Visualized detection of vancomycin by supramolecular hydrogelations†
Yongquan Huaa,
Caiwen Oua,
Guoqin Chenb,
Xiaoli Zhangc,
Yanbin Caic,
Zhimou Yangc,
Ling Wang*c and
Minsheng Chen*a
aDepartment of Cardiology, Zhujiang Hospital of Southern Medical University, Guangzhou 510280, P. R. China. E-mail: gzminsheng@vip.163.com
bCardiovascular Medicine Department of Guangzhou Panyu Central Hospital, 8 Fuyudonglu Qiaonanjie Panyu District, Guangzhou, P. R. China
cState Key Laboratory of Medicinal Chemical Biology, College of Pharmacy, Nankai University, Collaborative Innovation Center of Chemical Science and Engineering, Tianjin 300071, P. R. China. E-mail: chwling@nankai.edu.cn
First published on 16th September 2015
Abstract
The specific binding in the self-assembled peptide Nap-GFFYEGDADA and the antibiotic vancomycin leads to supramolecular hydrogelations. This sol–gel phase transition can be easily identified by the naked eye. Therefore, it may be developed into a versatile method to detect vancomycin in remote places and in house.
Antibiotics, the most important medicine in treating bacterial infectious diseases, have been widely used in the world since their occurrence and have made great contributions in clinical practice.1 However, the abuse of antibiotics may lead to antimicrobial resistance2 and other adverse health effects such as nephrotoxicity and ototoxicity. Vancomycin (VAN), a powerful broad spectrum antibiotic, is a third-line drug and is often used as the last defense against superstrain bacteria.3 Unfortunately, vancomycin resistant bacteria have been found worldwide since the first report in Japan.4 Moreover, vancomycin is extremely ototoxic and nephrotoxic compared to other antibiotics.5 Therefore, it is of great significance to develop analytical techniques for detecting antibiotic residues such as vancomycin. In the past few decades, several analytical techniques have been developed, including microbiological assays,6 immunoassays7 and instrument8 analysis. Although these techniques exhibit both high sensitivity and accuracy in detecting antibiotics, they are expensive, time-consuming, have complicated operations and are not suitable for real-time and on-site analysis. Thus, the invention of a cost-effective, rapid and simple method for detecting antibiotics remains a considerable challenge.
Supramolecular hydrogels based on peptides have been demonstrated as promising nano-materials due to their excellent properties, such as the ease of design and synthesis, biocompatibility, degradability and fast response to external stimuli.9 They have shown great potential in drug delivery,10 regenerative medicine,11 immune boosting,12 and analyte detection,13 etc. The sol–gel and gel–sol transitions of peptide hydrogels can be triggered by pH or temperature changes, photo-irradiation, and enzymes. Such kinds of phase transition can be easily recognized by the naked eye, which has been developed into a novel method for analyte detection. For instance, gel–sol or sol–gel phase transitions have been applied for the detection of enzymes,14 glucose,15 metal ions,16 melamine,17 etc. Compared to routine detection techniques and methods, there is no need to utilize any expensive equipment or to transport samples to a laboratory for this new method. These advantageous properties make the fast, real-time, and on-site analysis of analytes possible, which is especially useful for in house and remote place detections.
It’s well known that VAN binds to the terminal D-Ala-D-Ala unit of peptidoglycan precursors with specificity and high affinity, thereby preventing their integration into the bacterial cell wall which would result in lysis of the cell.18 Besides being used in the treatment of disease, the ligand–receptor interactions between vancomycin and D-Ala-D-Ala derivatives in aqueous solution has been well established. Xu and co-workers have reported on supramolecular hydrogels based on N-(fluorenyl-9-methoxycarbonyl)-D-Ala-D-Ala, which exhibited gel–sol transition upon binding to vancomycin via a ligand–receptor interaction.19 In addition, they have also reported that the addition of vancomycin into mechanically weak hydrogels of a derivative of D-Ala-D-Ala led to an increase of the storage modulus of the hydrogel.20 Inspired by this strategy and the intrinsic properties of the self-assembled peptide systems, we opted to design a peptide based D-Ala-D-Ala derivative with self-assembling properties for the detection of vancomycin. We envisioned that the derivative may self-assemble into short nanofibers but not hydrogels because of the relatively weak inter-fiber interactions. Upon the addition of the vancomycin, the ligand–receptor interactions between D-Ala-D-Ala and vancomycin may increase the inter-fiber interactions and therefore cross-link the nanofibers, leading to a sol–gel phase transition. Such a kind of phase transition might be developed into a suitable method for the detection of vancomycin.
In order to test our hypothesis, we designed two peptides, Nap-GFFYEG-D-Ala-D-Ala (1, Scheme 1) and its enantiomer, Nap-GFFYEG-L-Ala-L-Ala (2, Scheme 1). Many derivatives containing the moieties FF or FFY have been verified as molecular hydrogelators with excellent self-assembly properties.21 The D-Ala-D-Ala moiety can bind to its receptor (vancomycin, VAN). We assumed that our designed peptides might self-assemble into short nanofibers with good water solubility, but could not form hydrogels without vancomycin. We firstly prepared Nap-GFFYEG-D-Ala-D-Ala and Nap-GFFYEG-L-Ala-L-Ala by standard solid phase peptide synthesis (SPPS). The pure compounds were achieved by reverse-phase high performance liquid chromatography (HPLC).
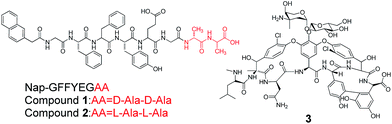 | ||
| Scheme 1 Chemical structures of Nap-GFFYEG-D-Ala-D-Ala (1), Nap-GFFYEG-L-Ala-L-Ala (2), and vancomycin (3). | ||
After obtaining the compounds, we studied their self-assembly properties in the presence or absence of VAN. We found that both peptides could form clear solutions in the phosphate buffer saline (PBS, pH = 7.4) solution at a concentration of 1.5 wt%, suggesting that the peptides have good water-dispersibility and would not form gels themselves at this concentration. We then tested whether they could form hydrogels or not after the addition of different equivalents of VAN. As shown in Fig. 1B, the solution of 1 formed a clear gel within 5 minutes after the addition of 0.005 equiv. of VAN (PBS, pH = 7.4), and it took a shorter time of about 30 seconds for hydrogelation when the concentration of VAN was higher than 0.005 equiv. Whereas, the solution of 2 could not form gels even in the presence of 0.1 equiv. of VAN (Fig. 1A). These results clearly demonstrate the success of our design. The minimum equiv. of VAN needed for gelation was about 0.005 in 0.01 M PBS. Such a value would be smaller when decreasing the pH value or increasing the ionic strength of the buffer (Fig. S8–S10†). For example, 0.0025 or 0.0017 equiv. of VAN (∼30 μg mL−1) was needed for gelation in pH = 5.5 or 0.05 M of PBS buffer solution. The minimum equiv. of VAN needed to gelate the PBS solution of the peptide (1.5 wt%, 0.05 M PBS, pH = 5.5) can be lowered to about 0.001 (20 μg mL−1) at 25 °C (Fig. S11†). Such sol–gel phase transitions can be easily recognized by the naked eye, which could be developed into a useful technique to detect VAN without any equipment.
 | ||
| Fig. 1 Optical images of the solutions of (A) Nap-GFFYEGDADA and (B) Nap-GFFYEGLALA without (left) or with (right) VAN in PBS. The temperature was 25 °C. | ||
Due to the intensive use of antibiotics for human (domestic and hospital use), veterinary and agricultural purposes, these compounds are continuously released into the environment without metabolism or in conjugate forms. Many studies have reported the occurrence of antibiotic residues in water sources including municipal wastewater effluents and surface waters.22 We then tested whether our method could be applied to detect VAN in wastewater effluents and biological fluids such as urine (Fig. 2). As shown in Fig. 2C, the mixture of PBS solution 1 and waste water without VAN would not form a gel. In the presence of VAN, a gel (Fig. 2D) would quickly form within 30 seconds when the concentration of vancomycin was higher than 0.005 equiv. Similar results were observed for the urine samples (Fig. 2E and F), and the lowest detection concentration of VAN was also about 0.005 equiv (0.1 mg mL−1). Although the detection limit of our method might be not as low as other methods, our method could be easily identified by the naked eye and there is no need to separate VAN from the samples or to send the samples to a laboratory and then measure them using equipment.
 | ||
| Fig. 2 Optical images of the solutions of Nap-GFFYEGDADA without (left) or with (right) VAN in PBS (A and B), waste water (C and D), and urine (E and F). The temperature was 25 °C. | ||
We then utilized a rheometer to investigate the mechanical properties of the hydrogels (Fig. S5–S7†). The mixed solutions containing 1 and different amounts of VAN were directly transferred to the rheometer. We then successively performed the dynamic time, strain, and frequency sweeps. As shown in Fig. 3B, the value of the storage moduli (G′, elasticity) for all the hydrogels was bigger than that of their corresponding loss moduli (G′′, viscosity), suggesting that all the samples behaved as viscoelastic materials. The G′ values of the hydrogels became bigger with an increased amount of VAN when the concentration of VAN was lower than 0.02 equiv. Once over 0.02 equiv., the G′ values began to decrease (Fig. 3C). These results were consistent with the optical images of the gels. As shown in Fig. 3A, the gels were very transparent under 0.02 equiv. of VAN. Once VAN exceeded 0.02 equiv., the gels turned to opaque.
We also characterized the morphology of the self-assembled nanostructures in the PBS solution and in the hydrogels with different equivalents of VAN by transmission electron microscopy (TEM). As shown in Fig. 4A, compound 1 itself self-assembled into short fibers with a low density of cross-linking points. With the addition of VAN, the fibers grew longer and bigger with a high density of cross-linking points (Fig. 4B and C). However, aggregated structures appeared when the concentration was higher than 0.1 equiv. (Fig. 4D). These observations were consistent with the results observed from the optical images and from the mechanical properties of the gels. These results also suggest that VAN helps to bring the peptides together and to extend the supramolecular chains to form longer fibers and their bundles. However, an excess amount of VAN disrupted the supramolecular interactions, thus resulting in partial aggregations.
 | ||
| Fig. 4 TEM images of (A) a solution of Nap-GFFYEGDADA, and gels with different equiv. of vancomycin: (B) 0.005 equiv., (C) 0.02 equiv., and (D) 0.1 equiv. | ||
To further understand the molecular arrangements in the hydrogels, we measured the emission spectra of the PBS solutions of 1 and 2 with and without VAN. As shown in Fig. 5, both the PBS solutions of 1 and of 2 exhibited distinct peaks centered at about 328 nm, suggesting monomeric naphthalene moieties in the solutions. In the presence of VAN, the peak showed a red shift to about 358 nm for 1 + 3 (gel), indicating that the naphthyl groups were stacked more efficiently. However, the emission spectrum of the solution of 2 showed little difference in the presence of VAN. The results also suggested that VAN helped to extend the supramolecular chains and the formation of hydrogels through the specific interaction between VAN and the dipeptide D-Ala-D-Ala.
Based on the above measurements, we have proposed a plausible molecular interaction between 1 and VAN in the gels. As indicated in Fig. 6, the peptide self-assembles into short fibers due to hydrogen bonds and the π–π interactions. With an increasing amount of VAN, VAN cross-links the short fibers via the specific ligand–receptor interaction and hydrogen bonds. Therefore, the short fibers grow into longer ones or form bundles, resulting in hydrogelation.
In conclusion, we have reported a supramolecular hydrogel-based system for the visualized detection of VAN. This simple method is applicable for detecting VAN in waste water effluent and urine by the naked eye. Vancomycin is generally used in humans to treat bacteria by oral administration or IV injection, and the therapeutic window of VAN is 20–5 μg mL−1. Immunoenzymatic techniques and chromatographic methods are generally used to detect the concentration of VAN in biological fluids, and their detection limit can be as low as 0.005 μg mL−1 (others are 2–5 μg mL−1).23 The detection limit of our method is about 20 μg mL−1 in 0.05 M of PBS solution at pH = 5.5. Although the detection limit of our method might not be as low as other techniques using complicated equipment, it may be a potential candidate for the rapid, simple, real-time and on-site screening of vancomycin in waste water effluents and biological fluids in house and in remote places. One limitation of our method is that the gelation will be greatly affected by both the pH value and the ionic strength. For the relatively accurate detection of VAN in a sample, the pH value is required to be adjusted to a certain value and the high ionic strength buffer solution of peptide (e.g. 0.1 M PBS) needs to be used to avoid the dilution effect from the addition of the sample. Our next step is to lower the detection limit of our method or to apply our method for the detection of bacteria, because the addition of bacteria might disrupt the binding between VAN and the peptide, which might lead to a gel–sol phase transition.
Acknowledgements
This work is supported by NSFC (31370964, 31470966) and the Guangdong Province Natural Science Fund (S2013010015314).Notes and references
- J. Davies and D. Davies, Microbiol. Mol. Biol. Rev., 2010, 74, 417–433 CrossRef CAS PubMed.
- H. C. Neu, Science, 1992, 257, 1064–1073 Search PubMed; J. L. Martínez, Science, 2008, 321, 365–367 CrossRef CAS PubMed; S. B. Levy, Sci. Am., 1998, 278, 32–39 CrossRef PubMed; F. Baquero, J.-L. Martínez and R. Cantón, Curr. Opin. Biotechnol., 2008, 19, 260–265 CrossRef PubMed.
- F. Bruniera, F. Ferreira, L. Saviolli, M. Bacci, D. Feder, G. P. M. da Luz, P. M. Sorgini, L. Azzalis, J. V. Campos and F. Fonseca, Eur. Rev. Med. Pharmacol. Sci., 2015, 19, 694–700 CAS.
- T. L. Smith, M. L. Pearson, K. R. Wilcox, C. Cruz, M. V. Lancaster, B. Robinson-Dunn, F. C. Tenover, M. J. Zervos, J. D. Band and E. White, N. Engl. J. Med., 1999, 340, 493–501 CrossRef CAS PubMed; M. Ploy, C. Grelaud, C. Martin, L. de Lumley and F. Denis, Lancet, 1998, 351, 1212 CrossRef; K. Hiramatsu, N. Aritaka, H. Hanaki, S. Kawasaki, Y. Hosoda, S. Hori, Y. Fukuchi and I. Kobayashi, Lancet, 1997, 350, 1670–1673 CrossRef; K. Hiramatsu, Lancet Infect. Dis., 2001, 1, 147–155 CrossRef; K. Hiramatsu, Drug Resist. Updates, 1998, 1, 135–150 CrossRef; S. Chang, D. M. Sievert, J. C. Hageman, M. L. Boulton, F. C. Tenover, F. P. Downes, S. Shah, J. T. Rudrik, G. R. Pupp and W. J. Brown, N. Engl. J. Med., 2003, 348, 1342–1347 CrossRef PubMed; Y. Cetinkaya, P. Falk and C. G. Mayhall, Clin. Microbiol. Rev., 2000, 13, 686–707 CrossRef.
- S. McKamy, E. Hernandez, M. Jahng, T. Moriwaki, A. Deveikis and J. Le, J. Pediatr., 2011, 158, 422–426 CrossRef CAS PubMed; A. H. Flannery and H. Bachmeier, Crit. Care Med., 2015, 43, e154 CrossRef PubMed; S. Elyasi, H. Khalili, S. Dashti-Khavidaki and A. Mohammadpour, Eur. J. Clin. Pharmacol., 2012, 68, 1243–1255 CrossRef PubMed.
- M. G. Pikkemaat, S. Oostra-van Dijk, J. Schouten, M. Rapallini and H. J. Van Egmond, Food Control, 2008, 19, 781–789 CrossRef CAS PubMed; M. G. Pikkemaat, Anal. Bioanal. Chem., 2009, 395, 893–905 CrossRef PubMed; L. Okerman, K. de Wasch and J. van Hoof, Analyst, 1998, 123, 2361–2365 RSC.
- E. Märtlbauer, E. Usleber, E. Schneider and R. Dietrich, Analyst, 1994, 119, 2543–2548 RSC; B. G. Knecht, A. Strasser, R. Dietrich, E. Märtlbauer, R. Niessner and M. G. Weller, Anal. Chem., 2004, 76, 646–654 CrossRef CAS PubMed; F. Davis and S. P. Higson, Pediatr. Res., 2010, 67, 476–480 CrossRef PubMed.
- M. Tajick and B. Shohreh, Poult. Sci., 2006, 5, 611–612 CrossRef CAS; H. Nakata, K. Kannan, P. D. Jones and J. P. Giesy, Chemosphere, 2005, 58, 759–766 CrossRef PubMed; L. Kantiani, M. Farré and D. Barceló, TrAC, TrendsAnal. Chem., 2009, 28, 729–744 CrossRef PubMed; H.-H. Chung, J.-B. Lee, Y.-H. Chung and K.-G. Lee, Food Chem., 2009, 113, 297–301 CrossRef PubMed; B. Chiavarino, M. E. Crestoni, A. Di Marzio and S. Fornarini, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 1998, 706, 269–277 CrossRef.
- S. Zhang, Acc. Chem. Res., 2012, 45, 2142–2150 CrossRef CAS PubMed; H. Wang and Z. Yang, Nanoscale, 2012, 4, 5259–5267 RSC; X. Du, J. Zhou and B. Xu, Chem.–Asian J., 2014, 9, 1446–1472 CrossRef PubMed; H. Cui, M. J. Webber and S. I. Stupp, J. Pept. Sci., 2010, 94, 1–18 CrossRef PubMed; Y. Li, M. Qin, Y. Cao and W. Wang, Sci. China: Phys., Mech. Astron., 2014, 57, 849 CrossRef; Y. Li, Y. Sun, M. Qin, Y. Cao and W. Wang, Nanoscale, 2015, 7, 5638 RSC; Y. Mao, T. Su, Q. Wu, C. Liao and Q. Wang, Chem. Commun., 2014, 50, 14429 RSC; S. Liu, A. Tan, M. Xie, Y. Zhao, J. Jiang and G. Liang, Angew. Chem., Int. Ed., 2015, 54, 3639 CrossRef PubMed.
- H. Wang and Z. Yang, Soft Matter, 2012, 8, 2344–2347 RSC; A. G. Cheetham, P. Zhang, Y.-a. Lin, L. L. Lock and H. Cui, J. Am. Chem. Soc., 2013, 135, 2907–2910 CrossRef CAS PubMed; X. Li, J. Li, Y. Gao, Y. Kuang, J. Shi and B. Xu, J. Am. Chem. Soc., 2010, 132, 17707–17709 CrossRef PubMed; F. Zhao, M. L. Ma and B. Xu, Chem. Soc. Rev., 2009, 38, 883–891 RSC.
- H. Hosseinkhani, P.-D. Hong and D.-S. Yu, Chem. Rev., 2013, 113, 4837–4861 CrossRef CAS PubMed; F. Gelain, L. D. Unsworth and S. Zhang, J. Controlled Release, 2010, 145, 231–239 CrossRef PubMed; R. N. Shah, N. A. Shah, M. M. D. R. Lim, C. Hsieh, G. Nuber and S. I. Stupp, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 3293–3298 CrossRef PubMed.
- J. S. Rudra, S. Mishra, A. S. Chong, R. A. Mitchell, E. H. Nardin, V. Nussenzweig and J. H. Collier, Biomaterials, 2012, 33, 6476–6484 CrossRef CAS PubMed; J. S. Rudra, T. Sun, K. C. Bird, M. D. Daniels, J. Z. Gasiorowski, A. S. Chong and J. H. Collier, ACS Nano, 2012, 6, 1557–1564 CrossRef PubMed.
- L. L. Lock, A. G. Cheetham, P. Zhang and H. Cui, ACS Nano, 2013, 7, 4924–4932 CrossRef CAS PubMed; K. Mizusawa, Y. Ishida, Y. Takaoka, M. Miyagawa, S. Tsukiji and I. Hamachi, J. Am. Chem. Soc., 2010, 132, 7291–7293 CrossRef PubMed; Y. Cai, Y. Shi, H. Wang, J. Wang, D. Ding, L. Wang and Z. Yang, Anal. Chem., 2014, 86, 2193–2199 CrossRef PubMed; T. Yoshii, S. Onogi, H. Shigemitsu and I. Hamachi, J. Am. Chem. Soc., 2015, 137, 3360–3365 CrossRef PubMed; C. Ren, H. Wang, D. Mao, X. Zhang, Q. Fengzhao, Y. Shi, D. Ding, D. Kong, L. Wang and Z. Yang, Angew. Chem., Int. Ed., 2015, 54, 4823–4827 CrossRef PubMed.
- S. Toledano, R. J. Williams, V. Jayawarna and R. V. Ulijn, J. Am. Chem. Soc., 2006, 128, 1070–1071 CrossRef CAS PubMed; M. Ikeda, T. Tanida, T. Yoshii, K. Kurotani, S. Onogi, K. Urayama and I. Hamachi, Nat. Chem., 2014, 6, 511 CrossRef PubMed; S. C. Bremmer, J. Chen, A. J. McNeil and M. B. Soellner, Chem. Commun., 2012, 48, 5482–5484 RSC; Y. Gao, Y. Kuang, Z.-F. Guo, Z. Guo, I. J. Krauss and B. Xu, J. Am. Chem. Soc., 2009, 131, 13576–13577 CrossRef PubMed.
- X. D. Xu, B. B. Lin, J. Feng, Y. Wang, S. X. Cheng, X. Z. Zhang and R. X. Zhuo, Macromol. Rapid Commun., 2012, 33, 426–431 CrossRef CAS PubMed.
- K. N. King and A. J. McNeil, Chem. Commun., 2010, 46, 3511–3513 RSC.
- J. W. Zhang, C. W. Ou, Y. Shi, L. Wang, M. S. Chen and Z. M. Yang, Chem. Commun., 2014, 50, 12873–12876 RSC.
- D. Kahne, C. Leimkuhler, W. Lu and C. Walsh, Chem. Rev., 2005, 105, 425–448 CrossRef CAS PubMed; D. H. Williams, A. J. Maguire, W. Tsuzuki and M. S. Westwell, Science, 1998, 280, 711–714 CrossRef; C. Walsh, Nature, 2000, 406, 775–781 CrossRef PubMed.
- Y. Zhang, H. Gu, Z. Yang and B. Xu, J. Am. Chem. Soc., 2003, 125, 13680–13681 CrossRef CAS PubMed.
- Y. Zhang, Z. Yang, F. Yuan, H. Gu, P. Gao and B. Xu, J. Am. Chem. Soc., 2004, 126, 15028–15029 CrossRef CAS PubMed.
- M. Zhou, A. M. Smith, A. K. Das, N. W. Hodson, R. F. Collins, R. V. Ulijn and J. E. Gough, Biomaterials, 2009, 30, 2523–2530 CrossRef CAS PubMed; W. Zheng, J. Gao, L. Song, C. Chen, D. Guan, Z. Wang, Z. Li, D. Kong and Z. Yang, J. Am. Chem. Soc., 2012, 135, 266–271 CrossRef PubMed; Y. Li, Y. Sun, M. Qin, Y. Cao and W. Wang, Nanoscale, 2015, 7, 5638 RSC; Y. Tian, H. Wang, Y. Liu, L. Mao, W. Chen, Z. Zhu, W. Liu, W. Zhen, Y. Zhao, D. Kong, Z. Yang, W. Zhang, Y. Shao and X. Jiang, Nano Lett., 2014, 14, 1439 CrossRef PubMed.
- K. Kümmerer, Chemosphere, 2009, 75, 417–434 CrossRef CAS PubMed; R. Hirsch, T. Ternes, K. Haberer and K.-L. Kratz, Sci. Total Environ., 1999, 225, 109–118 CrossRef.
- M. J. D. Valle, F. G. Lopez and A. S. Navarro, J. Pharm. Biomed. Anal., 2008, 48, 835–839 CrossRef CAS PubMed; D. Sym, C. Smith, G. Meenan and M. Lehrer, Ther. Drug Monit., 2001, 23, 441–444 CrossRef.
Footnote |
| † Electronic supplementary information (ESI) available: Synthesis and characterization, rheological data, and the effect of pH value, ionic strength, and the temperature on the gelation. See DOI: 10.1039/c5ra14045e |
| This journal is © The Royal Society of Chemistry 2015 |



