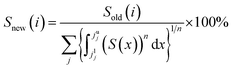AFM and NMR imaging of squid tropomyosin Tod p1 subjected to high hydrostatic pressure: evidence for relationships among topography, characteristic domain and allergenicity†
Yifeng Zhanga,
Bona Daib,
Yun Deng*a and
Yanyun Zhaoc
aKey Laboratory of Urban Agriculture (South), Ministry of Agriculture, SJTU-Bor S. Luh Food Safety Center, Department of Food Science and Technology, Shanghai Jiao Tong University, 800 Dongchuang Road, Shanghai 200240, China. E-mail: foodsjtu@sjtu.edu.cn; Fax: +86-21-34205755; Tel: +86-21-34205755
bInstrumental Analysis Center, Shanghai Jiao Tong University, 800 Dongchuan Road, 200240 Shanghai, China
cDepartment of Food Science & Technology, Oregon State University, Corvallis, OR 97331, USA
First published on 19th August 2015
Abstract
The surface topography, characteristic domain and allergenicity of squid tropomyosin Tod p1 (TMTp1) treated under single- and two-cycle high hydrostatic pressure (HHP) were analyzed. Atomic force microscopy (AFM) showed that the HHP treatment led to the formation of a rougher surface of squid TMTp1; the two-cycle 600 MPa HHP treatment produced the largest effect, with a mean roughness, maximum height, skewness and kurtosis of 6.56 nm, 94.83 nm, 3.05 and 23.13, respectively. The HHP treatment caused lower IgE and IgG-binding capacities, indicating a significant reduction of the allergenicity (p < 0.05) due to variations in the AFM surface topography. The peaks of the main allergenic characteristics affected were 0.99, 1.16, 1.21, 1.79, 1.82, 2.84, 2.88 and 3.37 ppm (in the initial 1H nuclear magnetic resonance (NMR) spectra). The HHP treatment changed the AFM surface topography and caused the migration or disappearance of the related 1H NMR peaks; these changes were significantly correlated (p < 0.05) with the reductions in allergenicity observed.
1. Introduction
Seafood allergies are one of the most common, severe, and long lasting food allergies, and receive extensive attention from people with a predisposition to allergies.1 Tropomyosin Tod p1 (TMTp1), a water soluble 38 kDa protein, is a major allergen which occurs extensively in crustacean and mollusc species.2 Many attempts have been made to reduce the presence of allergens in various foods during processing.3–5 High hydrostatic pressure (HHP), a valuable non-thermal food processing technology,6 can significantly reduce the activity of many allergens,7 such as β-lactoglobulin,8 and those found in soybean seeds9 and almond milk.10 In our previous study,11 single-cycle HHP treatments at 200, 400 or 600 MPa for 20 min decreased the allergenicity of squid TMTp1; the 400 and 600 MPa treatments were more effective than the 200 MPa treatment, according to indirect enzyme linked immunosorbent assays (ELISAs).HHP affects the structure of proteins, which can be shown by characterizing protein structures using circular dichroism spectra, as well as by analyzing the free sulfhydryl content and surface hydrophobicity index.8,9,11 However, the precise nature of the internal motions within protein macromolecules remains a mystery and is not easy to describe accurately.12 Highly sensitive and reliable methods to analyse squid TMTp1 are required to enable assessments of their allergenic properties.
Different proteins have different conformational properties which are important for the modification of their functional properties.13 Many studies have been done in recent years to explore the relationship between physical properties and biological functions, such as the research on the Pre-T-cell receptor structure and interactions,14 the physical properties of supramolecular peptides,15 and the antibacterial and/or antifouling property on the surface of peptides.16 Atomic force microscopy (AFM) is the principal method that allows complex chemical or biochemical changes and reactions to be clarified directly in many systems.17 Surfaces with varying AFM surface topography indices have different functional properties.18 For example, surface roughness, which directly corresponds to the sizes of proteins, could play an important role in defining different proteins’ characteristics.19 Altering the surface roughness influences the chemical reactivity of proteins,20 which affects properties like the sensitization response, in vitro osteoblast differentiation, and local factor production.21,22 HHP is likely to produce powerful percussive effects23–25 that might have an impact on the morphology of the squid TMTp1 molecules in solution and upon their adsorption onto a surface. Hence, AFM surface topography could be used to assess the effect of HHP treatments on squid TMTp1 and its associated allergenic properties.
Nuclear magnetic resonance (NMR) spectroscopy has become one of the most accepted methods for determining the structural properties of native and processed proteins.26 NMR data, especially 1H NMR spectra, provide information about the structure and dynamic properties of proteins, such as information relating to the positions, bonds and movements of specific atoms.27 This information could be used to describe the changes exhibited by a processed protein in terms of its structural, chemical and dynamic properties. Previous research27 illustrated that 1H NMR spectra can be used to generate a fingerprint, give insights into the molecular folding of allergen proteins, and offer an independent method for assessing the structural properties of proteins. Clearly, NMR could be used to analyse changes in the characteristic regions of squid TMTp1 when it is treated under HHP, as well as the positions, bonds and movements of the characteristic peaks which are related to the allergenic properties.
Compared with single-cycle HHP, multiple-cycle HHP may cause more significant structural damage to biomacromolecules,28 and could be more effective in inactivating microorganisms, improving the food quality, and would lower the cost in comparison to a single-cycle of the same dwell time.29,30 Previously we found that two-cycle HHP treatments were more effective in controlling microbial growth and reducing the deterioration of squids.23 However, there is limited information available to compare single- and two-cycle HHP treatments with respect to the nutritional, functional or other properties of seafood. Therefore, it is worthwhile to study the changes in allergenicity resulting from treating squid TMTp1 under both one- and two-cycle HHP.
The objectives of this study were to investigate the effect of one- and two-cycle HHP treatment on the allergenicity of squid TMTp1. To do this, the allergenic characteristic regions and peaks of the 1H NMR spectra of the HHP treated squid TMTp1 were analyzed. AFM was then used to evaluate the changes to the surface topography of the HHP treated squid TMTp1. Finally, the relationships among the allergenic properties, AFM surface topography and NMR characteristic regions in the HHP treated squid TMTp1 samples were explored.
2. Materials and methods
2.1 Materials and sample preparation
Squid (Todarodes pacificus; 310 ± 25 g per whole squid) were obtained from the Chinese Academy of Fishery Sciences (Shanghai, China) and stored at −80 °C until processing. The extraction, purification and identification of squid TMTp1 and the rabbit anti-squid TMTp1 polyclonal antibodies were carried out according to our previous methods.11 All procedures concerning animals were performed in accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals of Shanghai Jiao Tong University. The protocol was approved by the National Natural Science Foundation Commission of China (Permit number: 31271955) and the Committee on the Ethics of Animal Experiments of School of Agriculture and Biology, Shanghai Jiao Tong University. Individual human serum samples were collected from five patients (Xinhua Hospital, Shanghai, China) who were determined to have a squid allergy based on their history and the objective manifestations after ingestion of squid. The pooled sera of two non-allergic individuals from the same hospital were used as a negative control. Also, written informed consent was obtained from each human before the human serum was collected.The squid TMTp1 solution was diluted with 20 mM Tris–HCl (pH 7.5) to create a final concentration of 1 mg mL−1. Then, 20 mL samples of the diluted squid TMTp1 solution were individually packaged in polyamide/chlorinated polypropylene complex film bags (17 × 23 cm), the oxygen and water vapor permeabilities of which were 5.383 × 10−15 cm3 cm cm−2 s−1 Pa−1 and 1.383 × 10−12 g cm cm−2 s−1 Pa−1, respectively. The bags were sealed for the HHP treatments.
2.2 High hydrostatic pressure (HHP) treatments
The experimental design comprised of a control, three single-cycle treatments and three two-cycle treatments, with three replicates of each treatment performed. For the control, the squid TMTp1 samples were held at ambient pressure (0.1 MPa and ∼25 °C), with no HHP treatment. The three single-cycle treatments were carried out at 200, 400 or 600 MPa for 20 min and named as S-200, S-400 and S-600, respectively. For the three two-cycle treatments, two 10 min cycles at 200, 400 or 600 MPa were performed and named as T-200, T-400 and T-600, respectively. The treatments used in this study were based on our previous study.23The HHP treatments were carried out using an HHP device (HHP-750, Kefa High Pressure Food Processing Inc., Baotou, China) for a total holding time of 20 min. Water was used as the pressure transmitting medium, and the temperature was maintained at 25 ± 1 °C by a circulating water system. The pressure vessel (90 mm diameter × 320 mm height) had a volume of 2.5 L and a pressure range of 0–700 MPa. The rate for the treatments to reach the top pressure was set at approximately 200 MPa min−1 and the decompression time after the treatment was immediate (<4 s), in order to minimize adiabatic heating.29 After the HHP treatments, all the samples were freeze dried in a Freezone 2.5 L Triad system (Labconco Inc., Missouri, USA), and stored at −80 °C until further analysis.
2.3 AFM analysis
The squid TMTp1 morphology was characterized using AFM (Vecco Metrology Group, Digital Instruments, Santa Barbara, CA, USA). A 2 μL droplet of the squid TMTp1 (25 μg mL−1) was spread onto a freshly cleaved mica disk, which was adhered to a stainless plate using double-sided tape. The sample was air-dried for 60 min at ambient temperature. The sample surface topography was then measured using Multimode Nanoscope AFM. Imaging was performed in the tapping mode, with a cantilever resonant frequency of around 330 kHz and a scan rate of 2.441 Hz. Three subsamples of every treatment sample were scanned three times each under ambient atmospheric conditions. The original data was levelled, to remove the tilt, by applying a numerical second-order correction, and the mean values of surface roughness (Ra), maximum height (Rmax), skewness and kurtosis were determined using the NanoRule software (Pacific Nanotechnology, Santa Barbara, CA, USA).2.4 NMR analysis
1H NMR analyses of the squid TMTp1 samples were carried out on an Avance Bruker III HD 600 MHz NMR spectrometer (Bruker Biospin, Rheinstetten, Germany), which was equipped with a 5 mm TCl CryoProbe and maintained at 25 °C. The solvent used was D2O and a final sample concentration of 10 mg mL−1 and a 500 μL solution was used for each analysis. A solvent pre-saturation procedure was employed to suppress the water peak. The standard Carr-Purcell-Meiboom-Gill pulse sequence was used to record the spectra; the Carr-Purcell-Meiboom-Gill pre-saturation pulse sequence took the form of RD − 90° − (t − 180° − t)n − ACQ, where RD is the relaxation delay of 2 s, 90° and 180° represent the RF pulses that trip the magnetization vector, t is the spin-echo delay of 400 ms, n represents the number of loops (80 in this experiment), and ACQ is the data acquisition period of 1.36 s. In this experiment, the data points were acquired with 128 transients and the time delay was 5 s. The NMR spectrum was imported into the Chenomx NMR Suite 7.7 software (Chenomx, Inc., Alberta, Canada). Subsequently, the spectra were imported to AMIX (Bruker Biospin, Rheinstetten, Germany) and were all reduced to fixed integral regions (0.04 ppm) for further analysis. The normalization method was performed according to a previous study,31 based on the general equation:where Sold(i) and Snew(i) are the intensities of the variable i (spectral feature) before and after normalization, respectively; j is an index of the spectral regions used for normalization; and jlj and juj are the lower and upper borders of the spectral region j, for which the power n of the intensities S(x) is integrated.
2.5 Indirect ELISAs
An indirect ELISA was performed to analyse the allergenic properties of the squid TMTp1 samples by measuring their hydrolysates after in vitro digestion, according to our previous methods.11 An indirect ELISA with the human sera of five allergic patients (P1–P5) was performed. In addition, indirect ELISAs with rabbit anti-squid TMTp1 polyclonal antibodies (R), rabbit anti-squid TMTp1 polyclonal antibodies for hydrolysates after Simulated Gastric Fluid (SGF) digestion (R-SGF) and rabbit anti-squid TMTp1 polyclonal antibodies for hydrolysates after Simulated Intestinal Fluid (SIF) digestion (R-SIF) were performed. An automated ELISA plate reader (Thermo Co., USA) was used to monitor the absorbance at 450 nm. Three replicate measurements were carried out.2.6 Statistical analysis
Results are reported as mean ± standard deviation (SD). The statistical differences between treatments were assessed using analysis of variance (ANOVA), followed by Tukey’s HSD post hoc test (p < 0.05) using the SAS 9.2 software (SAS Institute Inc., Cary, NC, USA).3. Results and discussion
3.1 AFM surface topography of HHP treated squid TMTp1
The AFM topographic images revealed the structural differences among the squid TMTp1 samples subjected to different HHP treatments. In the AFM image of the control squid TMTp1 samples (Fig. 1A), the particles were distributed relatively evenly. The initial AFM image of the control squid TMTp1 samples presented similar characteristics to human cardiac α-tropomyosin.32,33 The S-200 treatment caused significant changes of the surface topography in the squid TMTp1 samples (Fig. 1B), in which the appearance of the surface roughness was darkened. The S-400 treatment caused a further increase in the maximum height and there was no uniformity between the surface topographies of the treated and control samples (Fig. 1C); the surface became even more rough and wrinkled when the pressure was increased to 600 MPa, and the AFM images produced from the S-600 treatment were entirely different to the control (Fig. 1D). This indicates that the higher pressures caused more changes to the surface topographical properties of the squid TMTp1 samples during the single-cycle treatments. Much more significant changes of the surface topography of squid TMTp1 were observed with the two-cycle HHP treatments (Fig. 1E–G). All of the two-cycle HHP treatments led to a much more rough and wrinkled surface topography in comparison to the S-600 treatment.To consider the AFM outputs further, the surface topography indices of Ra, Rmax, skewness and kurtosis of the different HHP treated squid TMTp1 samples were considered. Ra and Rmax are the most commonly used descriptors of surface roughness.34 Skewness is a measure of the symmetry of the statistical distribution, and when it is 0 there is an even distribution of peaks and troughs, of specific heights; a surface with larger peaks than troughs has a positive skewness, and vice versa.18 Kurtosis is a measure of the spikiness of the statistical distribution and a normal distribution has a kurtosis equal to 3. If the kurtosis is <3, it corresponds to a statistical distribution that is flatter than the normal distribution, and the opposite is true for distributions with a kurtosis value >3.18 Surfaces with varying Ra, Rmax, skewness or kurtosis values have different functional properties.20
In this study, the control had an Ra of 0.37 nm, Rmax of 15.20 nm, a skewness value of 7.10 and a kurtosis value of 91.51. In general, the HHP treated squid TMTp1 samples had significantly higher Rmax values (28.74–94.83 nm; p < 0.05), higher Ra values (except for S-200; 0.84–6.56 nm; p < 0.05) and different values of skewness and kurtosis. It is interesting that the single-cycle HHP treatments gave rise to higher skewness values, while the two-cycle HHP treatments led to lower values compared with the control. Similar results were obtained for the kurtosis values (apart from S-200, which was slightly lower). The T-600 treatment produced the biggest difference in the indices from the control, with a Ra of 6.56 nm, Rmax of 94.83, a skewness value of 3.05 and a kurtosis value of 23.13. There were significant differences between the indices of the of the squid TMTp1 samples produced by the single- and two-cycle HHP treatments at the same pressure level.
Biopolymers like proteins show a transformation of their native structure after HHP.35 However, there is limited information available about the chemistry behind the effect of HHP on the changes to the surface properties of proteins. HHP might have an impact on the hydrogen bonds, ionic or hydrophobic interactions with the modification of functional characteristics, thus changing the protein structure.29,35,36 When HHP was applied, the water molecules might have been squeezed into the free spaces between the semi-crystalline and amorphous lamellae, with greater forces produced by the higher pressure treatments,35 indicating that the higher pressures caused more changes to these intermolecular forces (hydrogen bonds, ionic and hydrophobic interactions) in the squid TMTp1 samples during the single-cycle treatments. Much more significant intermolecular changes to the squid TMTp1 samples were observed with the two-cycle HHP treatments (Fig. 1E–G). Compared with the control, the T-600 treatment produced the biggest intermolecular changes.
Surfaces with varying AFM surface topography indices have different functional properties.18 Altering the surface roughness influenced the chemical reactivity of proteins,20 which affected the protein properties. The hydrogen bonds, ionic and hydrophobic interactions of proteins are closely associated with their allergenic properties, for example, hydrogen bonds help the stabilization of allergenicity characteristics.37 Hence, these intermolecular and protein structure changes could potentially affect the allergenicity characteristics in the squid TMTp1 samples (Table 1).
| Treatment group | Mode | Pressure (MPa) | AFM indicesab | |||
|---|---|---|---|---|---|---|
| Mean roughness (Ra, nm) | Max height (Rmax, nm) | Skewness | Kurtosis | |||
| a Values are expressed as average ± standard deviation (SD) (n = 3).b Means in the same column with different lowercase letters (a–g) are significantly different (p < 0.05). | ||||||
| Control | — | 0.1 | 0.37 ± 0.02f | 15.20 ± 0.50g | 7.10 ± 0.26b | 91.51 ± 0.32b |
| S-200 | Single | 200 | 0.35 ± 0.02f | 28.74 ± 0.40f | 7.33 ± 0.36b | 85.58 ± 0.66c |
| S-400 | Single | 400 | 0.84 ± 0.04e | 33.02 ± 0.26e | 8.58 ± 0.24a | 102.81 ± 0.53a |
| S-600 | Single | 600 | 1.66 ± 0.01d | 29.31 ± 0.92d | 8.55 ± 0.55a | 100.06 ± 1.62a |
| T-200 | Two | 200 | 5.60 ± 0.16c | 53.98 ± 0.96c | 2.67 ± 0.20c | 22.95 ± 0.49d |
| T-400 | Two | 400 | 5.10 ± 0.04b | 85.29 ± 1.02b | 2.73 ± 0.56c | 24.31 ± 0.37d |
| T-600 | Two | 600 | 6.56 ± 0.06a | 94.83 ± 0.89a | 3.05 ± 0.26c | 23.13 ± 0.79d |
3.2 NMR analysis of HHP treated squid TMTp1
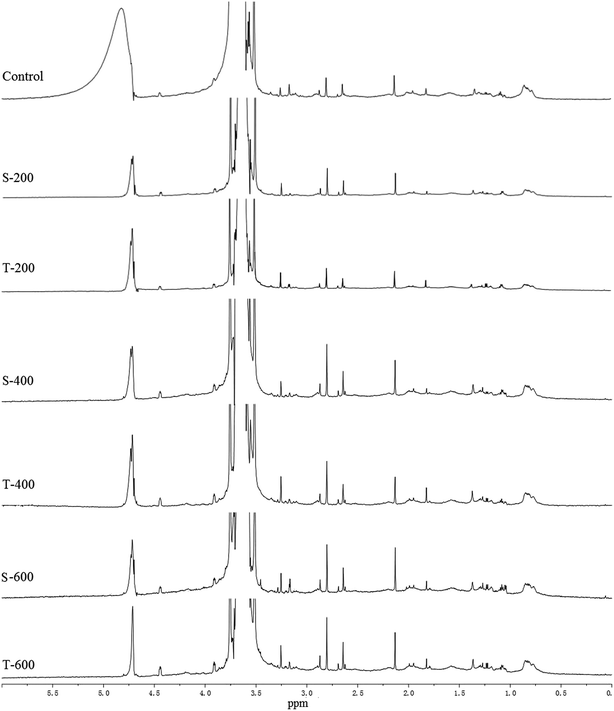
The 1H NMR spectra of the control and HHP treated squid TMTp1 show characteristic water resonance regions (4.70–5.02 ppm; Fig. 2), which are not analysed in this study. The sharp peak observed at 3.70 ppm in the 1H NMR spectra of the squid TMTp1 samples is due to residual Tris buffer.38 All of the squid TMTp1 samples showed multiple signals at approximately 0.85, 1.28, 1.88, 2.16, 2.68, 2.88, 3.27, and 3.51 ppm, amongst other chemical shifts (Fig. 2 and Table 2); the changes in the positions of the peaks within the 0.75–3.51 ppm range of the 1H NMR spectra for the different HHP treated squid TMTp1 samples could be attributed to the protons of the protein’s amino groups. The changes in the peaks between the different samples were attributed to the effect of the single- and two-cycle HHP treatments.| The position of peaks (ppm) | |||||||
|---|---|---|---|---|---|---|---|
| Control | S-200 | S-400 | S-600 | T-200 | T-400 | T-600 | |
| a S-200, S-400 and S-600 were the single-cycle high hydrostatic pressure (HHP) treatments, maintained at 200, 400 and 600 MPa for 20 min, respectively. T-200, T-400 and T-600 were the two-cycle HHP treatments comprised of two 10 min cycles at 200, 400 and 600 MPa, respectively. | |||||||
| 1 | 3.51 | 3.51 | 3.51 | 3.51 | 3.51 | 3.51 | 3.51 |
| 2 | 3.46 | 3.47 | 3.46 | 3.47 | |||
| 3 | 3.37 | 3.35 | 3.35 | 3.35 | 3.35 | 3.35 | 3.35 |
| 4 | 3.29 | 3.28 | 3.28 | 3.29 | 3.29 | 3.28 | 3.29 |
| 5 | 3.27 | 3.26 | 3.25 | 3.26 | 3.25 | 3.25 | 3.26 |
| 6 | 3.23 | 3.23 | 3.21 | 3.21 | 3.21 | 3.21 | 3.21 |
| 7 | 3.17 | 3.17 | 3.17 | 3.17 | 3.17 | 3.17 | 3.17 |
| 8 | 3.14 | 3.13 | 3.13 | 3.13 | |||
| 9 | 3.1 | 3.1 | 3.1 | 3.1 | 3.1 | 3.1 | |
| 10 | 3.07 | 3.06 | |||||
| 11 | 3.00 | 3 | |||||
| 12 | 2.88 | 2.87 | 2.87 | 2.87 | 2.87 | 2.87 | 2.87 |
| 13 | 2.84 | ||||||
| 14 | 2.81 | 2.8 | 2.8 | 2.8 | 2.8 | 2.8 | 2.8 |
| 15 | 2.78 | ||||||
| 16 | 2.68 | 2.69 | 2.69 | 2.69 | 2.69 | 2.69 | 2.69 |
| 17 | 2.65 | 2.64 | 2.64 | 2.64 | 2.64 | 2.64 | 2.64 |
| 18 | 2.63 | 2.63 | 2.62 | 2.62 | 2.62 | 2.62 | 2.62 |
| 19 | 2.57 | 2.53 | 2.53 | 2.53 | 2.52 | 2.52 | |
| 20 | 2.39 | ||||||
| 21 | 2.16 | 2.19 | 2.19 | 2.19 | 2.2 | 2.2 | 2.18 |
| 22 | 2.13 | 2.13 | 2.13 | 2.13 | 2.13 | 2.13 | 2.13 |
| 23 | 2.10 | ||||||
| 24 | 1.98 | 1.99 | 1.99 | 1.99 | 1.99 | 1.99 | 1.99 |
| 25 | 1.88 | 1.95 | 1.95 | 1.95 | 1.95 | 1.95 | 1.95 |
| 26 | 1.82 | 1.82 | 1.82 | 1.82 | 1.82 | 1.82 | 1.82 |
| 27 | 1.79 | 1.79 | 1.79 | 1.79 | 1.79 | 1.79 | |
| 28 | 1.70 | ||||||
| 29 | 1.59 | 1.56 | 1.59 | 1.58 | 1.58 | 1.58 | |
| 30 | 1.43 | ||||||
| 31 | 1.37 | 1.37 | 1.37 | 1.37 | 1.37 | 1.37 | |
| 32 | 1.34 | ||||||
| 33 | 1.32 | ||||||
| 34 | 1.28 | 1.29 | 1.29 | 1.29 | 1.29 | 1.29 | 1.29 |
| 35 | 1.26 | 1.27 | 1.27 | 1.27 | 1.27 | 1.27 | 1.27 |
| 36 | 1.23 | 1.23 | 1.23 | 1.23 | 1.23 | 1.23 | 1.23 |
| 37 | 1.21 | 1.22 | 1.22 | 1.22 | 1.22 | 1.22 | 1.22 |
| 38 | 1.16 | 1.18 | 1.18 | 1.19 | 1.19 | 1.19 | 1.18 |
| 39 | 1.09 | 1.1 | 1.09 | 1.1 | 1.1 | 1.09 | 1.1 |
| 40 | 1.08 | 1.08 | 1.08 | 1.08 | 1.08 | 1.08 | 1.08 |
| 41 | 1.06 | 1.07 | 1.07 | 1.07 | 1.07 | 1.07 | 1.07 |
| 42 | 1.05 | 1.05 | 1.05 | 1.05 | |||
| 43 | 0.99 | ||||||
| 44 | 0.85 | 0.85 | 0.85 | 0.85 | 0.85 | 0.85 | 0.85 |
| 45 | 0.83 | 0.83 | 0.83 | 0.83 | 0.83 | 0.83 | 0.83 |
| 46 | 0.82 | 0.82 | 0.81 | 0.82 | 0.82 | 0.81 | 0.82 |
| 47 | 0.78 | 0.78 | 0.78 | 0.78 | 0.77 | 0.78 | |
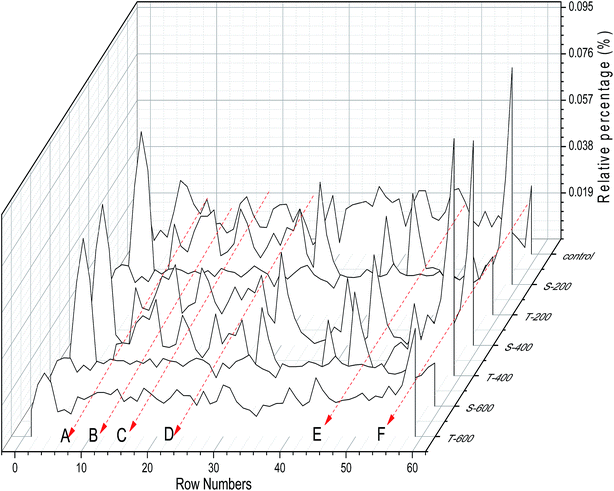
The NMR spectra (0.71–3.51 ppm) were divided into 0.04 ppm integral regions and subjected to normalization analysis. The relative percentage of each integral region after normalization of 1H NMR spectra for the squid TMTp1 samples after each HHP treatment is shown in Fig. 3; the regions of 1, 2, 58 and 59 represented 0.71–0.75, 0.75–0.80, 3.43–3.47 and 3.47–3.51 ppm in the 1H NMR spectra, respectively (ESI Table S1†). The HHP treatments clearly increased P0.75–0.80 and P0.80–0.84, but decreased P2.95–3.00 compared with the control. P0.75–0.80 increased from 0.0058% (control) to 0.0215% (T-600), but P2.95–3.00 decreased significantly from 0.0261% (control) to 0.0014% (T-600) (Fig. 3). The single- and two-cycle HHP treatments conducted at the same pressure level also produced significantly different results (Fig. 3).
These results indicated that the HHP treatments modified the structure of squid TMTp1 by impacting on some characteristic groups and controlling the allergenic properties, which was consistent with previously published studies.39,40 In general, the NMR spectroscopic analysis of squid TMTp1 demonstrated that the HHP treatments changed some related NMR characteristic regions and resulted in variations in the protein’s allergenic properties. The changes in the protein residues, which were related to the allergenic properties, could be found by NMR data through information about the positions, bonds, and movements of specific atoms and the normalization method.39,40 These residues, which were found in many allergens by NMR,39,40 play an important role in IgE/G binding and the allergenicity of these allergens by reacting with epitopes.42 The epitopes of allergens are essential to an allergen and the development of specific allergen immunotherapy.39,40 This demonstrated that the higher pressures and two-cycle treatments caused more changes to these residues of squid TMTp1 during the HHP treatments. From the NMR data (ESI Table S1 and Fig. S3†), there were the biggest changes to these residues in the T-600 treated squid TMTp1 sample. However, the exact mechanisms causing the changes in the samples should be further studied.
3.3 Allergenic properties
Indirect ELISAs were performed to study the allergenic properties of squid TMTp1 using the human sera of five allergic patients (IgE-binding capacity; IgE) or rabbit anti-squid TMTp1 polyclonal antibodies (IgG-binding capacity; IgG). Overall, the HHP treatments resulted in lower IgE and IgG values, representing a decrease in allergenicity, and the two-cycle HHP treatments (Table 3) caused significant decreases. Significant differences (p < 0.05) in the IgE values were observed between the control and two-cycle HHP treated samples; the higher the pressure of the HHP treatment, the lower the IgE values. As such, the T-600 samples had the lowest IgE values of 0.87, 0.24, 0.25, 0.22 and 0.25 for the five human sera, respectively. The IgG (R) value of the control squid TMTp1 samples was 1.66, which was in good agreement with our previous report.11 The IgG (R) values clearly show that the two-cycle HHP treatments resulted in significantly lower binding capacities compared with the control treatment (Table 3). The changes in the IgG values of hydrolysates after in vitro digestion (R-SGF and R-SIF) were subject to the variations of the control and HHP treated samples, and the control samples had the highest IgG (R-SIF) value (0.67). Previously, we reported the IgE and IgG values of squid TMTp1 treated with HHP at 200, 400 or 600 MPa for 20 min.11 The two-cycle HHP treatments showed significantly lower IgE and IgG values of the squid TMTp1 samples compared to our previous study.11 For example, the IgG (R) value of squid TMTp1 treated with a single-cycle of HHP at 200 MPa was 1.34, while the two-cycle treatment at the same pressure resulted in a significantly lower binding of 1.15.| Treatment group | IgE binding* | IgG binding# | ||||||
|---|---|---|---|---|---|---|---|---|
| P1 | P2 | P3 | P4 | P5 | R | R-SGF | R-SIF | |
| a ■T-200, T-400 and T-600 are the two-cycle HHP treatments, comprised of two 10 min cycles at 200, 400 and 600 MPa, respectively. *P1–P5: indirect ELISAs performed with human sera of five allergic patients. #R: indirect ELISA performed with rabbit anti-squid TMTp1 polyclonal antibodies. R-SGF: indirect ELISA performed with rabbit anti-squid TMTp1 polyclonal antibodies for hydrolysates after Simulated Gastric Fluid (SGF) digestion. R-SIF: indirect ELISA performed with rabbit anti-squid TMTp1 polyclonal antibodies for hydrolysates after Simulated Intestinal Fluid (SIF) digestion. a–d Values are expressed as mean ± standard deviation (SD) (n = 3); means in the same column with different lowercase letters (a–d) are significantly different (p < 0.05). | ||||||||
| Control | 1.67 ± 0.04a | 0.82 ± 0.01a | 0.74 ± 0.01a | 0.55 ± 0.06a | 0.47 ± 0.02a | 1.66 ± 0.03a | 1.37 ± 0.06a | 0.67 ± 0.03a |
| T-200■ | 1.08 ± 0.07b | 0.28 ± 0.01b | 0.31 ± 0.02b | 0.24 ± 0.03b | 0.28 ± 0.01b | 1.15 ± 0.02bc | 0.90 ± 0.03bc | 0.41 ± 0.02b |
| T-400 | 0.98 ± 0.05c | 0.26 ± 0.02bc | 0.28 ± 0.03bc | 0.23 ± 0.04b | 0.25 ± 0.01c | 1.12 ± 0.03cd | 0.86 ± 0.04cd | 0.42 ± 0.03b |
| T-600 | 0.87 ± 0.05d | 0.24 ± 0.01c | 0.25 ± 0.02c | 0.22 ± 0.02b | 0.25 ± 0.01c | 1.09 ± 0.01d | 0.84 ± 0.02d | 0.37 ± 0.02c |
Recent research has shown that the activity of phytoferritin (i.e., the iron release activity) is dramatically enhanced after HHP treatment,41 and the immunoreactivity of soybean seeds is reduced by HHP treatment at 300 MPa for 15 min, maintained at 40 °C.9 Also, HHP has been shown to induce structural unfolding, protein denaturation, and even dissociation of some proteins into subunits to change their functional properties.36,42 Meanwhile, multiple cycles of HHP have been shown to produce a more powerful percussive action and shear effect, resulting in a significant enhancement of the HHP effectiveness.23,43,44
In this study, the two-cycle HHP treatment caused a more pronounced alteration of the protein structure of squid TMTp1 than single-cycle HHP (Fig. 1–3) and these changes caused lower IgE and IgG values. These results indicate that HHP could decrease the allergenic properties of squid TMTp1, and that increasing the pressure level, as well as performing two-cycles of HHP, creates more significant effects on controlling the allergenic properties.
3.4 Correlation among AFM surface topography, NMR characteristic domain and allergenicity of HHP treated squid TMTp1
Correlation analyses among the pressure level, cycle numbers, IgE, IgG and AFM surface topography of the squid TMTp1 samples are presented in ESI Table S2.† The correlations among the IgE and IgG values (i.e., the allergenic properties) are really high (R ≥ 0.79, p < 0.001). The negative correlations between the pressure levels and the IgE values were all significant (p < 0.001), and there was also a significant negative correlation between the pressure levels and the IgG values (p < 0.01). There were negative correlations between the cycle numbers and both skewness (R = −0.80, p < 0.001) and kurtosis (R = −0.85, p < 0.001), but the cycle numbers were positively correlated with Ra (R = 0.90, p < 0.001) and Rmax (R = 0.88, p < 0.001). These results demonstrated that the pressure level and cycle number had a significant correlation with the modification of the allergenic and AFM surface topography measurements in squid TMTp1. In accordance with this, the changes in the allergenic properties, caused by the HHP treatments, were significantly correlated to the AFM surface topography indices.Correlation analyses between the NMR results, pressure level, cycle numbers, allergenic and AFM surface topography for the squid TMTp1 samples are presented in ESI Table S1.† The relationships between the HHP pressure and NMR P0.99–1.03, P1.79–1.84, P2.85–2.90, P3.36–3.39 and P3.40–3.43 indices were all significant (p < 0.05); the correlations of these NMR indices with the HHP mode were also significant (p < 0.05). This indicates that the HHP pressure and HHP mode significantly impacted upon some NMR characteristic regions of the squid TMTp1 samples. The main ranges of the characteristic regions for the allergenic properties were 0.99–1.03, 1.19–1.24, 1.44–1.49, 1.79–1.84, 2.85–2.90, 3.36–3.39 and 3.40–3.43 ppm (Fig. 3). For example, there was a peak at 0.99 ppm, in the range of 0.99–1.03 ppm, for the squid TMTp1 sample of the control treatment, however, there was no peak in this range for the HHP treated samples (Table 2). The peak at 0.99 ppm in this study was the trimethylamine signal, a proton of –CH3, and a multiple of (t, 7).45 This means that HHP changed the –CH3 group, which led to the modification of the allergenic properties (IgE and IgG values). The main allergenic characteristic peaks that were affected by HHP were 0.99, 1.16, 1.21, 1.79, 1.82, 2.84, 2.88 and 3.37 ppm, as observed in the control 1H NMR spectra (Table 2). It is important to note that the HHP mode (cycle numbers) is highly relevant to the changes in the NMR data, which resulted in the variation of the allergenic properties and AFM surface topographies.
There were negative correlations between P0.99–1.03 and both Ra and Rmax (R ≥ 0.76, p < 0.001), as well as P1.49–1.53 and both Ra and Rmax (R ≥ 0.74, p < 0.001); however, P3.32–3.36 and P3.36–3.39 were positively correlated with Ra and Rmax (R ≥ 0.84, p < 0.001). The AFM indices of skewness (S) and kurtosis (K) contradicted the results, compared to the tendencies of the variation in Ra and Rmax. The ranges of the main characteristics for the AFM surface topography were 1.04–1.08, 1.49–1.53, 1.64–1.69, 1.74–1.79, 3.15–3.20, 3.24–3.27, 3.32–3.36 and 3.36–3.39 ppm. Furthermore, the peaks of the main characteristics for the AFM surface topography that were affected by HHP were 1.06, 1.79, 3.23, 3.27 and 3.37 ppm, as identified in the control 1H NMR spectra (Table 2). For example, there was a peak at 3.37 ppm, in the range of 3.32–3.39 ppm, for the control squid TMTp1 samples, however, there were peaks at 3.35 ppm in this range for all of the HHP treated samples (Table 2). The peak at 3.37 ppm in this study was a characteristic signal of dimethyl ether, a proton of –CH3, and a multiple of (s).45 This means that HHP induced this –CH3 group migration and led to changes in the AFM surface topography (Ra, Rmax, S and K indices).
The results of this study reveal that the changes (by migration or disappearance) of related NMR characteristic regions, caused by the HHP treatments, significantly impacted upon the allergenic properties and AFM surface topography of squid TMTp1. The HHP mode significantly correlated (p < 0.05) with the variation in the NMR indices, which resulted in a reduction of the allergenicity and changes to the structure of the squid TMTp1 samples.
4. Conclusion
HHP treatments resulted in the modification (by migration or disappearance) of related NMR characteristic regions in squid TMTp1 samples, which in turn caused a reduction in the allergenicity and changes in the AFM surface topography. The 400 MPa and 600 MPa single-cycle HHP treatments to the squid TMTp1 samples caused significant increases in the roughness and maximum height, and the surface topography was no longer uniform in comparison with the control. However, the single-cycle HHP treatments gave rise to higher skewness and kurtosis values, while two-cycle HHP treatments led to lower values. In addition, the two-cycle HHP treatments significantly controlled the allergen (lower R values of 1.15–1.09).The HHP treatments affected squid TMTp1 by modifying some of its related NMR characteristic regions, which was apparent from the NMR spectroscopic analysis. The cycle numbers (mode) of the HHP treatments significantly correlated (p < 0.05) with the NMR indices, which resulted in reductions in the allergenicity and changes in the surface topography of the squid TMTp1 samples. The peaks of the main allergenic characteristics affected by HHP were 0.99, 1.16, 1.21, 1.79, 1.82, 2.84, 2.88, and 3.37 ppm (as identified in the control 1H NMR spectra), and the peaks of the AFM characteristics were 1.06, 1.79, 3.23, 3.27, and 3.37 ppm. The NMR spectra demonstrated that HHP treatments affected the structural characteristics of the squid TMTp1 samples by impacting on the main characteristic regions; by increasing the pressure level and performing two cycles (instead of one), the HHP treatment was more effective. This study provided meaningful information for the use of HHP as a non-thermal, minimal processing technology to change some NMR characteristic regions and control seafood allergens.
Acknowledgements
This research was supported by the National Natural Science Foundation of China (No. 31271955). The authors would like to give their thanks to SJTU-Instrumental Analysis Center for expert assistance with the AFM and NMR experiments.References
- S. B. Lehrer, R. Ayuso and G. Reese, Mar. Biotechnol., 2003, 5, 339–348 CrossRef CAS PubMed.
- C. Gamez, M. Zafra, M. Boquete, V. Sanz, C. Mazzeo, M. D. Ibanez, S. Sanchez-Garcia, J. Sastre and V. del Pozo, Mol. Nutr. Food Res., 2014, 58, 1915–1925 CAS.
- N. Y. Leung, C. Y. Wai, S. Shu, J. Wang, T. P. Kenny, K. H. Chu and P. S. Leung, Clin. Rev. Allergy Immunol., 2014, 46, 180–197 CrossRef CAS PubMed.
- S. Benede, I. L. Exposito, E. Molina and R. L. Fandino, Food Funct., 2015, 6, 694–713 CAS.
- E. Zhou, Y. Fu, Z. Wei and Z. Yang, Food Funct., 2014, 5, 2106–2112 CAS.
- S. Jung, M.-S. Lee, Y. Shin, C. T. Kim, I.-H. Kim, Y. S. Kim and Y. Kim, J. Funct. Foods, 2014, 10, 169–177 CrossRef CAS PubMed.
- J. Somkuti and L. Smeller, Biophys. Chem., 2013, 183, 19–29 CrossRef CAS PubMed.
- R. Chicon, J. Belloque, E. Alonso, P. J. M. Alvarez and R. L. Fandino, J. Food Prot., 2008, 71, 1453–1459 CAS.
- E. Penas, R. Gomez, J. Frias, M. L. Baeza and C. V. Valverde, Food Chem., 2011, 125, 423–429 CrossRef CAS PubMed.
- S. Dhakal, C. Liu, Y. Zhang, K. H. Roux, S. K. Sathe and V. Balasubramaniam, Food Res. Int., 2014, 62, 215–222 CrossRef CAS PubMed.
- Y. Jin, Y. Deng, B. Qian, Y. Zhang, Z. Liu and Y. Zhao, Food Chem. Toxicol., 2015, 76, 86–93 CrossRef CAS PubMed.
- A. L. Lee and A. J. Wand, Nature, 2001, 411, 501–504 CrossRef CAS PubMed.
- J. Kinsella, Food Proteins, 1982, 1, 51–103 Search PubMed.
- X. Yang and R. A. Mariuzza, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 8166–8167 CrossRef CAS PubMed.
- L. A. Abramovich and E. Gazit, Chem. Soc. Rev., 2014, 43, 7236 RSC.
- K. Bazaka, M. Jacob, W. Chrzanowski and K. Ostrikov, RSC Adv., 2015, 5, 48739–48759 RSC.
- H. Yang, Y. Wang, S. Lai, H. An, Y. Li and F. Chen, J. Food Sci., 2007, 72, R65–R75 CrossRef CAS PubMed.
- R. Kuisma, E. P. Leinonen, I. Redsven, H. R. Kymalainen, I. Saarikoski, A. M. Sjoberg and M. Hautala, Surf. Sci., 2005, 584, 119–125 CrossRef CAS PubMed.
- R. A. Gittens, T. McLachlan, R. O. Navarrete, Y. Cai, S. Berner, R. Tannenbaum, Z. Schwartz, K. H. Sandhage and B. D. Boyan, Biomaterials, 2011, 32, 3395–3403 CrossRef CAS PubMed.
- M. Christensen, J. T. Rasmussen and A. C. Simonsen, Food Hydrocolloids, 2015, 45, 168–174 CrossRef CAS PubMed.
- K. Kieswetter, Z. Schwartz, T. Hummert, D. Cochran, J. Simpson, D. Dean and B. Boyan, J. Biomed. Mater. Res., 1996, 32, 55–63 CrossRef CAS.
- A. L. Raines, R. O. Navarrete, M. Wieland, D. L. Cochran, Z. Schwartz and B. D. Boyan, Biomaterials, 2010, 31, 4909–4917 CrossRef CAS PubMed.
- Y. Zhang, S. Jiao, Z. Lian, Y. Deng and Y. Zhao, J. Food Sci., 2015, 80, E1012–E1020 CrossRef CAS PubMed.
- Y. Zhang, G. Wang, Y. Jin, Y. Deng and Y. Zhao, Food Control, 2016, 60, 189–195 CrossRef CAS PubMed.
- J. Yue, Y. Zhang, Y. Jin, Y. Deng and Y. Zhao, Food Chem., 2016, 194, 12–19 CrossRef CAS PubMed.
- M. Kainosho, T. Torizawa, Y. Iwashita, T. Terauchi, A. M. Ono and P. Guntert, Nature, 2006, 440, 52–57 CrossRef CAS PubMed.
- S. Alessandri, A. Sancho, S. Vieths, C. E. Mills, J. M. Wal, P. R. Shewry, N. Rigby and K. H. Sommergruber, PLoS One, 2012, 7, e39785 CAS.
- G. Hu, Y. Zheng, D. Wang, B. Zha, Z. Liu and Y. Deng, High Pressure Res., 2015, 35, 330–338 CrossRef CAS PubMed.
- Y. Deng, Y. Jin, Y. Luo, Y. Zhong, J. Yue, X. Song and Y. Zhao, J. Cereal Sci., 2014, 60, 302–310 CrossRef CAS PubMed.
- H. Hsu, S. Sheen, J. Sites, J. Cassidy, B. Scullen and C. Sommers, Food Microbiol., 2015, 48, 1–7 CrossRef CAS PubMed.
- P. Giraudeau, I. Tea, G. S. Remaud and S. Akoka, J. Pharm. Biomed. Anal., 2014, 93, 3–16 CrossRef CAS PubMed.
- C. K. Loong, H.-X. Zhou and P. B. Chase, PLoS One, 2012, 7, e39676 CAS.
- C. K. Loong, H. X. Zhou and P. B. Chase, FEBS Lett., 2012, 586, 3503–3507 CrossRef CAS PubMed.
- A. Rodriguez, W. R. Autio and L. A. McLandsborough, J. Food Prot., 2008, 71, 170–175 Search PubMed.
- D. Lampe, U. Strijowski, V. Heinz and U. Bindrich, J. Cereal Sci., 2013, 57, 411–417 CrossRef CAS PubMed.
- J. Somkuti and L. Smeller, Biophys. Chem., 2013, 183, 19–29 CrossRef CAS PubMed.
- S. D. Halleux, E. Stura, L. VanderElst, V. Carlier, M. Jacquemin and J. M. S. Remy, J. Allergy Clin. Immunol., 2006, 117, 571–576 CrossRef PubMed.
- C. de Kruif, R. Tuinier, C. Holt, P. Timmins and H. Rollema, Langmuir, 2002, 18, 4885–4891 CrossRef CAS.
- S. L. Chan, S. T. Ong, S. Y. Ong, F. T. Chew and Y. K. Mok, J. Immunol., 2006, 176, 4852–4860 CrossRef CAS.
- D. P. Uceda, O. Palomares, M. Bruix, M. Villalba, R. Rodriguez, M. Rico and J. Santoro, Biochemistry, 2004, 43, 16036–16045 CrossRef PubMed.
- T. Zhang, C. Lv, S. Yun, X. Liao, G. Zhao and X. Leng, Food Chem., 2012, 130, 273–278 CrossRef CAS PubMed.
- J. L. Silva, Y. Cordeiro and D. Foguel, Biochim. Biophys. Acta,Proteins Proteomics, 2006, 1764, 443–451 CrossRef CAS PubMed.
- A. L. Curl and E. F. Jansen, J. Biol. Chem., 1950, 185, 713–723 CAS.
- A. L. Curl and E. F. Jansen, J. Biol. Chem., 1950, 184, 45–54 CAS.
- G. R. Fulmer, A. J. M. Miller, N. H. Sherden, H. E. Gottlieb, A. Nudelman, B. M. Stoltz, J. E. Bercaw and K. I. Goldberg, Organometallics, 2010, 29, 2176–2179 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra13655e |
| This journal is © The Royal Society of Chemistry 2015 |