in subsequent measurements, a self-assembly time of 10 h was used. Similarly, optimization of the hybridization conditions includes the time of the DNAzyme hybridization (DNA S2) reaction. The hybridization time is an important factor to ensure the adequacy of a contact reaction. As shown in Fig. 2B, the response current increased sharply with increasing hybridization time from 30 to 70 min, and then leveled off.
The reaction time between DNA with Cu2+ would have a profound effect on the performance of the biosensor. Upon exposure to 200 nM Cu2+, the ΔRCT increased within 50 min of incubation and then remained constant when the action time was increased further (Fig. 2C). Therefore, an incubation time of 50 min was used as the optimal reaction time between DNA with Cu2+. The effect of the reaction temperature on the response of the system was also investigated. Fig. 2D depicts the ΔRCT response of the sensor at varying reaction temperatures ranging from 25 to 40 °C. The peak current increased with increasing temperature from 25 to 40 °C and then decreased rapidly as the reaction temperature was increased from 40 to 50 °C. Therefore, 40 °C was chosen as the optimal reaction temperature.
3.3. Detection of Cu2+
Under optimized experimental conditions, the modified electrode in 0.1 M PBS (pH 7.4) containing 5 mM Fe(CN)63−/4− (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and 10 mM KCl after the electrodes were incubated with different concentrations of Cu2+, were examined using Nyquist plots. As shown in Fig. 3A, upon decreasing the concentration of Cu2+ from 400 nM to 0.1 nM, less trans-acting catalytic strands cleaved the sessile of the substrate into two fragments, which led to a decrease in ΔRCT. The change in the ΔRCT was linear with the logarithm of the Cu2+ concentration within a concentration range from 0.1 nM to 400 nM. The linear regression equation was Z = (−602.69 ± 28.50)X + (6182.16 ± 225.72) (Z is the ΔRCT (Ω), where X is the common value of the target concentration (M) with a correlation coefficient of r2 = 0.9945). The detection limit (LOD) of this sensor was estimated to be 0.0725 nM (based on S/N = 3). As shown in Table 1, this novel impedimetric biosensor showed a comparable LOD and linear detection range compared to other enzyme-based electrochemical DNA sensors for Ag+, Hg2+ and Cu2+ using EIS, and the linear range and detection limit of this novel electrochemical sensor was comparable to some of the other methods. Moreover, this method was relatively simple.
1) and 10 mM KCl after the electrodes were incubated with different concentrations of Cu2+, were examined using Nyquist plots. As shown in Fig. 3A, upon decreasing the concentration of Cu2+ from 400 nM to 0.1 nM, less trans-acting catalytic strands cleaved the sessile of the substrate into two fragments, which led to a decrease in ΔRCT. The change in the ΔRCT was linear with the logarithm of the Cu2+ concentration within a concentration range from 0.1 nM to 400 nM. The linear regression equation was Z = (−602.69 ± 28.50)X + (6182.16 ± 225.72) (Z is the ΔRCT (Ω), where X is the common value of the target concentration (M) with a correlation coefficient of r2 = 0.9945). The detection limit (LOD) of this sensor was estimated to be 0.0725 nM (based on S/N = 3). As shown in Table 1, this novel impedimetric biosensor showed a comparable LOD and linear detection range compared to other enzyme-based electrochemical DNA sensors for Ag+, Hg2+ and Cu2+ using EIS, and the linear range and detection limit of this novel electrochemical sensor was comparable to some of the other methods. Moreover, this method was relatively simple.
| Method | Materials | Linear range (mol L−1) | LOD (mol L−1) | References |
|---|---|---|---|---|
| Electrochemiluminescence/DNAzyme | 6-Carboxyfluorescein | 8 × 10−8–2 × 10−6 | 3.5 × 10−9 | 10 |
| Fluorescence detection/DNAzyme | Microarray | 1 × 10−8–1 × 10−4 | 9.5 × 10−9 | 15 |
| Fluorescence detection/DNAzyme | SYBR Green I (SG) | 4 × 10−8–1.2 × 10−6 | 1 × 10−8 | 16 |
| Colorimetric detection/DNAzyme | Horseradish peroxidase | 5 × 10−8–1.2 × 10−6 | 5.9 × 10−9 | 17 |
| Colorimetric detection/DNAzyme | Gold nanoparticles | 1 × 10−10–2 × 10−9 | 6 × 10−11 | 18 |
| Colorimetric detection/DNAzyme | Gold nanoparticle | 1 × 10−9–2 × 10−8 | 4.7 × 10−10 | 19 |
| Detection Ag+/EIS/DNA | Ordered mesoporous carbon nitride material | 1 × 10−10–1 × 10−5 | 5 × 10−11 | 20 |
| Detection Hg2+/EIS/DNA | Gold electrode | 1 × 10−10–1 × 10−3 | 1 × 10−10 | 21 |
| Detection Cu2+/EIS/DNAzyme | Avidin–graphite epoxy composite | 1 × 10−5–4 × 10−5 | 6.5 × 10−6 | 22 |
| Detection Pb2+/EIS/DNAzyme | OMC–GNPs | 5 × 10−10–5 × 10−5 | 2 × 10−10 | 9 |
| Detection Cu2+/EIS/DNAzyme | GNCs | 1 × 10−10–4 × 10−7 | 7.25 × 10−11 | This work |
3.4. Reproducibility, stability and selectivity of the biosensor
The reproducibility of this biosensor was investigated. Six biosensors were fabricated with six different GCEs using the same steps independently and used to detect 200 nM Cu2+, as presented in Fig. 4. The RSD was 4.91% with five biosensors prepared independently, indicating that the fabrication procedure was reliable, and this biosensor had good reproducibility. | ||
| Fig. 4 Six different GCEs constructed by the same procedure on response of biosensor for Cu2+ (200 nM). The error bars indicate the standard deviations from three replicative tests. | ||
The stability of the biosensor was also explored. We investigated the stability of this sensor through the response to 200 nM Cu2+ for 1 month (as shown in Fig. S-2B†). Beyond this period, the experiment was carried out every 5 days. When not in use, the electrode was stored in a moist state at 4 °C. The result showed that the biosensor retained approximately 81% of its original ΔI after 1 month. The result indicated that this biosensor has relatively good stability because the DNAzyme-based sensor is sensitively and specifically responsive to its target ion, and the film (GNCs) could provide a biocompatible microenvironment.
Methods related to the sensing of metal ions by the DNAzyme-based sensor concern the specificity of the system. Thus, the selectivity of this detection method was tested using the impedimetric Cu2+ sensor in 0.1 M PBS (pH 7.4) containing 5 mM Fe(CN)63−/4− (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and 10 mM KCl. Under the optimal experimental conditions, 200 nM of Cu2+, 2000 nM of Fe3+, Zn2+, Mn2+, Co2+, Hg2+, Pb2+, Cd2+, Ca2+, and their mixture containing 200 nM of Cu2+, and their mixture without Cu2+ were measured. As shown in Fig. 5, a negligible signal response was observed upon the addition of the other tested ions, whereas significant response of ΔRCT as observed for Cu2+. Therefore, the results showed excellent selectivity toward Cu2+ over other ions due to the specificity of DNAzyme for Cu2+ ions. Cu2+ and other ions were then mixed to form a mixture solution as a sample for the anti-jamming capability testing of this sensor (Fig. 5). The ΔRCT was obviously higher than the other samples without Cu2+. These results clearly indicated that the approach is not only insensitive to other ions but also selective toward Cu2+ in their presence. As discussed above, the present sensor had excellent selectivity and anti-jamming capability.
1) and 10 mM KCl. Under the optimal experimental conditions, 200 nM of Cu2+, 2000 nM of Fe3+, Zn2+, Mn2+, Co2+, Hg2+, Pb2+, Cd2+, Ca2+, and their mixture containing 200 nM of Cu2+, and their mixture without Cu2+ were measured. As shown in Fig. 5, a negligible signal response was observed upon the addition of the other tested ions, whereas significant response of ΔRCT as observed for Cu2+. Therefore, the results showed excellent selectivity toward Cu2+ over other ions due to the specificity of DNAzyme for Cu2+ ions. Cu2+ and other ions were then mixed to form a mixture solution as a sample for the anti-jamming capability testing of this sensor (Fig. 5). The ΔRCT was obviously higher than the other samples without Cu2+. These results clearly indicated that the approach is not only insensitive to other ions but also selective toward Cu2+ in their presence. As discussed above, the present sensor had excellent selectivity and anti-jamming capability.
 | ||
| Fig. 5 Interference study in the analysis of Cu2+ by this biosensor. The data is average of three replicate measurements. The error bars indicate standard deviations. | ||
3.5. Analysis of real samples
To evaluate the practicality of the present method, the biosensor was applied to detect the recovery of Cu2+ with water samples taken from Xiangjiang River, Hunan province. These Cu2+ concentrations in environmental samples were adjusted to fall in the concentration range from 1 to 200 nM by spiking with Cu2+ stock solutions, followed by measuring the Cu2+ content using the proposed method and ICP/MS. The results summarized in Table 2 show good agreement with those achieved by ICP/MS, indicating that the present sensor can also work in environmental water samples.| Sample number | Addition concentration (nM) | Biosensor (meana ± SDb) (nM) | ICP/MS (meana ± SDb) (nM) | Relative standard deviation (%) |
|---|---|---|---|---|
| a An average of three replicate measurement.b SD = standard deviation. | ||||
| 1 | 0 | 12.12 ± 0.89 | 13.01 ± 0.76 | 5.00 |
| 2 | 1 | 20.17 ± 1.3 | 19.54 ± 1.22 | 2.24 |
| 3 | 50 | 71.20 ± 2.4 | 73.92 ± 1.5 | 2.65 |
| 4 | 100 | 113.47 ± 5.1 | 117.85 ± 6.9 | 2.68 |
4. Conclusions
A novel impedimetric biosensor was reported to determine Cu2+ concentration using the difference in charge-transfer resistance (ΔRCT) before and after DNA interactions with Cu2+; ΔRCT is sufficiently sensitive to detect Cu2+ as low as 0.0725 nM, and the linear range was from 0.1 M to 400 nM. Moreover, because of the signal amplification on the GNCs platform and the merits of high specificity of the DNAzymes, the sensor maintained high selectivity over the other nonspecific metal ions. Considering the good performance of this electrochemical Cu2+ sensor, it is expected to obtain great success in environmental monitoring.Acknowledgements
The authors gratefully acknowledge the Program for Changjiang Scholars (T2011116), the Key Projects of Science and Technology of Hunan Province, China (2012FJ1010 and 2014FJ1011), the Natural Science Foundation of China (51474247), the National High Technology Research and Development Program of China (2011AA061001) for financial support and the Natural Science Foundation of China (51474247).References
- E. L. Que, D. W. Domaille and C. J. Chang, Chem. Rev., 2008, 108, 1517–1549 CrossRef CAS PubMed.
- P. G. Georgopoulos, A. Roy, M. J. Yonone-Lioy, R. E. Opiekun and P. J. Lioy, J. Toxicol. Environ. Health, Part B, 2001, 4, 341–394 CAS.
- A. P. S. Gonzáles, M. A. Firmino, C. S. Nomura, F. R. P. Rocha, P. V. Oliveira and I. Gaubeur, Anal. Chim. Acta, 2009, 636, 198–204 CrossRef PubMed.
- N. G. Beck, R. P. Franks and K. W. Bruland, Anal. Chim. Acta, 2002, 455, 11–22 CrossRef CAS.
- M. Etienne, J. Bessiere and A. Walcarius, Sens. Actuators, B, 2001, 76, 531–538 CrossRef CAS.
- A. Mohadesi and M. A. Taher, Talanta, 2007, 72, 95–100 CrossRef CAS PubMed.
- B. High, D. Bruce and M. M. Richter, Anal. Chim. Acta, 2001, 449, 17–22 CrossRef CAS.
- Q. Suyan, G. Sen, Z. Xi, L. Zhenyu, Q. Bin and C. Guonan, Analyst, 2011, 136, 1580–1585 RSC.
- Y. Y. Zhou, L. Tang, G. M. Zeng, C. Zhang, X. Xie, Y. Y. Liu, J. J. Wang, J. Tang, Y. Zhang and Y. C. Deng, Talanta, 2016, 146, 641–647 CrossRef CAS PubMed.
- L. Juewen and L. Yi, J. Am. Chem. Soc., 2007, 129, 9838–9839 CrossRef PubMed.
- J. J. Yao, J. S. Li, J. Owensc and W. W. Zhong, Analyst, 2011, 136, 764–768 RSC.
- Y. Zhang, G. M. Zeng, L. Tang, Y. P. Li, Z. M. Chen and G. H. Huang, RSC Adv., 2014, 4, 18485–18492 RSC.
- L. Chen, G. Zeng, Z. Yi, T. Lin, D. Huang, C. Liu, Y. Pang and L. Jie, Anal. Biochem., 2010, 407, 172–179 CrossRef CAS PubMed.
- E. L. S. Wong, P. Erohkin and J. J. Gooding, Electrochem. Commun., 2004, 6, 648–654 CrossRef CAS.
- Z. Peng, B. C. Yin and B. C. Ye, Biosens. Bioelectron., 2009, 25, 935–939 CrossRef PubMed.
- L. Zhang, Y. Zhang, M. Wei, Y. Yi, H. Li and S. Yao, New J. Chem., 2013, 37, 1252–1257 RSC.
- G. Chenchen, L. Quan, W. Dou, Z. Shiming, L. Xiaoling, Y. Luxin, X. Xuerong and Z. Lingwen, Anal. Chem., 2014, 86, 6387–6392 CrossRef PubMed.
- X. Miao, L. Ling, D. Cheng and X. Shuai, Analyst, 2013, 137, 3064–3069 RSC.
- L. Lu, F. Jie, Y. Fan, T. Bo and A. Chem, Anal. Chem., 2015, 87, 4829–4835 CrossRef PubMed.
- Y. Zhou, L. Tang, X. Xia, G. Zeng, J. Wang, Y. Deng, G. Yang, C. Zhang, Y. Zhang and J. Chen, Analyst, 2014, 139, 6529–6535 RSC.
- R. G. Cao, B. Zhu, J. Li and D. Xu, Electrochem. Commun., 2009, 11, 1815–1818 CrossRef CAS.
- O. A. Cristina, M. Natalia, D. V. Manel and P. Valeri, Analyst, 2013, 138, 1995–1999 RSC.
Footnote |
| † Electronic supplementary information (ESI) available: More information is detailed regarding the SEM images of GNCs, cyclic voltammetry diagrams of GCE, GCE/GNCs, electrochemical impedance spectra of GCE and GCE/GNCs. See DOI: 10.1039/c5ra20641c |
| This journal is © The Royal Society of Chemistry 2016 |