Safflomin A inhibits neuraminidase activity and influenza virus replication
Miao Yua,
Ye Wangb,
Li Tianc,
Yanyan Wanga,
Xizhu Wanga,
Weiguo Lianga,
Jiyu Yanga,
Dahai Yua,
Tonghui Ma*d and
Xuexun Fang*a
aKey Laboratory for Molecular Enzymology and Engineering of Ministry of Education, College of Life Sciences, Jilin University, 2699 Qianjin Street, Changchun 130012, China. E-mail: fangxx@jlu.edu.cn; Fax: +86-431-85155249; Tel: +86-431-85155249
bSchool of Life Science, Jilin University, 2699 Qianjin Street, Changchun 130012, China
cState Key Laboratory of Electroanalytical Chemistry, Changchun Institute of Applied Chemistry, Chinese Academy of Sciences, Changchun 130022, China
dCollege of Basic Medical Sciences, Dalian Medical University, Dalian 116044, China. E-mail: tonghuima@dlmedu.edu.cn; Fax: +86-0411-86110378; Tel: +86-0411-86110378
First published on 26th October 2015
Abstract
Neuraminidase (NA) is a glycoprotein on the surface of the influenza virus that plays an important role in the early processes of virus infection and viral release from the infected cells. NA inhibitors are currently the most effective drugs to treat influenza virus infection. Many traditional Chinese medicines (TCMs) in various formulations have been used in Chinese clinics to treat influenza, however, the effective constituents and the mechanism of action are mostly unknown. In this paper, we have tested almost 300 natural compounds from a Chinese medicinal herbal compound library to evaluate their anti-NA activities in vitro. Safflomin A (SA) was one of the compound detected with NA inhibitory activities. It showed inhibition against neuraminidases from H1N1 and H3N2 of type A and neuraminidase of type B influenza viruses. Enzyme kinetic tests using SA revealed that the types of inhibition against N1 and N2 neuraminidases were noncompetitive. The interaction of SA with the N1 and N2 neuraminidases was analyzed using molecular simulation and docking, which showed that SA was bound in the non-active sites of N1 and N2. SA was also analyzed for its cytotoxicity and anti-viral activities in cell culture. It showed inhibitory effect for viral replication of H1N1 and H3N2 influenza viruses in MDCK cells. Enzymatic analysis indicated that a SA and oseltamivir carboxylate combination treatment was synergistic with combination index (CI) values ranging between 0.49 and 0.52 against neuraminidase of A/New Caledonia/20/1999 (H1N1), and ranging between 0.56 and 0.88 against neuraminidase of A/Fujian/411/2002 (H3N2). These results suggest that an herbal formulation containing SA in combination with oseltamivir carboxylate may serve as a potential therapeutic option to currently available anti-influenza therapeutics.
1 Introduction
Influenza viruses annually cause epidemics and complications, and even significant morbidity and fatality, especially in senior citizens who have underlying cardiopulmonary diseases.1,2 In 2009, worldwide H1N1 influenza epidemics infected millions and caused severe fatality. In February 2013, a novel avian influenza H7N9 virus emerged in China and was found to infect humans.3 There is evidence that influenza pandemics occurred four times in the 20th century and caused 20 to 50 million deaths worldwide.4 Influenza like symptoms typically include headache, sore throat, muscle ache, fatigue and fever. Most patients were critically ill and presented sever pneumonia, acute kidney failure, acute respiratory distress syndrome, diffuse intravascular coagulation and septic shock.4–6Influenza virus is a highly infectious respiratory virus which could infect human and other animals and is classified into three genera (i.e. influenza A, B and C).7 There are two different types of viral integral glycoproteins, named hemagglutinin (HA) and neuraminidase (NA). The influenza A virus is called by the type of HA (H1-H16) and NA (N1-N9) it contains. The structure of the two proteins change easily through a process of antigenic drift that could result in influenza virus variations. According to the “150-cavity” adjacent to the active site, the NA of influenza A has been divided into two different classes, known as group-1 (comprising N1, N4, N5, N8) and group-2 (comprising N2, N3, N6, N7, N9).8 The NA plays an important role in influenza A virus replication and facilitates the early process of influenza virus infection in human airway epithelium, which can cleave terminal sialic acids of sialoglycans and promote the release of progeny virus from the infected host cell.7–10 Hence, as an ideal target for design of anti-influenza drugs, the NA presents on the surface of the influenza virus has attracted global attention.
Since vaccination is less effective or not available upon the influenza virus variants outbreak, the primary focus of attention has been on drugs. There are only two classes of anti-influenza drugs approved by U.S. FDA for the clinical treatment and prophylaxis: M2 ion channel blockers (amantadine and rimantadine) and neuraminidase inhibitors (oseltamivir, zanamivir and peramivir). According to the U.S. Centers for Disease Control and Prevention, 100% of seasonal H3N2 and 2009 pandemic flu have shown resistance to adamantanes and rimantadine, thus amantadine and rimantadine are no longer recommended for treatment of influenza.11 The neuraminidase inhibitors are currently the only clinically effective drugs to treat influenza. However, all of these synthetic drugs have the side effects and the widely used of these drugs resulted in the emergence of drug-resistant mutant viruses in recent years.12–14 Studies have shown that influenza virus isolated in human resistant to oseltamivir were frequent in Japan than in Europe.15 There were 20% isolates in Europe which was oseltamivir-resistant H1N1 mutants.16 Influenza viruses which has resistant have universal shape of the NA catalytic site. The active site of the NA content 8 functional residues (R118, D151, R152, R224, E276, R292, R371 and Y406) and surrounded by 11 framework residues (E119, R156, W178, S179, D198, I222, E227, H274, E277, N294 and E425).17 Both oseltamivir and zanamivir inhibited NA by competing with the substrate at the catalytic site. One single change of catalytic site of NA could reduce the inhibition of oseltamivir or zanamivir to the influenza virus. H274Y mutation of influenza virus (H1N1) was reported to be oseltamivir resistant.18 Vasiliy P. et al. found neuraminidase D151G mutation of influenza A (H3N2) was more resistant to the zanamivir and the IC50 was almost increased 1500-fold.19 In recent years, the novel avian H7N9 caused more than 227 deaths in the world. This novel H7N9 virus is a triple reassortant virus taking along genes from H7N3, H9N2 and H11N9 or H2N9 avian influenza A viruses.20 Zhou et al. found that the H7N9 virus can bind to both avian-type and human-type receptors.21 The isolated H7N9 with neuraminidase R292K mutation is broad resistance to neuraminidase inhibitors.20,22 Due to lack of the commercial drugs and emergence of oseltamivir-resistant virus, new drugs should be presented to oppose this new fatal influenza virus.
Favipiravir (T-705) is an antiviral drug that selectively inhibits the RNA-dependent RNA polymerase of influenza virus.21 Sleeman et al. found that favipiravir inhibits in vitro replication of a wide range of influenza viruses, including those resistant to currently available drugs.23 Other interesting drug targets, such as nucleoprotein, viral polymerase, and non-structural protein NS1A, have recently been well reviewed by Das and colleagues.24 In 2009, Dias et al. showed that a intrinsic RNA and DNA endonuclease is located in the PA subunit of virus polymerase. Inhibitors targeting of this endonuclease may also emerge as potential new anti-influenza drugs.25
Herbal remedies with the natural and active substances play a role as anti-influenza agents in controlling and treatment of influenza virus. A series of traditional Chinese medicinal (TCM) has been reported with antiviral effects, such as Melia toosendan could inhibit influenza virus through suppression of NA both in vitro and vivo.26 Forsythia suspensa extract against influenza A virus (H1N1) in vitro and in vivo using the cytopathic effect and neutral red dye uptake assays.27 Based on a large amount of studies, a variety of efficiently bioactive compounds isolated from TCMs have been identified to treat influenza.7 In the last few years, researchers had focused on the components of herbs to anti-influenza. Jin-Yuan Ho et al. extracted the ethanolic component from Paeonia lactiflora, which exhibited low IC50 values against influenza virus A/WSN/33 (H1N1) in vitro and vivo. It also has inhibitory activity against H1N1pdm strain which was isolated from clinical oseltamivir-resistant virus.28 The aqueous extracts of H. erectum, M. cochinchinensis and T. chebula reduced the titre of A/Teal/Tunka/7/2010 (H3N8) in vitro.29 Wang et al. used a system pharmacology approach to detect 50 compounds from two herbs aim to map to target-disease and drug-target-pathway networks in silico.30
Computer-based modeling methodology has been widely used to design new drugs. It enhanced efficiency of detection and reduced the cost of experiment. Molecular docking has been usually utilized to drug discovery especially against influenza virus.31 An et al. utilized computational molecular docking to identify potential inhibitors of the H5N1 NA in 20 compounds.32 Tambunan et al. found two ligands that have good IC50 values of the target interaction to H1N1 NA and toxicological properties using simulation results of molecular docking. These two ligands are predicted with no mutagencity or carcinogenicity and have good oral bioavailability.33 We have also taken advantage of homology modeling to simulate the structure of H1N1 NA, the model structure of N1 was used to screen compounds of NPD database34 for potential inhibitors. We found 34 of compounds with low affinity energy, among which, two natural compounds (ZINC02128091 and ZINC02098378) possessed the most favorable interaction energy.35
In this study, we tested 289 natural compounds by NA inhibition assay. Among these compounds, safflomin A (SA) showed NA inhibitory activity and was analyzed for its anti-viral effect by measuring cytopathic effect (CPE) of the influenza A virus. We also determined the type of inhibition of SA towards NA by enzyme kinetics. Moreover, the interaction of SA and influenza NA was analyzed by molecular docking. Finally, combination studies were undertaken to investigate whether SA could act synergistically with oseltamivir carboxylate against the NA of H1N1 and H3N2.
2 Materials and methods
2.1 Reagents
2-(N-Morpholino)ethanesulfonic acid (MES, >99%), dimethylsulfoxide (DMSO), 2′-(4-methylumbelliferyl)-α-D-N-acetylneuraminic acid, sodium salt hydrate (MU-NANA, CAS no. M8639) and 3-[4,5-dimethyl-thiazol-2-yl]-2,5-diphenyl tetrazolium bromide (MTT, >98%) were purchased from Sigma-Aldrich Corporation. Fetal bovine serum (FBS), Dulbecco's modified Eagle's medium (DMEM), TPCK-treated trypsin and trypsin–EDTA were purchased from the Gibco Company. Oseltamivir carboxylate was purchased from Dalian meilun biology technology co. Other chemical reagents were purchased from Beijing Chemical Works.2.2 Compounds from traditional Chinese medicines (TCMs)
All compounds (analytical reagent, see Appendix) which are ingredients of TCMs were purchased from the Chinese Standard Chemical Networks (http://www.crmrm.com/). Each compound was dissolved in DMSO to the final concentration of 20 mM and stored in 96-well plates numbered α, β, γ, and δ at −80 °C.2.3 Cells and virus
Madin–Darby canine kidney (MDCK) cells were obtained from Norman Bethune Health Science Center of Jilin University and grown in DMEM supplemented with 10% FBS, 100 μg mL−1 streptomycin, 100 U mL−1 penicillin. Influenza viruses (A/New Caledonia/20/1999 (H1N1), A/Fujian/411/2002 (H3N2), B/Jiangsu/10/2003) which were used as the source of NAs was obtained from Changchun Institute of Biological Products. Each type was propagated in 9 days old embryonated chicken eggs. The allantoic fluid was collected, then centrifuged at 3000 rpm min−1 for 5 min (ref. 36) and the supernatant was stored at −80 °C.2.4 NA activity assay
NA activity was assayed with MU-NANA as a substrate.37 The supernatant of chicken egg embryonic allantoic fluid were first dissolved in pH 6.5 MES buffer (32.5 mM), which contains 4 mM calcium chloride. For NA activity inhibition assay, 49 μL of the supernatant was incubated with 1 μL DMSO dissolved TCM compound at different concentrations at 37 °C for 30 min in a black 96-well ELISA plate. Then 50 μL of 20 μM MU-NANA were added. TCM compounds with or without 20 μM MU-NANA were also detected the fluorescence intensity as control. Fluorescence intensity was measured using the FLX800 fluorescence microplate reader (Bio-Tek) with excitation wavelength at 360 nm and emission wave length at 450 nm. The detection lasted for 8 min. The fluorescence intensity changing rate of was determined as the NA activity. The level of inhibition was determined by the initial rates of the reaction with or without the compounds added. The median inhibitory concentration (IC50) was defined as the concentration of detected compounds to reduce the NA activity by 50% relative to the reaction mixture containing NA without compounds.2.5 Median tissue culture infective dose (TCID50) determination
MDCK cells were cultivated in DMEM and re-plated 2 days before infection in 96 wells plates. Serial 10-fold dilutions of the allantonic fluid (H1N1 or H3N2) were prepared, then the diluted fluid was added in MDCK cells which contain 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well to incubate for 2 h for virus to infect. The media for dilution was DMEM supplemented with 1 μg mL−1 TPCK-treated trypsin. Treated MDCK cells were washed 3 times by DMEM and cultured in DMEM with TPCK-treated trypsin (1 μg mL−1) for 24 h. 24 h later, MDCK cells were washed 3 times and cultured in DMEM with 2% FBS. After 3 days, MTT assay was used to determinate the virus-induced cytopathic effect (CPE), which indicated the presence of virus, TCID50 was calculated by the Reed–Muench method.
000 cells per well to incubate for 2 h for virus to infect. The media for dilution was DMEM supplemented with 1 μg mL−1 TPCK-treated trypsin. Treated MDCK cells were washed 3 times by DMEM and cultured in DMEM with TPCK-treated trypsin (1 μg mL−1) for 24 h. 24 h later, MDCK cells were washed 3 times and cultured in DMEM with 2% FBS. After 3 days, MTT assay was used to determinate the virus-induced cytopathic effect (CPE), which indicated the presence of virus, TCID50 was calculated by the Reed–Muench method.
2.6 Cytotoxicity assay
MTT assay was also used to determinate the cytotoxic effects of natural compounds on MDCK cells. MDCK cells were grown in 96 wells plate in DMEM with 10% FBS and subsequently incubated with concentration gradient of compounds for 24 h at 37 °C. Cell viability was detected by 15 μL MTT (5 mg mL−1) in 100 μL DMEM for 4 h and measured the absorbance with spectrophotometer at 570 nm. The half-maximum cytotoxic concentration (CC50) was defined as the concentration which reduced the 50% OD570 of treated cells to untreated cells.2.7 Viral growth inhibition assay
To detect the antiviral effect of the samples on influenza virus (H1N1 and H3N2), MDCK cells were grown in DMEM medium supplemented with 10% FBS and antibiotics. For the assay, MDCK cells were seeded into 96-well culture plates at a density of 3 × 103 per well and incubated for 24 h until 75–85% confluency. Monolayer MDCK cells were washed with DMEM medium twice. There are two types of treatment: (1) we used 200 μL aliquot of virus which is at the optimal dose (100 TCID50) mixed with 200 μL concentration gradient of compounds for 2 h at 37 °C. The mixture was added into the wells to infect the MDCK cells. 2 h later, the virus-infected MDCK cells were washed with DMEM medium three times and cultured with DMEM medium with trypsin (1 μg mL−1) and after 24 h, DMEM which contained the 2% FBS was changed; (2) we used 200 μL aliquot of virus at the optimal dose for 2 h at 37 °C in medium without serum. The virus-infected MDCK cells were washed with DMEM medium three times and added different concentrations of compounds for 2 h.26 The cells in the absence of compound without or with virus added were used as control.For both types of treatment, 2 h later, MDCK cells were also washed with DMEM medium three times and cultured with DMEM medium with trypsin (1 μg mL−1) and after 24 h, DMEM which contained the 2% FBS was changed. Following 3 days incubation, the monolayers were examined for CPE. For the CPE, the monolayers were washed to remove dead cells and debris, MTT assay was used to evaluate the antiviral effect of the anti-NA compounds on influenza virus. Concentration for 50% of maximal effect (EC50) was defined as the concentration of detected compounds to enhance the cell viability by 50% between the cell viability of cells infected influenza virus without compounds and the cell viability of cells with no infection.
After 3 days of incubation, the cells contained cell-associated virus and extracellular virus with DMEM culture solution were frozen and unfrozen 3 times to rupture the cells. Then the solution was centrifuged with 2000 rpm for 5 min.38 The supernatant was transferred, and the quantity of virus was represented by the NA activity. 50 μL of supernatant was added into black 96-well ELISA plate and 50 μL of 20 μM MU-NANA were also added, the rest of operation was carried out as described above. The solution in the absence of compound with virus was used as control.
2.8 The inhibition type of SA on the activity of NA
To determinate the type of inhibition of NA by SA, we performed enzyme kinetics with NAs from influenza virus (H1N1 and H3N2). In NA activity assay, we used 50 μL allantonic fluid which contain of NA of influenza virus (H1N1 or H3N2) as the control and added 100, 150 or 200 μM SA respectively, the mixtures were added into a black Elisa plate and incubated at 37 °C for 30 min. Then 50 μL MU-NANA substrate solution which concentration were 0.5, 1, 1.5, 2 nmol were added into the control or the experiment wells. The fluorescence intensity measured as mentioned in NA activity assay. Based on the reciprocal concentration (X) and the reciprocal fluorescence intensity (Y), the double-reciprocal plot was obtained.2.9 Molecular docking simulation
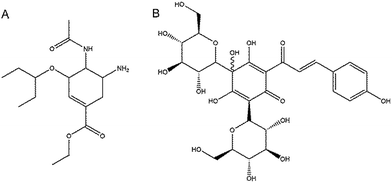
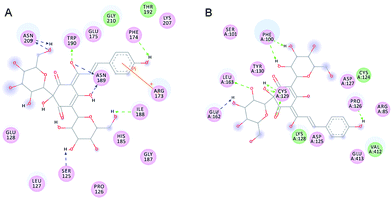
To study the interaction and the conformation of the protein–ligand complex, SA was docked into the N1, N2 NA protein using the AutoDock 4.0 program based on Lamarkian Genetic Algorithm (Scripps Research Institute, La Jolla, CA).39 Due to the lack formation of NA which we used, the crystal structure of N1, N2 which are highly homology (PDB ID: 3TI6_A and PDB ID: 4GZP_A) were obtained from the RCSB Protein Data Bank (http://www.rcsb.org/pdb). The structure of oseltamivir (Fig. 1A) was obtained from PubChem (CID: 65028). The structure of SA (Fig. 1B) was obtained from PubChem (CID: 6443665). The predicted complexes were optimized and ranked according to the empirical scoring function, ScreenScore, which estimated the binding free energy of the ligand receptor complex. Each docking was performed twice, and each operation screened 250 conformations for the protein–ligand complex that were advantageous for docking; each docking had 500 preferred conformations. The most stable conformation distinguished which had the minimal binding energy was shown by Discovery Studio (DS) 3.5 Visualizer.2.10 Drug synergism studies
To analyze the drug synergism on the effect of treatment of NA activities of N1 and N2 with SA and the oseltamivir, alone or in combination, isobologram analysis were used, according to the results by CalcuSyn Windows software (Biosoft, Cambridge, United Kingdom) for dose-effect analysis and synergism/antagonism quantification.40 The conservative isobologram assumes that two drugs have independent or dissimilar modes of actions.41 For two-drug combination design, the combination of the drugs at their equipotent ratio (the ratio of their IC50s) was chosen. A mixture of the two drugs was prepared and the mixture was serially diluted (2-fold dilutions, SA: oseltamivir carboxylate is equal to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 0.5
1, 0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 0.25
1 and 0.25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 at the concentration of their IC50 values) to obtain a wide dosage range, as indicated in the CalcuSyn standard protocol. For single-drug treatment, serial dilutions were prepared from the serial dilutions of 0.1–0.5 times and 1–4 times of each compound's IC50 value, using the same protocol. 30 μL of dissolved NAs were added into a 96-black plate. After NAs were added, 10 μL of SA at the concentration mentioned above of was added and 10 μL of the oseltamivir carboxylate in the different proportional concentrations were added at 37 °C for 30 min, each concentration was repeated 3 times. Then 50 μL of 20 μM MU-NANA substrate solution was added. The fluorescence intensity was detected same as the NA activity assay.
1 at the concentration of their IC50 values) to obtain a wide dosage range, as indicated in the CalcuSyn standard protocol. For single-drug treatment, serial dilutions were prepared from the serial dilutions of 0.1–0.5 times and 1–4 times of each compound's IC50 value, using the same protocol. 30 μL of dissolved NAs were added into a 96-black plate. After NAs were added, 10 μL of SA at the concentration mentioned above of was added and 10 μL of the oseltamivir carboxylate in the different proportional concentrations were added at 37 °C for 30 min, each concentration was repeated 3 times. Then 50 μL of 20 μM MU-NANA substrate solution was added. The fluorescence intensity was detected same as the NA activity assay.
Dose inhibition curves were drawn for SA and oseltamivir carboxylate, used alone or in combination. The IC50 values were plotted against the fractional concentrations of oseltamivir carboxylate and SA on the x axis and y axis, respectively. The evaluation of drug synergism based on a median-effect equation has been widely used. Using the median-effect equation, dose-effect curves for each drug and combination of drugs were generated and the combination index (CI) were calculated. To determinate the interactions between SA and oseltamivir carboxylate, CI was calculated. CI analysis provides qualitative information on the drug interaction. Its value calculated as described in eqn (A).
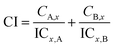 | (A) |
CA,x and CB,x represent the concentrations of drug SA and drug oseltamivir carboxylate used in combination to achieve x% single drug effect. ICx,A and ICx,B are the concentrations for single drug to achieve the same effect.
For two-drug combinations, in the isobologram plot, if the combination data point falls on the diagonal, an additive effect is indicated; if it falls on the lower left, synergism is indicated, whereas if it falls on the upper right, antagonism is indicated. A CI of >1 denotes antagonism, a CI equal to 1 denotes additivity, and a CI of <1 denotes synergism. In particular, CI values in the range of 0.1 to 0.3, 0.3 to 0.7, and 0.7 to 0.85 are considered to indicate strong synergism, synergism, and moderate synergism, respectively.
2.11 Statistical analysis
The results were expressed as mean ± S.E.M. for three independent experiments.3 Results
3.1 SA inhibit the NA activities of influenza virus
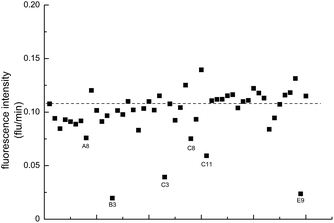
NA is the most effective drug target for influenza virus. There are numbers of herbal medicines used to treat influenza viral infection in traditional Chinese medicinal clinics, however, the compounds that could inhibit NA activity in these herbs have not been systematically tested. We used a natural compound library of 289 compounds which derived from TCMs to screen their NA inhibitory activities. First, NA of HIN1 virus was used in the NA activity assay and each compound was tested once. A representative result was shown in Fig. 2. An inhibition of the NA activity was detected when the fluorescent intensity of the product in the presence of the compound was lower than the control reaction. Then, we used selected compounds (as marked in Fig. 2) which showed inhibitory activity toward the NA of H1N1 to detect IC50 values of NAs for H1N1 and H3N2 of type A and type B influenza virus. The IC50 values were shown in Table 1. In order to eliminate the false positive caused by the natural compound, the fluorescence intensity of selected compounds with or without 20 μM MU-NANA were also detected. No influence on the fluorescence intensity in the presence of these compounds with or without substrate was observed. One compound, SA, showed inhibitory activities toward these NAs. The IC50 values were 143.80 ± 9.13, 155.33 ± 17.37, and 104.72 ± 8.91 μM toward NAs for H1N1 and H3N2 of type A and type B influenza virus, respectively.| Well no. in Plate δ | IC50 value towards NA of influenza virus H1N1 (μM) | IC50 value towards NA of influenza virus H3N2 (μM) | IC50 value towards NA of type B influenza virus (μM) |
|---|---|---|---|
| A8 | ≥200 | — | ≥200 |
| B3 | 164.53 ± 4.05 | ≥200 | 136.51 ± 16.20 |
| C3 | 143.80 ± 9.13 | 155.33 ± 17.37 | 104.72 ± 8.91 |
| C8 | ≥200 | — | ≥200 |
| C11 | ≥200 | — | ≥200 |
| E9 | ≥200 | ≥200 | ≥200 |
3.2 Inhibition of influenza A virus growth in MDCK cells by the SA
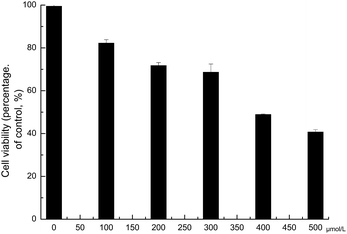
Since SA showed anti-NA activities, we then tested whether it could protect cells from viral infection. We first examined the effect of SA on the cell viability. MDCK cells were used in the assay with concentration gradient of SA (from 100 to 500 μM). Half-maximum cytotoxic concentration (CC50) of SA after 24 h exposure of MDCK cells to it were shown in Fig. 3 and the CC50 is 414.12 ± 12.58 μM. This result showed that SA had minimal cytotoxicity in low concentrations.In the presence of the virus, the MDCK cells viability was reduced. To determine the median tissue culture infective dose (TCID50) of H1N1 or H3N2, MDCK cells were cultivated in DMEM and a serial 10-fold dilutions of the allantonic fluid (containing H1N1 or H3N2) were added for virus to infect. Three days after infection, MTT assay was used to determinate the virus-induced cytopathic effect (CPE), and TCID50 was calculated by the Reed–Muench method. The TCID50 values were 10−4.2 (H1N1) and 10−7.5 (H3N2), respectively.
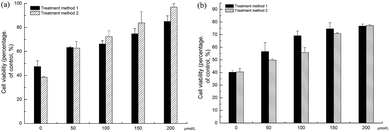
To determine the anti-viral activity of SA, the MDCK cells were infected 100 TCID50 of virus. Compared with control which has no influenza virus infection, the cell viability was greatly reduced when the MDCK cells were infected with 100 TCID50 of virus, which indicated the H1N1 virus caused severe damage to the MDCK cells (Fig. 4a). To determinate whether the SA protected the MDCK cells from the infection of H1N1 influenza virus, we used two types of treatments. As mentioned in the methods section, SA was added to the MDCK cells upon or post viral infection. The results demonstrated that under both conditions, SA protected MDCK cells from viral damage, the protecting effects increased with the increase of the compound concentration (Fig. 4a). The EC50 of SA measured is 82.36 ± 11.74 μM for the viral infected MDCK cells (Table 2). Thus in agreement with that SA inhibited NA activities, the compound could reduce cell damage caused by influenza virus infection.
| IC50(μM) | CC50 (μM) | EC50 (μM) | |||
|---|---|---|---|---|---|
| H1N1 | H3N2 | B | H1N1 | H3N2 | |
| 143.80 ± 9.13 | 155.33 ± 8.91 | 104.72 ± 8.91 | 414.12 ± 12.58 | 82.36 ± 11.74 | 160.33 ± 2.06 |
We also used H3N2 virus to infect the MDCK cells in the same way as the H1N1 virus (Fig. 4b). The results were similar to H1N1 virus infected MDCK cells. The EC50 of SA measured is 160.33 ± 2.06 μM for the viral infected MDCK cells (Table 2).
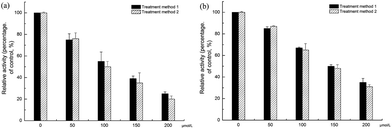
The results of cell viability indicated that SA could protect the MDCK cells from viral damage. Nonetheless, it did not show whether SA suppress the viral replication in MDCK cells. As viral numbers could be represented by the NA activities they carry, we measured the NA activity which reflect the multiplication of virus.42 The NA activities in MDCK cells infected the H1N1 and H3N2 without SA was regarded as 100%. When SA were added with an increased concentration, the cellular NA activities were reduced in both treatment methods against H1N1 or H3N2. Which represented the inhibition of the viral reproduction (Fig. 5). The results also showed that SA reduced the virus yield in MDCK cells in a dose-dependent manner and it were consistent with the results of CPE assay.
3.3 The inhibition type of SA on the activity of NA
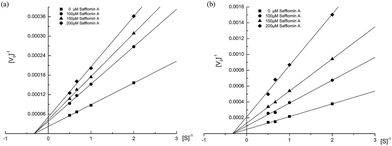
The inhibition kinetics of NA by SA were studied. Under the experimental conditions employed, the reaction of NA with SA followed Michaelis–Menten kinetics. The double-reciprocal plots of the enzyme inhibited by SA were shown in Fig. 6a (H1N1) and Fig. 6b (H3N2). The results showed that the plots of 1/v versus 1/[S] gave straight lines with different slopes, and the lines intersected at one point on the X-axis. These results suggested that the types of inhibition by SA were noncompetitive. It indicated that SA interacts with NA in a site other than the catalytic center, and this interaction may change the conformation of the NA active site and in turn reduce the activity of NA.3.4 SA bind to non-catalytic site of NA by molecular docking
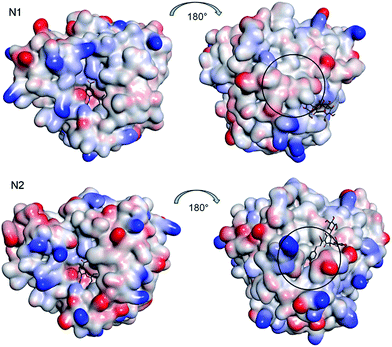
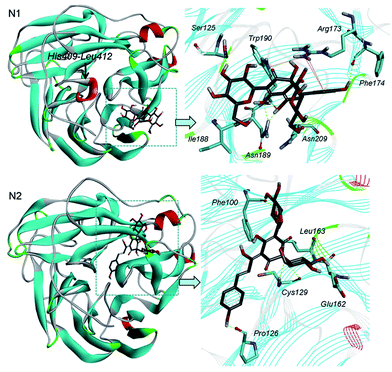
To study the possible interaction sites of SA and the NAs, we used molecular simulation analysis. The crystal structure of N1, N2 were obtained from the RCSB Protein Data Bank and SA was docked into the N1 or N2 NA protein using the AutoDock 4.0 program based on Lamarkian Genetic Algorithm as described in materials and methods. Our docking results showed that SA bound in the non-catalytic site of N1 and N2, which is different from the control drug, oseltamivir carboxylate. Oseltamivir docked to the center of the active site of N1 and N2. Meanwhile, SA bound at the backside of N1 or N2 (Fig. 7). Fig. 8 showed the details of the compound–NAs complexes. Notably, the locations where SA bound to N1 and N2 were different. The residues His409-Leu412 of N1 formed an alpha helix structure, which was the absent in N2 (Fig. 8). Such structural differences created a deep groove on N2 surface for SA to bind (Fig. 7 black circle).To identify the binding forces contributed to the compound–NAs complexes, the estimated free energy of binding, which including the van der Waals energy, electrostatic energy, H-bond energy, dissolving energy, final total internal energy, torsional free energy, and unbound system's energy, were calculated. The values of the estimated free energy of binding for SA with NAs shown in Table 3 were consistent with the experimental values: the inhibition constant of SA to N1 (143.80 ± 9.13 μM) were lower than to N2 (155.33 ± 8.91 μM), and the calculated free energy of binding for SA with NAs were −6.19 kcal mol−1 for N1 and −5.66 kcal mol−1 for N2.
| NAs | Energy (kcal mol−1) | IC50 (μM) |
|---|---|---|
| N1 | −6.19 | 143.80 ± 9.13 |
| N2 | −5.66 | 155.33 ± 8.91 |
Hydrogen bonds formed between compound and protein were usually contributed to the stability of the substrate–enzyme complexes, and more hydrogen bonds form a more stable complex. In the present study, the compound–NAs complexes models were visualized by DS4.0 molecular graphics system. Our results shown that SA formed eight hydrogen bonds with both N1 (Fig. 9A) and N2 (Fig. 9B), respectively. From analyzing the hydrogen bonds interaction, some critical NAs residues were identified (Tables 4 and 5). The residues of Ser125, Phe174, Ile188, Asn189, Trp190 and Asn209 of N1 were considered to be important in the interaction. Meanwhile, the presence of five residues, Phe100, Pro126, Cys129, Glu162 and Leu163, were previously identified as conserved residues in SA-N2 interaction.
| Donors atom | Receptors atom | Distances (Å) |
|---|---|---|
| Asn189: HD21 | SA: O12 | 2.12 |
| Asn189: HD21 | SA: O15 | 2.26 |
| Trp190: HN | SA: O15 | 1.70 |
| Asn209: HD22 | SA: O13 | 2.04 |
| SA: H66 | Asn209: OD1 | 2.08 |
| SA: H67 | Ile188: O | 2.22 |
| SA: H63 | Ser125: OG | 1.91 |
| SA: H75 | Phe174: O | 2.14 |
| Donors atom | Receptors atom | Distances (Å) |
|---|---|---|
| Cys129: HN | SA: O11 | 1.91 |
| Cys129: HN | SA: O3 | 2.42 |
| Leu163: HN | SA: O4 | 2.17 |
| SA: H56 | Cys129: O | 2.13 |
| SA: H60 | Glu162: OE1 | 1.75 |
| SA: H63 | Phe100: O | 1.90 |
| SA: H62 | Phe100: O | 1.97 |
| SA: H75 | Pro126: O | 2.13 |
SA bound to N1 not only by hydrogen bond interaction, but also by a strong conjugation effect (Fig. 9). Normal conjugated effect, also known as pi–pi interaction, was due to the formation of conjugated Pi bond caused by the effect of the molecular nature of the charge. By analyzing the pi–pi interaction results (Table 6), Arg173 of N1 was identified to be important amino acids in the pi–pi interaction.
| End1 | End2 | Distances (Å) |
|---|---|---|
| SA | N1: Arg173 | 5.08 |
| SA | N1: Arg173 | 6.78 |
By analyzing the estimated free energy of binding, hydrogen bonds interaction and pi–pi interaction, we consider that SA has a strong binding effect with N1 than N2.
3.5 SA and oseltamivir act synergistically against N1 and N2
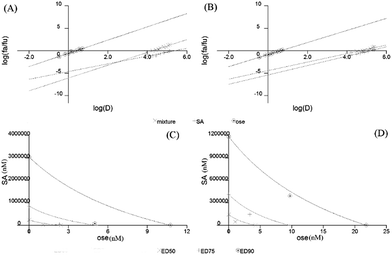
Oseltamivir carboxylate is widely used in the clinics for the treatment of influenza. It is well know as a potent competitive inhibitor for NA. Since SA inhibit noncompetitively of NA, it is of great interest to know if these two drugs can act synergistically. NAs were mixed with SA and oseltamivir carboxylate to detect the rate of inhibition. The effect of inhibitory rate on SA and oseltamivir carboxylate in combination against N1 and N2 were shown in Fig. 10. The SA-oseltamivir carboxylate combination treatment was significantly more effective than when used alone.To further analyze the synergistic effect of the two drugs, a log dose-effect curve and median-effect plot were generated for both drugs using the CalcuSyn software (Fig. 11A). The x intercept of the median-effect plot determines the median-effect dose (Dm), which, under the conditions described, is equivalent to the IC50 values of the drug; the Dm were found to 140.89 μM and 2.42 nM for SA and oseltamivir carboxylate against N1, respectively, when administered separately. The Dm values of the two drug were 18 μM and 1.07 nM for SA and oseltamivir against N1, respectively, when the drugs are used in combination, indicating that the doses needed to achieve 50% inhibitory of N1 were approximately 7.8-fold and 2.26-fold lower than those needed in monotherapy for SA and oseltamivir carboxylate, respectively.
We also detected the effect of SA and oseltamivir carboxylate against N2 (Fig. 11B). The Dm values were 144.07 μM and 4.248 nM for SA and oseltamivir carboxylate against N2, respectively, when administered separately. However, the Dm values of the two-drug combination were 45.437 μM and 1.05 nM for SA and oseltamivir carboxylate against N2, respectively, when the drugs are used in combination, indicating the doses needed to achieve 50% inhibitory N2 were approximately 3.17-fold and 4.05-fold lower than those needed in monotherapy for SA and oseltamivir carboxylate, respectively (Fig. 11B).
The evaluation of drug synergism based on a median-effect equation has been widely used. CalcuSyn-generated isobolograms based on the two drugs administered in combination at fixed ratios toward N1 and N2 are shown in Fig. 11C and D. The ED50, ED75, and ED90 (50%, 75%, and 90% equipotent doses) plots for each drug ratio fell below the line indicating a synergistic effect of the drug combination on NA activities. CI values were calculated for all conditions (Table 7). Under the used concentrations, the SA-oseltamivir carboxylate combination treatment was synergistic against both N1 and N2, with CI values ranging between 0.49 and 0.52 for N1, and CI values ranging between 0.56 and 0.88 for N2, respectively (Table 7). The CI values are consistent with conservative isobologram plots.
| CI at indicated ED | |||
|---|---|---|---|
| ED50 | ED75 | ED90 | |
| H1N1 | 0.52 | 0.50 | 0.49 |
| H3N2 | 0.56 | 0.70 | 0.88 |
4 Discussion
In this study, we tested 289 natural compounds for their inhibitory activities towards NAs of influenza virus. The NAs in the allantoic fluid of H1N1, and H3N2 of type A virus and type B virus were used in the assay. Among these compounds, we found that SA exhibited inhibitory activities to these NAs (Table 1). However, the IC50 values were different toward these NAs, which was mostly possible due to difference of the residues and structure at the active site of the influenza virus NAs.Next, we tested the viral inhibitory activity of the NA inhibiting compound SA in H1N1 or H3N2 infected MDCK cells. Although higher concentration of SA caused a reduction in the cell viability (Fig. 3), it also reduced CPE caused by influenza virus in MDCK cells in a dose-dependent manner (Fig. 4 and 5). These results suggest that SA could inhibit viral growth and protect cells from viral induced CPE by NA inhibition. Based on the structural characteristics of the NAs, type A influenza were divided into group-1 (comprising N1, N4, N5, N8) and group-2 (comprising N2, N3, N6, N7, N9). Our results also suggest that SA may reduce cell damage caused by either group-1 or group-2 type A influenza virus.
We also investigated the type of inhibition caused by SA with N1 or N2. According to the results shown in Fig. 6, SA inhibits NAs by a noncompetitive fashion. Molecular docking results were consistent with the enzyme kinetic analysis (Fig. 7). From analyzing the hydrogen bonds interaction, the residues of Ser125, Phe174, Ile188, Asn189, Trp190 and Asn209 of N1 were considered to be important (Table 4), and Phe100, Pro126, Cys129, Glu162 and Leu163 were identified as key residues in SA-N2 interaction (Table 5).
Traditional Chinese medicines (TCMs) have a long history in China to treat influenza. In the past, researchers found some TCMs to treat influenza. Heartleaf Houttuynia herb (Houttuynia cordata Thunb.) was found to inhibit MDCK cell apoptosis induced by H3N2 virus.43 Asiatic toddalia root (Toddalia asiatica Lam.) showed potent antiviral activities against H1N1 virus, with a 50% effective concentration valued of 4.7 mg L−1 in the MTS assay and 0.9 mg L−1 in the quantitative PCR assay.44 Pu et al. used extractive from Hypericum perforatum known as a TCM for anti-bacteria and defervescence against H1N1 virus in vivo and vitro, they found that it could inhibit the H1N1 in MDCK cells with an EC50 value of 40 μg mL−1.45 Despite a number of reports showed that TCMs could be inhibit influenza virus, limited effective compounds in TCMs were reported. As a consequence, the anti-influenza mechanisms for these TCMs were not known. In recent years, researchers have started to detect compounds in TCMs in cells or in animal models. A variety of polyphenols, flavonoids, saponins, glucosides and alkaloids isolated from TCMs have been studied and tested for anti-influenza activity. Hayashi et al. showed the inhibitory effect of trans-cinnamaldehyde (CA), one of the principal constituents of essential oil derived from cinnamomi cortex, on the growth of influenza A virus in vitro and in vivo, they found CA inhibit the influenza growth in dose-dependent manner and it could also inhibit the viral protein synthesis.46 Four diarylheptanoids from the seed of Alpinia katsumadai showed NA inhibitory activities in micromolar range in vitro against H1N1. Katsumadain A which inhibit the NA of H1N1 with IC50 value between 0.9 and 1.64 μM was also a constituent from the seed of Alpinia katsumadai.43
Dao et al. found 4 chalcone derived compounds from Cleistocalyx operculatus exhibited stronger NA inhibitory activities than flavanones, dihydroflavonols and isoflavones. These 4 chalcones had inhibitory effect against NAs from the wild-type and oseltamivir-resistant H1N1 influenza virus (H274Y mutant). The 4 chalcones inhibited N1 in noncompetitive manner and they could protect MDCK cells with a strong protective effect for viral infection and low toxicity to the host MDCK cells.47 In another report, they found 8 chalcone derived compounds from Glycyrrhiza inflata could inhibit NA of influenza virus (H1N1 and H9N2). The 8 chalcones were all noncompetitive inhibitions.48 SA is a chalcone glycoside. Our results showed that SA inhibited N1 and N2 in a noncompetitive manner. We determined the binding site of SA using molecular simulation (Fig. 8), and the result is in agreement with enzyme kinetics.
Some recent studies showed that the inhibitory effect of the anti-influenza drug bind to the catalytic site could be enhanced by another compound binding in the non-catalytic site. For example, the inhibition of on NAs (H9N2, H1N1, and NA mutants) by oseltamivir in the presence of ehinantin binding with NA in a non-catalytic site was strengthened by several folds.46 Because of binding in non-catalytic site, it is likely that SA may have potential to enhance the inhibitory effect of oseltamivir carboxylate against the oseltamivir-resistant mutant.
In contrast to the widely developed anti-influenza drugs, SA interact with NA at a non-catalytic site and it inhibit NA in micromolar range. It protects cell from viral damage and hinders virus replication in vitro. A synergistic antiviral effect was acquired when SA was administered in combination with oseltamivir carboxylate against the neuraminidase activities of N1 and N2 (Fig. 10). Median-effect curves and conservative isobologram analysis also indicated that the SA-oseltamivir carboxylate combination treatment was synergistic, with CI values in the 0.49–0.52 against N1 and 0.56–0.88 against N2 (Fig. 11). Chalcones have been reported to inhibit NA of H1N1, in a noncompetitive manner and enhance the inhibitory effect of oseltamivir.48 SA is a chalcone glycoside derivative. Oseltamivir located in the center of the active site of NAs while SA bound at the backside of NAs (Fig. 7). The different binding to the NA resulted different mode of action. It is most likely that the SA binding caused a change of NA conformation, which lead to a reduced activity of NA and a more efficient inhibition of NA by oseltamivir, thus resulted as synergism of inhibition. SA and oseltamivir carboxylate combination treatment were both synergistic against N1 and N2.
These findings suggest that SA exerts synergistic antiviral activity when using in combination with oseltamivir against NAs of H1N1 and H3N2. It is worthwhile to further test whether the synergism of SA and oseltamivir can overcome the drug resistance caused by the NA mutations and investigate the treatment strategy to add SA to oseltamivir as effective anti-influenza therapeutics.
5 Conclusions
We identified SA from a Chinese medicinal herbal compound library as one of neuraminidase inhibitor. It showed a significant reduction for viral replication in MDCK cells against H1N1 and H3N2 influenza virus. The type of inhibition between SA and N1 or N2 was noncompetitive by the enzyme kinetic tests. The interaction of SA with N1 and N2 was simulated through molecular simulation and docking, which showed that SA bound in the non-active site of N1 and N2. Combination therapy studies were undertaken to investigate the effect of SA when administered in combination with oseltamivir. The results showed that SA-oseltamivir combination treatment resulted in synergism against N1 or N2. These results suggest that an herbal formulation containing SA may serve as an effective supplementary strategy to currently available anti-influenza therapeutics.Appendix
| ID no. | Name | Molecular weight |
|---|---|---|
| 110759 | Bornyl acetate | 196.286 |
| 110776 | Venillic acid | 168.147 |
| 11688 | Natural borneol | 154.249 |
| 111512 | Isoborneol | 154.249 |
| 110881 | Borneol | 154.2 |
| 110809 | Protocatechuic acid | 154.12 |
| 100815 | Curcumol | 236.35 |
| 110728 | Menthol | 198.302 |
| 110845 | Salicylic acid | 138.121 |
| 110792 | Butyl-phydroxybenzoate | 216.209 |
| 110831 | Gallic acid 1 | 70.12 |
| 110747 | Camphor | 152.233 |
| 110885 | Caffeic acid | 180.157 |
| 111735 | Liquidambaric acid | 440.65 |
| 100528 | Guaifenesin | 198.216 |
| 110808 | Catanol | 362.329 |
| 111541 | Fumaric acid | 116.072 |
| 110810 | Protocatechuic aldehyde | 179.5 |
| 110812 | E-10-Hydroxy-3-decylennic acid | 186.25 |
| 110708 | Paeonol | 166.174 |
| 110755 | Ursodeoxycholic acid | 392.572 |
| 110851 | β-Sitosterol | 414 |
| 110816 | Sodium tauroursodeoxycholate | 392.58 |
| 110815 | Sodium Taurocholate | 537.686 |
| 110806 | Chenodesoxuchalic acid | 392.572 |
| 110803 | Cinobufagin | 442.545 |
| 110744 | Sarsasapogenin | 416.636 |
| 110846 | Sodium taurochenodeoxycholic | 522.694 |
| 110724 | Deoxycholic acid | 392.572 |
| 110718 | Recibufogenin | 384.508 |
| 111638 | Ecdysterone | 480.634 |
| 111731 | Fraxetin | 208 |
| 109711 | Columbianadin | 328.36 |
| 110741 | Aesculetin | 178.142 |
| 110837 | Isofraxidin | 222.194 |
| 110822 | Osthol | 244.29 |
| 110900 | Daphnetin | 178.142 |
| 110826 | Imperatorin | 270.28 |
| 110827 | Isoimperatorin | 270.28 |
| 111511 | Scoparone | 206.195 |
| 42096 | 6,7-Dimethoxyl-coumarin | 206.19 |
| 110723 | Glycyrrhetinic acid | 470.64 |
| 110738 | Isopsoralen | 186.16 |
| 111513 | 7-Methoxy coumarin | 176.17 |
| 111741 | 6-Hydroxy-7-methoxycoumarin | 192 |
| 111711 | Praeruptorin A | 386 |
| 110836 | Sesamine | 354.353 |
| 110835 | Ethyl-p-methoxycinnamate | 206.238 |
| 111568 | Linderane | 260.285 |
| 110854 | Dehydroandrographolide | 332.434 |
| 110797 | Andrographolide | 350.449 |
| 110865 | Bilobalide | 326.299 |
| 111747 | Protopanaxadiol | 460.6 |
| 111655 | Andrographolidi natril bisulfis | 262.13 |
| 110701 | Panaxadiol | 460.732 |
| 111664 | Epifriedelanol | 428.733 |
| 111525 | Dehydrocostuslactone | 230.302 |
| 110760 | Alantolactone | 232.318 |
| 110800 | Evodin | 470.512 |
| 110702 | Panaxatriol | 476.731 |
| 100200 | Artesunate | 384.43 |
| 110761 | Isoalantolactone | 232.318 |
| 110863 | Ginkgolide B | 424.399 |
| 110862 | Ginkgolide A | 408.399 |
| 110864 | Ginkgolide C | 440.398 |
| 1566 | Triptolide | 360.401 |
| 111665 | Germacrone | 218.335 |
| 111518 | Cucurbitacin IIa | 574.702 |
| 110742 | Ursolic acid | 456.7 |
| 110753 | Chlorogenic acid | 354.309 |
| 110880 | Pseudolaric acid | 430.491 |
| 100271 | Artemether | 298.375 |
| 111663 | Friedelin | 426.717 |
| 111517 | Gossypol | 518.554 |
| 111581 | Shionone | 426.717 |
| 1539 | Diosgenin | 414.621 |
| 110709 | Oleanolic acid | 456.7 |
| 111524 | Costunolide | 232.33 |
| 110787 | Aloin | 418.394 |
| 111575 | Polydatin | 390.384 |
| 110794 | Rhaponticin | 420.41 |
| 1110893 | Madecassicside | 975.121 |
| 111582 | Cynanchagenin | 500.584 |
| 110887 | Uridine | 244.201 |
| 111574 | Syringin | 372.367 |
| 111590 | Chonglou saponin I | 855.021 |
| 110892 | Asiaticoside | 959.122 |
| 111536 | Tubeimiside I | 1319.43 |
| 110819 | Arctiin | 534.552 |
| 111571 | Isorhamnetin-3-O-nehesperidin | 624.56 |
| 110745 | Notoginsenoside R1 | 919.101 |
| 111523 | 5-o-Methyl visammioside | 452.453 |
| 111685 | Asperosaponin VI | 636.856 |
| 110820 | Amygdalin | 457.429 |
| 110841 | Pseuoginsenoside F11 | 801.013 |
| 111640 | Loganin | 390.382 |
| 111515 | Stevioside | 804.872 |
| 110844 | 2,3,5,4′-Tetrahedroxystilbene-2-O-D-glucoside | 406.383 |
| 111538 | Quercitroside | 448.377 |
| 111522 | prim-O-glucosylcimifugin | 468.451 |
| 111528 | Buddleoside | 592.55 |
| 111553 | Ginserg stem and leaf faponins | — |
| 110778 | Saikosaponin D | 780.982 |
| 110770 | Gentiopicrin | 356.325 |
| 110777 | Saikosaponin A | 780.982 |
| 110736 | Paeoniflorin | 480.462 |
| 111530 | Acteoside | 624.587 |
| 110785 | Swertiamain | 374.34 |
| 111610 | Liguiritin | 418.394 |
| 110821 | Forsythin | 534.552 |
| 110818 | Salidroside | 300.304 |
| 110781 | Astragaloside | 784.97 |
| 110782 | Clinodiside A | 959.125 |
| 110754 | Ginsenoside Re | 947.154 |
| 111580 | Methyl Hesperidin | 624.587 |
| 111592 | Chonglou saponin VI | 738.906 |
| 110870 | Sanqi total saponins | 872.0 |
| 111537 | Pinoresinol diglucoside | 682.67 |
| 110891 | Huangshanyao saponins extract | — |
| 111596 | Picroside-II | 512.461 |
| 111607 | Mangiferin | 422.34 |
| 111695 | Sophoricoside | 432.37 |
| 111670 | Echinacoside | 786 |
| 111714 | Sec-o-glucosylhamaudol | 438 |
| 111745 | Picfeltrarraenin IA | 762.9 |
| 111748 | Ginsenoside Rh2 | 622.6 |
| 111742 | Sweroside | 358.34 |
| 111727 | Picroside I | 492 |
| 111738 | Daidzin | 416 |
| 111719 | Ginsenoside Rf | 801 |
| 111686 | Ginsenoside Rb3 | 1079 |
| 111629 | 2′′-O-galloylhyperin | 616.48 |
| 110871 | Sanqi stem and leaf saponins | — |
| 111593 | Chonglou saponin VII | 1049.2 |
| 100080 | Rutin | 664.563 |
| 110703 | Ginsenoside RgI | 1602.03 |
| 110749 | Geniposide | 388.366 |
| 110715 | Baicalin | 446.361 |
| 110721 | Hesperidin | 610.561 |
| 110734 | Jujuboside A | 1207.35 |
| 111573 | Typhaneoside | 770.685 |
| 110722 | Naringin | 580.535 |
| 110740 | Ginsenoside Rb1 | 1109.29 |
| 110804 | Ginsenoside Rg3 | 785.013 |
| 111668 | Rhamnosylvitexin | 578.519 |
| 110737 | Icariin | 676.662 |
| 111521 | Hyperoside | 464.376 |
| 110842 | Scutellarin | 492.473 |
| 110879 | Adenosine | 267.242 |
| 111508 | D-Xylose | 150.13 |
| 111506 | Arabinose | 150.131 |
| 110833 | Glucose anhydrous | 180.16 |
| 111504 | D-Fructose | 180.156 |
| 111507 | Sucrose | 342.296 |
| 110795 | Aloe-emodin | 270.237 |
| 110758 | Physcion | 284.263 |
| 110756 | Emodin | 270.237 |
| 110829 | 1,9-Dihydroxyanthraquinone | 240.211 |
| 110769 | Shikonin | 288.295 |
| 110852 | Cryptonshinone | 296.36 |
| 111605 | Sulfotanshinone sodium | 396 |
| 110796 | Chrysophanol | 254.237 |
| 110766 | Tanshinonel IIA | 702.616 |
| 110867 | Tanshinonel I | 294.34 |
| 100228 | Menadiol acetate | 258.269 |
| 110884 | Mollugin | 284.307 |
| 110757 | Rhein | 284.22 |
| 111514 | Wogonin | 284.263 |
| 110856 | Silymarin | 482.436 |
| 110762 | Alpinetin | 270.28 |
| 111502 | Daidzein | 254.237 |
| 111554 | Vitexicarpin | 374.3 |
| 111595 | Baicalein | 270.237 |
| 111555 | Lysionotin | 344.315 |
| 110860 | Isorhanetin | 316.262 |
| 110861 | Kaempferol | 286.236 |
| 111557 | Irisflorentin | 386.352 |
| 110850 | Farrerol | 300.306 |
| 100081 | Quercetin | 302.236 |
| 110752 | Puerarin | 416.378 |
| 110763 | Cardamonin | 270.28 |
| 110772 | Patchoulialcohol | 222.366 |
| 111701 | Chrysin | 254 |
| 111520 | Luteolin | 286.23 |
| 110872 | Arginine | 174.201 |
| 111578 | L-Hydroxyproline | 131.13 |
| 110735 | Glycine | 75.067 |
| 110875 | L-Citrulline | 175.186 |
| 110890 | Histidine | 155.155 |
| 110876 | Leucine | 131.173 |
| 110873 | DL-Aminopropionic acid | 89.09 |
| 110889 | Threomine | 119.18 |
| 110874 | DL-Aminovaleic acid | 117.15 |
| 111533 | Homoharringtonine | 545.621 |
| 110779 | Papaverine hydrochloride | 375.846 |
| 100747 | Vincamine | 354.443 |
| 110894 | Betaine | 117.146 |
| 111666 | Dihydrocapsaicin | 307.428 |
| 110886 | Adenine | 135.127 |
| 110775 | Piperine | 285.338 |
| 110784 | Sophoridine | 248.364 |
| 110780 | Oxymatrine | 264.363 |
| 100249 | Raceanisodamine | 305.38 |
| 110805 | Matrine | 248.364 |
| 100121 | Theophylline | 180.164 |
| 100243 | Huperzine A | 258.316 |
| 110774 | Sinomenine | 329.39 |
| 100526 | Hydroxycamptothecin | 364.352 |
| 110801 | Rutaecarpine | 287.315 |
| 100476 | Methscopolamine bromide | 398.292 |
| 111654 | Dictamnine | 199.205 |
| 111652 | Oxysophocarpine | 262.347 |
| 110802 | Evodiamine | 303.358 |
| 100382 | Paclitaxel | 853.906 |
| 110733 | Jatrorrhizine hydrochloride | 374.887 |
| 111718 | Chelerythrine | 348.36 |
| 111651 | Cavidine | 353.412 |
| 111647 | Cepharanthin | 606.707 |
| 100270 | Nimodipine | 418.44 |
| 100336 | Tinidazole | 247.273 |
| 110839 | Capsaici | 305.412 |
| 100135 | Glibenclamide | 494.004 |
| 100412 | Sulfagunidine | 232.26 |
| 110732 | Palmatine hydrochloride | 391.888 |
| 110717 | Indirabin | 262.263 |
| 100547 | Lappaconitine adhesive | 584.7 |
| 110746 | Aristolochic acid A | 341.272 |
| 11071 | Indigotin | 262.27 |
| 110726 | Tetrahydropalmatine | 355.428 |
| 110895 | Bullatine A | 343.503 |
| 100250 | Isosorbide dinitrate | 236.136 |
| 110750 | Peimine | 431.651 |
| 110751 | Peinine | 429.64 |
| 111667 | Dehydrocavidine | 353 |
| 110711 | Tetrandrine | 622.75 |
| 110793 | Fangchinoline | 608.723 |
| 110720 | Aconitine | 645.737 |
| 110883 | Trigonelline | 137.136 |
| 111566 | Nuciferine | 295.376 |
| 110712 | Stachydrine hydrochloride | 179.644 |
| 110853 | Protopine | 353.369 |
| 100465 | Methylcantharidinimde | 209 |
| 111608 | Convza Blinii extract | — |
| 111501 | Allantoin | 158.116 |
| 110807 | Gastrodin | 286.278 |
| 110704 | Cordycepin | 251.242 |
| 110866 | Ginkgo biloba extract | — |
| 100385 | Hilieidum | 284.26 |
| 110877 | Catechin | 290.268 |
| 110843 | Diphenyl | 154 |
| 110878 | Epicatechin | 290.268 |
| 110811 | Dracohodin perochlorate | 366.75 |
| 110896 | Succinic acid | 118.088 |
| 110817 | Chuangxiongzine hydrochloride | 208.69 |
| 100247 | Sodium houttuyfonate | 302.36 |
| 100845 | Ligustrazine phosphate | 242 |
| 110855 | Sodium danshensu | 220 |
| 110857 | Schisandrin | 432.5 |
| 110859 | Hupehenine | 415.345 |
| 110888 | Cycloyirobuxinum D | 402.364 |
| 111535 | Resveratrol | 390 |
| 111637 | Safflomin A | 612.533 |
| 111689 | β,β-Dimethylacrylalkannin | 370.39 |
| 111696 | Liensinine perchlorate | 610.74 |
| 111699 | Galangin | 270.25 |
| 111700 | Fraxinellon | 232 |
| 111702 | Sinapine cyanide sulfonate | 368 |
| 111703 | Formonontin | 268.27 |
| 111712 | RaddeaninA | 896 |
| 111717 | 1,3-O-dicaffeoylquinic acid | 516.45 |
| 111721 | Oridonin | 364.43 |
| 111744 | 1-Hydroxy-3,4,5-trimethoxyxanthone | 302 |
| 110765 | γ-Schisandrin | 400.46 |
| 110764 | Deoxyschizandrin | 416.507 |
| 110882 | Magnoshinin | 414.491 |
| 110773 | Ferulic acid | 194.18 |
| 111532 | Bergenin | 328.271 |
| 111645 | Podophyllotoxin | 414.405 |
| 111529 | Schisantherin | 536.57 |
| 111660 | Loureirin A | 286.322 |
| 111558 | Loureirin B | 316.35 |
| 111539 | Toddalolatone | 308.326 |
| 110729 | Magnolol | 266.334 |
| 110730 | Honokiol | 266.334 |
| 111561 | Fargesin | 370.396 |
| 100202 | Artemisinin | 282.332 |
| 111559 | Cochinchinenin C | 514.566 |
| 110823 | Curcumin | 368.38 |
| 111594 | Total ginkgolic acid | 346.49 |
Acknowledgements
The authors gratefully acknowledge financial support from the National Natural Science Foundation of China (No. 31170742 and No. 31370742) and Fund from Science and Technology Department of Jilin Province (No. 20130206066YY).Notes and references
- A. E. Fiore, D. K. Shay, K. Broder, J. K. Iskander, T. M. Uyeki, G. Mootrey, J. S. Bresee and N. J. Cox, MMWR Recommendations & Reports–Morbidity & Mortality Weekly Report Recommendations & Reports, 2009, vol. 58, pp. 1–52.
- K. R. Short, P. C. Reading, L. E. Brown, J. Pedersen, B. Gilbertson, E. R. Job, K. M. Edenborough, M. N. Habets, r. A. Zome, P. W. Hermans, D. A. Diavatopoulos and O. L. Wijburg, Infect. Immun., 2013, 81, 645–652 CrossRef CAS PubMed.
- R. Gao, B. Cao, Y. Hu, Z. Feng, D. Wang, W. Hu, J. Chen, Z. Jie, H. Qiu, K. Xu, X. Xu, H. Lu, W. Zhu, Z. Gao, N. Xiang, Y. Shen, Z. He, Y. Gu, Z. Zhang, Y. Yang, X. Zhao, L. Zhou, X. Li, S. Zou, Y. Zhang, X. Li, L. Yang, J. Guo, J. Dong, Q. Li, L. Dong, Y. Zhu, T. Bai, S. Wang, P. Hao, W. Yang, Y. Zhang, J. Han, H. Yu, D. Li, G. F. Gao, G. Wu, Y. Wang, Z. Yuan and Y. Shu, N. Engl. J. Med., 2013, 368, 1888–1897 CrossRef CAS PubMed.
- N. L. La Gruta, K. Kedzierska, J. Stambas and P. C. Doherty, Immunol. Cell Biol., 2007, 85, 85–92 CrossRef PubMed.
- H. N. Gao, H. Z. Lu, B. Cao, B. Du, H. Shang, J. H. Gan, S. H. Lu, Y. D. Yang, Q. Fang, Y. Z. Shen, X. M. Xi, Q. Gu, X. M. Zhou, H. P. Qu, Z. Yan, F. M. Li, W. Zhao, Z. C. Gao, G. F. Wang, L. X. Ruan, W. H. Wang, J. Ye, H. F. Cao, X. W. Li, W. H. Zhang, X. C. Fang, J. He, W. F. Liang, J. Xie, M. Zeng, X. Z. Wu, J. Li, Q. Xia, Z. C. Jin, Q. Chen, C. Tang, Z. Y. Zhang, B. M. Hou, Z. X. Feng, J. F. Sheng, N. S. Zhong and L. J. Li, N. Engl. J. Med., 2013, 368, 2277–2285 CrossRef CAS PubMed.
- Q. Li, L. Zhou, M. Zhou, Z. Chen, F. Li, H. Wu, N. Xiang, E. Chen, F. Tang and D. Wang, N. Engl. J. Med., 2013, 14, 1668–1677 Search PubMed.
- X. Wang, W. Jia, A. Zhao and X. Wang, Phytother. Res., 2006, 20, 335–341 CrossRef CAS PubMed.
- R. J. Russell, L. F. Haire, D. J. Stevens, P. J. Collins, Y. P. Lin, G. M. Blackburn, A. J. Hay, S. J. Gamblin and J. J. Skehel, Nature, 2006, 443, 45–49 CrossRef CAS PubMed.
- T. Betakova, Curr. Pharm. Des., 2007, 13, 3231–3235 CrossRef CAS PubMed.
- M. N. Matrosovich, T. Y. Matrosovich, T. Gray, N. A. Roberts and H.-D. Klenk, J. Virol., 2004, 78, 12665–12667 CrossRef CAS PubMed; K. Das, J. M. Aramini, L. C. Ma, R. M. Krug and E. Arnold, Nat. Struct. Mol. Biol., 2010, 17, 530–538 Search PubMed.
- L. M. Surhone, M. T. Timpledon and S. F. Marseken, Rimantadine, Betascript Publishing, 2010 Search PubMed.
- P. K. Cheng, A. P. To, T. W. Leung, P. C. Leung, C. W. Lee and W. W. Lim, Emerging Infect. Dis., 2010, 16, 155 CrossRef PubMed.
- A. C. Hurt, J. Ernest, Y. M. Deng, P. Iannello, T. G. Besselaar, C. Birch, P. Buchy, M. Chittaganpitch, S. C. Chiu, D. Dwyer, A. Guigon, B. Harrower, I. P. Kei, T. Kok, C. Lin, K. McPhie, A. Mohd, R. Olveda, T. Panayotou, W. Rawlinson, L. Scott, D. Smith, H. D'Souza, N. Komadina, R. Shaw, A. Kelso and I. G. Barr, Antiviral Res., 2009, 83, 90–93 CrossRef CAS PubMed.
- J. Boon-Long, K. Ikuta, G.-R. Bai and N. Takeda, Bulletin of the Department of Medical Sciences, 2014, vol. 55, pp. 117–127 Search PubMed.
- G. Orozovic, K. Orozovic, J. D. Jarhult and B. Olsen, PLoS One, 2014, 9, e89306 Search PubMed.
- A. Meijer, A. Lackenby, O. Hungnes, B. Lina, S. van der Werf, B. Schweiger, M. Opp, J. Paget, J. van de Kassteele and A. Hay, Emerging Infect. Dis., 2009, 15, 552 CrossRef PubMed.
- M. Samson, A. Pizzorno, Y. Abed and G. Boivin, Antiviral Res., 2013, 98, 174–185 CrossRef CAS PubMed.
- T. Baranovich, R. Saito, Y. Suzuki, H. Zaraket, C. Dapat, I. Caperig-Dapat, T. Oguma, I. I. Shabana, T. Saito and H. Suzuki, J. Clin. Virol., 2010, 47, 23–28 CrossRef CAS PubMed.
- V. P. Mishin, K. Sleeman, M. Levine, P. J. Carney, J. Stevens and L. V. Gubareva, Antiviral Res., 2014, 101, 93–96 CrossRef CAS PubMed.
- Q. Liu, J. Ma, D. R. Strayer, W. M. Mitchell, W. A. Carter, W. Ma and J. A. Richt, Expert Rev. Anti-Infect. Ther., 2014, 12, 165–169 CrossRef CAS PubMed.
- J. Zhou, D. Wang, R. Gao, B. Zhao, J. Song, X. Qi, Y. Zhang, Y. Shi, L. Yang, W. Zhu, T. Bai, K. Qin, Y. Lan, S. Zou, J. Guo, J. Dong, L. Dong, Y. Zhang, H. Wei, X. Li, J. Lu, L. Liu, X. Zhao, X. Li, W. Huang, L. Wen, H. Bo, L. Xin, Y. Chen, C. Xu, Y. Pei, Y. Yang, X. Zhang, S. Wang, Z. Feng, J. Han, W. Yang, G. F. Gao, G. Wu, D. Li, Y. Wang and Y. Shu, Nature, 2013, 499, 500–503 CrossRef CAS PubMed.
- R. Hai, M. Schmolke, V. H. Leyva-Grado, R. R. Thangavel, I. Margine, E. L. Jaffe, F. Krammer, A. Solorzano, A. Garcia-Sastre, P. Palese and N. M. Bouvier, Nat. Commun., 2013, 4 Search PubMed.
- K. Sleeman, V. M. Mishin VPDeyde, Y. Furuta, A. I. Klimov and L. V. Gubareva, Antimicrob. Agents Chemother., 2010, 54, 2517–2524 CrossRef CAS PubMed.
- K. Das, J. M. Aramini, L. C. Ma, R. M. Krug and E. Arnold, Nat. Struct. Mol. Biol., 2010, 17, 530–538 CAS.
- A. Dias, D. Bouvier, T. Crépin, A. A. Mccarthy, D. J. Hart, F. Baudin, S. Cusack and R. W. H. Ruigrok, Nature, 2009, 458, 914–918, DOI:2010.1038/nature07745.
- L. Tian, Z. Wang, H. Wu, S. Wang, Y. Wang, Y. Wang, J. Xu, L. Wang, F. Qi and M. Fang, J. Ethnopharmacol., 2011, 137, 534–542 CrossRef PubMed.
- H. Ge, Y. F. Wang, J. Xu, Q. Gu, H. B. Liu, P. G. Xiao, J. Zhou, Y. Liu, Z. Yang and H. Su, Nat. Prod. Rep., 2010, 27, 1758–1780 RSC.
- J. Y. Ho, H. W. Chang, C. F. Lin, C. J. Liu, C. F. Hsieh and J. T. Horng, Viruses, 2014, 6, 1861–1875 CrossRef PubMed.
- N. Oyuntsetseg, M. A. Khasnatinov, P. Molor-Erdene, J. Oyunbileg, A. V. Liapunov, G. A. Danchinova, S. Oldokh, J. Baigalmaa and C. Chimedragchaa, BMC Complementary Altern. Med., 2014, 14, 235 CrossRef PubMed.
- X. Wang, X. Xu, Y. Li, X. Li, W. Tao, B. Li, Y. Wang and L. Yang, Integrative biology : quantitative biosciences from nano to macro, 2013, vol. 5, pp. 351–371 Search PubMed.
- D. A. Gschwend, A. C. Good and I. D. Kuntz, J. Mol. Recognit., 1996, 9, 175–186 CrossRef CAS PubMed.
- J. H. An, D. C. W. Lee, A. H. Y. Law, C. L. H. Yang, L. L. M. Poon, A. S. Y. Lau and S. J. M. Jones, J. Med. Chem., 2009, 52, 2667–2672 CrossRef CAS PubMed.
- U. S. Tambunan, A. A. Parikesit, Y. Dephinto and F. R. Sipahutar, Acta Pharm., 2014, 64, 157–172 CrossRef CAS PubMed.
- Natural Products Database (NPD) http://wiki.compbio.ucsf.edu/wiki/index/php/Natural_products _database. 2011.
- Y. Wang, D. Wu, D. Yu, Z. Wang, L. Tian, Y. Wang, W. Han and X. Fang, J. Mol. Model., 2012, 18, 3445–3453 CrossRef CAS PubMed.
- P. Ramanujam, W. Tan, S. Nathan and K. Yusoff, Arch. Virol., 2002, 147, 981–993 CrossRef CAS PubMed.
- M. Potier, L. Mameli, M. Belisle, L. Dallaire and S. B. Melancon, Anal. Biochem., 1979, 94, 287–296 CrossRef CAS PubMed.
- D. F. Smee, J. H. Huffman, A. C. Morrison, D. L. Barnard and R. W. Sidwell, Antimicrob. Agents Chemother., 2001, 45, 743–748 CrossRef CAS PubMed.
- G. M. Morris, D. S. Goodsell, R. S. Halliday, R. Huey, W. E. Hart, R. K. Belew and A. J. Olson, J. Comput. Chem., 1998, 19, 1639–1662 CrossRef CAS.
- T. C. Chou, M. Hayball and L. Cw., CalcuSyn-windows software for dose-effect analysis and synergism/antagonism quantification, and user's manual, Biosoft, Cambridge, United Kingdom, 1996 Search PubMed.
- T. C. Chou and P. Talalay, Eur. J. Biochem., 1981, 115, 207–216 CrossRef CAS PubMed.
- D. P. Nayak and U. Reich, J. Virol. Methods, 2004, 122, 9–15 CrossRef CAS PubMed.
- U. Grienke, M. Schmidtke, J. Kirchmair, K. Pfarr, P. Wutzler, R. Dürrwald, G. Wolber, K. R. Liedl, H. Stuppner and J. M. Rollinger, J. Med. Chem., 2009, 53, 778–786 CrossRef PubMed.
- S. Lu, Y. Qiao, P. Xiao and X. Tan, China J. Chin. Mater. Med., 2005, 30, 998–1001 Search PubMed.
- X.-y. Pu, J.-p. Liang, X.-h. Wang, T. Xu, L.-y. Hua, R.-f. Shang, Y. Liu and Y.-m. Xing, Virol. Sin., 2009, 24, 19–27 CrossRef.
- K. Hayashi, N. Imanishi, Y. Kashiwayama, A. Kawano, K. Terasawa, Y. Shimada and H. Ochiai, Antiviral Res., 2007, 74, 1–8 CrossRef CAS PubMed.
- T. T. Dao, B. T. Tung, P. H. Nguyen, P. T. Thuong, S. S. Yoo, E. H. Kim, S. K. Kim and W. K. Oh, J. Nat. Prod., 2010, 73, 1636–1642 CrossRef CAS PubMed.
- T. T. Dao, P. H. Nguyen, H. S. Lee, E. Kim, J. Park, S. I. Lim and W. K. Oh, Bioorg. Med. Chem. Lett., 2011, 21, 294–298 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2015 |