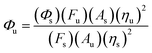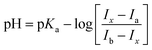A new fluorescent pH probe for acidic conditions†
Xuan Zhangab,
Guang-Jie Songc,
Xiang-Jian Caoab,
Jin-Ting Liua,
Ming-Yu Chenab,
Xiao-Qun Caoc and
Bao-Xiang Zhao*a
aInstitute of Organic Chemistry, School of Chemistry and Chemical Engineering, Shandong University, Jinan 250100, PR China. E-mail: bxzhao@sdu.edu.cn; Fax: +86-531-88564464; Tel: +86-531-88366425
bTaishan College, Shandong University, Jinan 250100, PR China
cTaishan Medical University, Tai'an 271000, PR China
First published on 15th October 2015
Abstract
A new imidazo[1,5-a]pyridine-based pH fluorescent probe has been synthesized and characterized. It is the first time that imidazo[1,5-a]pyridine is used as a fluorescent probe. This probe responds to acidic pH with fast response (within 3 min), high selectivity and sensitivity. It has good reversibility and nearly no interference from common metal ions. The probe is suitable for acidic conditions and can quantitatively detect pH value based on the equilibrium equation, pH = pKa − log[(Ia − Ix)/(Ix − Ib)], with a good linear relationship. Moreover, we verified that the protonation of the 2-nitrogen in the imidazole decreases the electron density on the fused ring by 1H NMR analysis and DFT calculation. We proposed that intramolecular charge transfer and solvation of the probe in aqueous solutions might be the mechanism of the fluorescence enhancement under strongly acidic conditions.
Introduction
Intracellular pH plays a significant role in cellular behaviors and pathological conditions. The fluctuation of pH can affect many cellular behaviors obviously, such as cellular metabolism,1–3 endocytosis, apoptosis, ion transport and enzymatic activity.4–9 Abnormality of cellular pH can cause many cellular functional changes. When pH changes from acidic to neutral, for example, the activities of enzymes in lysosomes decrease. Moreover, serious diseases such as cancer, stroke and Alzheimer can be caused when pH changes dramatically.10–12 So it is important for us to monitor intracellular pH change.It is reported that many detection methods including acid–base indicator titration13 and potentiometric titration14 have been used to measure pH value. But these detection methods are expensive, easily interfered by metal ions and complex environment. Compared with these methods, fluorescent probes have many advantages such as high sensitivity, selectivity, fast response, low cost and convenient operation.15,16 Fluorescent probes are widely used not only in pH detection but also in other analyte detections including metal ions and fluoride ions in the environment.17–22
Lots of pH fluorescent probes have been designed. They can be divided into two categories. Probes belonging to one category respond to neutral pH ranging from 6 to 8.23,24 Others respond to weak acid pH ranging from 4 to 6.25,26 Beyond these two categories very few probes respond to pH value below 4 (ref. 27) and are suitable for extremely acid conditions.28–30
Imidazo[1,5-a]pyridine derivatives exhibited good biological activity and have great potential for developing new drug. Imidazo[1,5-a]pyridine derivatives have a wider range of anti-inflammation activity.31 Up to now, only several ways for synthesizing imidazo[1,5-a]pyridine derivatives have been reported. One main way involved the reaction of 2-(aminomethyl)pyridine with CHCl3 and alkaline hydroxide in the phase transfer catalysis condition.32 The tandem reaction has been successfully used to synthesize imidazo[1,5-a]pyridine derivatives with moderate condition, easy work-up and high yield.33 However, no literature has been reported that imidazo[1,5-a]pyridine derivative can be used as a fluorophore and can be used as a pH probe. As a continuation of our work on developing fluorescent probe for pH,34–38 we found that imidazo[1,5-a]pyridine derivatives have strong fluorescence intensity and high quantum yield. Here, we report an imidazo[1,5-a]pyridine derivative, 3-butyl-1-chloroimidazo[1,5-a]pyridine-7-carboxylic acid, as a new pH probe for acid condition based on intramolecular charge transfer (ICT), which is one of the most important signaling mechanisms in designing fluorescent probes.39,40 This probe with pKa 4.04 can respond to acid pH ranging from 3 to 5 with excellent linear relationship, so it can be used to image living cells because many cells can still be alive in this pH condition.
Materials and methods
Materials
All reagents and solvents were purchased from commercial sources and used without further purification. The solutions of metal ions were prepared from nitrate salts which were dissolved in deionized water. Deionized water was used throughout the process of absorption and fluorescence determination. All samples were prepared at room temperature, shaken for 10 s and rested for 3 h before UV-vis and fluorescence determination. Britton–Robinson (B–R) buffer was prepared with 40 mM acetic, boric acid, and phosphoric acid. Dilute hydrochloric acid or sodium hydroxide was used for tuning pH values.Instruments
Thin-layer chromatography (TLC) involved silica gel 60 F254 plates (Merck KGaA). Melting points were detected on an XD-4 digital micro melting point apparatus. 1H NMR spectra were recorded on a Bruker Avance 400 (400 MHz) spectrometer and 13C NMR spectra were recorded on a Bruker Avance 300 (75 MHz) spectrometer, using DMSO-d6 as solvent and tetramethylsilane (TMS) as an internal standard. HRMS spectra were recorded on a Q-TOF6510 spectrograph (Agilent). Fluorescence measurements were recorded on a Perkin-Elmer LS-55 luminescence spectrophotometer and UV-vis spectra were recorded on a U-4100 UV-Vis-NIR Spectrometer (Hitachi). The pH measurements were measured by the use of a PHS-3C digital pH-meter (YouKe, Shanghai). The single crystals were measured on a Bruker-AXS CCD single-crystal diffractometer with graphite-monochromated MoKα radiation source (λ = 0.71073 Å). The structure was solved with direct methods using the SHELXS-97 program, and refined on F2 by full-matrix least-squares with the SHELXL-97 package.41 Molecular graphics were designed by using DIAMOND 3.2.42 PLATON program was also used for structure analysis.43The molecular geometries of the probe in neutral and acid solution were optimized using density functional theory and time dependent density functional theory (DFT and TDDFT) at the B3LYP/6-31G level. The solvent effect on molecular geometries is included by means of the polarizable continuum model (PCM). Based on the optimized geometry, the molecular orbitals of the probe in neutral and acid solution were calculated at the same level. All the calculations were performed in Gaussian09 software.
Synthesis
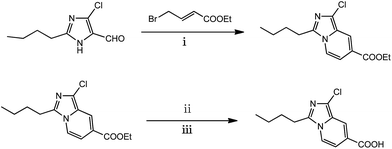
Compound 1 (2-butyl-4-chloro-1H-imidazole-5-carbaldehyde) and ethyl(E)-4-bromobut-2-enoate were achieved commercially. Compound 2 was synthesized as described in the literature.33Compound 2 (2.8 g, 10 mmol) and NaOH (0.8 g, 20 mmol) were dissolved in a mixture of ethanol (20 mL) and water (15 mL). The mixture was stirred under reflux for 6 h. Afterward, the resulting solution was poured into water (200 mL) followed by neutralization with hydrochloric acid (2 mol L−1). The precipitated yellow solid was separated out and washed with water (2 × 5 mL). After dried in an oven at 50 °C, a yellow solid was obtained in 92% yield (2.32 g). mp: 170–172 °C. 1H NMR (DMSO-d6, 400 MHz), δ 13.19 (s, 1H), 8.26 (d, J = 7.6 Hz, 1H), 7.98 (s, 1H), 7.04 (d, J = 7.6 Hz, 1H), 2.99 (t, J = 7.6 Hz, 2H), 1.67–1.77 (m, 2H), 1.36 (dt, J = 15.2, 7.6 Hz, 2H), 0.91 (t, J = 7.6 Hz, 3H). 13CNMR (DMSO-d6, 75 MHz) δ (ppm) 13.59, 21.67, 25.18, 28.49, 139.54 (2C), 119.44, 120.73, 121.08, 122.05, 123.58, 165.73. HRMS (C12H13ClN2O2): calcd [M − H]−: 251.0593; found value: [M − H]− 251.0635.
Fungi culture and imaging
Saccharomyces cerevisiae (S. cerevisiae, a species of yeast used in making wine, baking, and brewing since ancient times) was resuscitated at 30 °C in Yeast Extract Peptone Dextrose (YPD) medium (tryptone 2%, yeast extract 1%, glucose 2%) for 12 h in a table concentrator (ZHI CHU ZQZY-70B, China) at 200 rpm. Then the culture was centrifuged (Thermo Micro 21, German) in 2 mL Eppendorf tubes at 4500 g for 2 min to collect S. cerevisiae cells. The sediment was resuspended with 1 mL B–R buffer at different pH (3, 4, 5), respectively. After that, the tubes were placed in a table concentrator as mentioned above. After 2 h, the probe dissolved in DMSO was added to every tube to make the probe concentration achieve 10 μM and continually incubated for 30 min. Then, smeared on slides and observed by laser confocal microscopy (Carl Zeiss LSM-700, Germany) at the wavelength of 350 nm.Results and discussion
Synthesis of the probe
The general synthetic route of the probe was shown in Scheme 1. The structure of the probe was characterized by 1H NMR, 13C NMR and HRMS spectra, especially, by X-ray single crystal diffraction.Crystal structure
The crystals suitable for X-ray analysis were obtained from the slow evaporation of ethanol solutions of 3-butyl-1-chloro imidazo [1,5-a]pyridine-7-carboxylic acid (the probe) at room temperature. The compound crystallized in the monoclinic space group P21/c (Tables S1 and S2, ESI†). The X-ray analysis confirmed the molecular structure and atom connectivity as illustrated in Fig. 1. The crystal structure of the compound clearly revealed its well-defined geometry due to the rigidity of the fused rings. The five-membered ring [N(1), C(6), N(2), C(4) and C(5)] (Cg1) and six-membered ring [N(2), C(7), C(8), C(2)–C(4)] (Cg2) were almost coplanar with dihedral angle of 0.9(4)°, indicating that they are conjugated.The crystal packing of the compound was dominated by O–H⋯O, C–H⋯N and Cl⋯π interactions (Tables S3 and S4, ESI†). Interestingly, two O1–H1⋯O2 intermolecular hydrogen bonds generated centrosymmetric R22(8) dimmers (Fig. 1(b)), which were linked to each other through C8–H8⋯N1 intermolecular hydrogen bonds and the Cl⋯π stacking interactions, forming a three-dimensional structure.
Spectroscopic properties and optical responses to pH
We studied the spectroscopic properties of the probe. Fig. 2(a) shows that the fluorescence intensity of the probe in the buffer solution is low when the pH value is over 6. But the fluorescence intensity increases dramatically when the pH value goes down from 6 to 2. According to the literature,44 we calculated that the quantum yield (Φ) is 0.399 at pH 3.In the formula Φ and F are the fluorescence quantum yield and the integral area under the emission spectrum (the excitation wavelength 380 nm) respectively. A is absorbance at the excitation wavelength and η is the refractive index of the solution. Subscripts u and s are the unknown and the standard, respectively. Quinine sulphate dehydrate (99.0%) in 0.1 N H2SO4 is used as the main standard.
pKa value of the new probe was determined in Britton–Robison buffer/ethanol (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). In Fig. 2(b), we can see the “Z” shaped coordinate figure where the X axis represents the pH value and the Y axis represents the fluorescence intensity (emission wavelength 480 nm). We can use the following equilibrium from the literature45 to describe the relationship between the pH value and the fluorescence intensity at λem = 480 nm quantitatively.
1). In Fig. 2(b), we can see the “Z” shaped coordinate figure where the X axis represents the pH value and the Y axis represents the fluorescence intensity (emission wavelength 480 nm). We can use the following equilibrium from the literature45 to describe the relationship between the pH value and the fluorescence intensity at λem = 480 nm quantitatively.
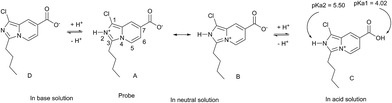
In the equilibrium, Ia = 902.3 and Ib = 571 represent the fluorescence intensity of the probe in its acid (pH = 2) and conjugate base (pH = 10), respectively. According to the equation in the literature,46 we calculate that pKa1 is 4.02 which stems from carboxylic acid. Similarly, pKa2 is calculated to be 5.50 which belongs to ammonium ion (Scheme 2). Considering the fluorescence intensity is very sensitive in the range of pH from 2 to 5, the probe can be used to detect pH value in this range. In Fig. 2(b), we can see that when the pH value is between 3.2 and 4.8, the fluorescence intensity (Y axis) and the pH value (X axis) can be described by a perfect linear regression relationship (R2 = 0.990). In Fig. 3, we get the linear regression relationship between the pH value and “log[(Ia − Ix)/(Ix − Ib)]”. They can be described by the following formula with R2 = 0.998.
| pH = 1.0889X + 4.0254 |
In the formula, X represents “log[(Ia − Ix)/(Ix −Ib)]”. Any sample with pH ranged from 3.2 to 4.8 can be calculated by the formula based on the fluorescence intensity with high preciseness. We believe this probe is a useful pH sensor.
In Fig. 4 the fluorescence intensity between pH 2 and 5 is reversible, which means it is suitable for the detection of a system with a shifty pH value. Moreover, the time course analysis (Fig. 5) shows that the probe can respond to pH less than three minutes in different conditions. Therefore, we can use this probe to get pH value of a sample immediately. Furthermore, the interference from metal ions that are common in biological system is negligible (Fig. 6). Therefore, the probe has a potential to detect the pH value inside living cells. Also when the probe is used to detect complex environment, this feature guarantees the effective use of the probe.
 | ||
Fig. 5 The time course of fluorescence intensity of the probe in solution (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, B−R-EtOH, v/v) with various pH value by continuous irradiation. 1, B−R-EtOH, v/v) with various pH value by continuous irradiation. | ||
The mechanism detecting the strong acidity
Density functional theory calculations
To gain insight into the ICT mechanism, density functional theory (DFT) based theoretical calculations was carried out. The carboxyl group of the probe could be deprotonated and exists as the carboxylate anion at about neutral pH, while a nitrogen atom could be protonated and exists as the ammonium cation. Thus, the probe exists in aqueous solution primarily in the form of a dipolar ion, or zwitterion, i.e. A or B, which are resonance structures. However, the probe (A or B form) is protonated to form C (AH+ or BH+) in acidic aqueous solutions (Scheme 2). As shown in Fig. 7, for the probe both in neutral and acid condition, the S1–S0 transitions (L–H) were electron density redistributions, thus there were ICT processes in the two cases. Furthermore, the energy gap between the LUMO and HOMO of the protonated probe was lower than that of the probe in neutral condition, in good agreement with the red shift in the emission spectra observed. The absorption spectra of the probe showed a red shift (Fig. S2, ESI†).Fluorescence imaging in S. cerevisiae
To verify the potential biological application of the probe, we detected the acidic condition in S. cerevisiae. We used buffer with pH 3, 4, 5 to incubate S. cerevisiae. We added the probe to measure the pH value and imaging it. From image captured with the microscope (Fig. 8), it is clear that from pH 3 to 5, the fluorescence intensity decreases. Furthermore, fluorescence intensity quantitation was analyzed by the ImageJ (Fig. 8(b)). Based on the fluorescence intensity extracted from the images, we could calculate a specific pH to an image of S. cerevisiae according to the equilibrium equation,| pH = pKa − log[(Ia − Ix)/(Ix − Ib)] |
 | ||
| Fig. 8 (a) Imaging in acidity in S. cerevisiae cells with the probe (5 μM) incubated with pH 3, 4, 5. (b) The fluorescence intensity quantitation was analyzed by the Image J. | ||
Conclusion
In summary, a new pH fluorescent probe based on imidazo[1,5-a]pyridine derivative was developed to detect pH value in acid conditions. It is the first time imidazo[1,5-a]pyridine is used as a fluorophore. With the help of the “Z” shaped plot we can detect pH value quantitatively according to equilibrium equation pH = pKa − log[(Ia − Ix)/(Ix − Ib)]. This probe has good selectivity, sensitivity, excellent reversibility and extremely short response time so it can be used to monitor real time pH value. Also it can resist interfering ions greatly so we can detect pH value in complicated environment. More importantly, the cell assay proved that the probe had good effect in imaging acidity in S. cerevisiae and could be able to detect the pH value via analyzing the fluorescence intensity extracted from the images. Furthermore, the detection mechanism has been verified. The probe existed in the zwitterion form in neutral condition and was protonated in acid condition to cause ICT change. All in all, the probe has good ability in detecting low pH conditions in solutions and we believe it will be great beneficial to study chemical and biological systems.Acknowledgements
This study is supported by the Natural Science Foundation of Shandong Province (ZR2014BM004) and Foundation of Talent Training of Fundamental Subject of China (Grant No.: J1103314).Notes and references
- M. Montgomery, T. Boyd, C. Osburn, R. Plummer, S. Masutani and R. Coffin, Desalination, 2009, 249, 861–864 CrossRef CAS PubMed.
- T. Otomo, K. Higaki, E. Nanba, K. Ozono and N. Sakai, J. Biol. Chem., 2011, 286, 35283–35290 CrossRef CAS PubMed.
- X. Wang, Z. Pi, W. Liu, Y. Zhao and S. Liu, Chin. J. Chem., 2010, 28, 2494–2500 CrossRef CAS PubMed.
- D. Pérez-Sala, D. Collado-Escobar and F. Mollinedo, J. Biol. Chem., 1995, 270, 6235–6242 CrossRef PubMed.
- J. R. Casey, S. Grinstein and J. Orlowski, Nat. Rev. Mol. Cell Biol., 2010, 11, 50–61 CrossRef CAS PubMed.
- C. He, K. Lu and W. Lin, J. Am. Chem. Soc., 2014, 136, 12253–12256 CrossRef CAS PubMed.
- Y. Li, Y. Wang, S. Yang, Y. Zhao, L. Yuan, J. Zheng and R. Yang, Anal. Chem., 2015, 87, 2495–2503 CrossRef CAS PubMed.
- M. Lakadamyali, M. J. Rust, H. P. Babcock and X. Zhuang, Proc. Natl. Acad. Sci. U. S. A., 2003, 100, 9280–9285 CrossRef CAS PubMed.
- M. H. Lee, J. H. Han, J. H. Lee, N. Park, R. Kumar, C. Kang and J. S. Kim, Angew. Chem., Int. Ed., 2013, 52, 6206–6209 CrossRef CAS PubMed.
- T. Davies, R. Fine, R. Johnson, C. Levesque, W. Rathbun, K. Seetoo, S. Smith, G. Strohmeier, L. Volicer and L. Delva, Biochem. Biophys. Res. Commun., 1993, 194, 537–543 CrossRef CAS.
- G. K. Vegesna, J. Janjanam, J. Bi, F. T. Luo, J. Zhang, C. Olds, A. Tiwari and H. Liu, J. Mater. Chem. B, 2014, 2, 4500–4508 RSC.
- F. Yu, Y. Wang, W. Zhu, Y. Huang, M. Yang, H. Ai and Z. Lu, RSC. Adv., 2014, 4, 36849–36853 RSC.
- Z. Simi, Z. D. Stani and M. Antonijevi, J. Braz. Chem. Soc., 2011, 22, 709–717 CrossRef.
- N. Balazs and P. Sipos, Carbohydr. Res., 2007, 342, 124–130 CrossRef CAS PubMed.
- T. Hasegawa, Y. Kondo, Y. Koizumi, T. Sugiyama, A. Takeda, S. Ito and F. Hamada, Bioorg. Med. Chem. Lett., 2009, 17, 6015–6019 CrossRef CAS PubMed.
- M. Zhang, S. Zheng, L. Ma, M. Zhao, L. Deng, L. Yang and L. J. Ma, Spectrochim. Acta, Part A, 2014, 124, 682–686 CrossRef CAS PubMed.
- D. Jeyanthi, M. Iniya, K. Krishnaveni and D. Chellappa, Spectrochim. Acta, Part A, 2015, 136, 1269–1274 CrossRef CAS PubMed.
- M. Iniya, D. Jeyanthi, K. Krishnaveni and D. Chellappa, J. Lumin., 2015, 157, 383–389 CrossRef CAS PubMed.
- M. Iniya, D. Jeyanthi, K. Krishnaveni and D. Chellappa, RSC. Adv., 2014, 4, 25393–25397 RSC.
- T. Anand, G. Sivaraman and D. Chellappa, Spectrochim. Acta, Part A, 2014, 123, 18–24 CrossRef CAS PubMed.
- M. Iniya, D. Jeyanthi, K. Krishnaveni, A. Mahesh and D. Chellappa, Spectrochim. Acta, Part A, 2014, 120, 40–46 CrossRef CAS PubMed.
- G. Sivaraman, V. Sathiyaraja and D. Chellappa, J. Lumin., 2014, 145, 480–485 CrossRef CAS PubMed.
- R. Gui, X. An and W. Huang, Anal. Chim. Acta, 2013, 767, 134–140 CrossRef CAS PubMed.
- B. K. McMahon, R. Pal and D. Parker, Chem. Commun., 2013, 49, 5363–5365 RSC.
- X. Du, N. Y. Lei, P. Hu, Z. Lei, D. H. Ong, X. Ge, Z. Zhang and M. H. Lam, Anal. Chim. Acta, 2013, 787, 193–202 CrossRef CAS PubMed.
- H. S. Lv, S. Y. Huang, B. X. Zhao and J. Y. Miao, Anal. Chim. Acta, 2013, 788, 177–182 CrossRef CAS PubMed.
- M. Yang, Y. Song, M. Zhang, S. Lin, Z. Hao, Y. Liang, D. Zhang and P. R. Chen, Angew. Chem., Int. Ed. Engl., 2012, 51, 7674–7679 CrossRef CAS PubMed.
- H. Li, H. Guan, X. Duan, J. Hu, G. Wang and Q. Wang, Org. Biomol. Chem., 2013, 11, 1805–1809 CAS.
- Y. Xu, Z. Jiang, Y. Xiao, F. Z. Bi, J. Y. Miao and B. X. Zhao, Anal. Chim. Acta, 2014, 820, 146–151 CrossRef CAS PubMed.
- X. Zhang, S. Y. Jing, S. Y. Huang, X. W. Zhou, J. M. Bai and B. X. Zhao, Sens. Actuators, B, 2015, 206, 663–670 CrossRef CAS PubMed.
- K. C. Rupert and J. H. Henry, Bioorg. Med. Chem. Lett., 2003, 13, 347–350 CrossRef CAS.
- K. C. Langry, J. Org. Chem., 1991, 56, 2400–2404 CrossRef CAS.
- H. M. Li, C. Wang and J. W. Wang, Chin. Chem. Lett., 2000, 11, 949–950 CAS.
- X. X. Zhao, X. P. Chen, S. L. Shen, D. P. Li, S. Zhou, Z. Q. Zhou, Y. H. Xiao, G. Xi, J. Y. Miao and B. X. Zhao, RSC. Adv., 2014, 4, 50318–50324 RSC.
- H. S. Lv, S. Y. Huang, Y. Xu, X. Dai, J. Y. Miao and B. X. Zhao, Bioorg. Med. Chem. Lett., 2014, 24, 535–538 CrossRef CAS PubMed.
- S. L. Shen, X. F. Zhang, S. Y. Bai, J. Y. Miao and B. X. Zhao, RSC. Adv., 2015, 5, 13341–13346 RSC.
- X. F. Zhang, T. Zhang, S. L. Shen, J. Y. Miao and B. X. Zhao, RSC. Adv., 2015, 5, 49115–49121 RSC.
- X. F. Zhang, T. Zhang, S. L. Shen, J. Y. Miao and B. X. Zhao, J. Mater. Chem. B, 2015, 3, 3260–3266 RSC.
- X. D. Liu, Y. Xu, R. Sun, Y. J. Xu, J. M. Lu and J. F. Ge, Analyst, 2013, 138, 6542–6550 RSC.
- L. Zhang, D. Duan, X. Cui, J. Sun and J. Fang, Tetrahedron, 2013, 69, 15–21 CrossRef CAS PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Crystallogr., 2007, 64, 112–122 CrossRef PubMed.
- K. Brandenburg, Acta Crystallogr., Sect. A: Found. Crystallogr., 1996, 52, C562 Search PubMed.
- A. L. Spek, Acta Crystallogr., Sect. D: Biol. Crystallogr., 2009, 65, 148–155 CrossRef CAS PubMed.
- Z. L. Gong, B. X. Zhao, W. Y. Liu and H. S. Lv, J. Photochem. Photobiol., A, 2011, 218, 6–10 CrossRef CAS PubMed.
- S. Derinkuyu, K. Ertekin, O. Oter, S. Denizalti and E. Cetinkaya, Anal. Chim. Acta, 2007, 588, 42–49 CrossRef CAS PubMed.
- A. Lobnik, I. Oehme, I. Murkovic and O. S. Wolfbeis, Anal. Chim. Acta, 1998, 367, 159–165 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available: Supplementary absorption, fluorescent spectra, characterization of the compound: NMR, HRMS spectra. CCDC 1409600. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/c5ra14174e |
| This journal is © The Royal Society of Chemistry 2015 |