LTQ-Orbitrap-based strategy for traditional Chinese medicine targeted class discovery, identification and herbomics research: a case study on phenylethanoid glycosides in three different species of Herba Cistanches†
Jiayu Zhanga,
Chen Lib,
Yanyun Chec,
Jiarui Wua,
Zijian Wanga,
Wei Caia,
Yun Lia,
Zhiguo Ma*d and
Pengfei Tu*a
aBeijing University of Chinese Medicine, Beijing 100029, China. E-mail: pengfeitu@vip.163.com; Tel: +86 01082802750
bThermo Fisher Scientific, Shanghai 201206, China
cYunnan University of Traditional Chinese Medicine, Kunming 650500, China
dCollege of Pharmacy, Jinan University, Guangzhou 510632, China. E-mail: mzg79@hotmail.com; Tel: +86 02085223784
First published on 17th September 2015
Abstract
Traditional Chinese Medicine (TCM) usually contains one specific class of components that could represent its phytochemical properties and therapeutic effects. However, the unclear specific class constituents hinders the sufficient interpretation of its bioactivities. In this paper, an HPLC-LTQ-Orbitrap-based strategy focused on the TCM targeted class was developed. This strategy was successfully applied in the discovery and identification of phenylethanoid glycosides (PhGs) in Herba Cistanches, and herbomics research of Cistanche deserticola, C. sinensis, and C. tubulosa. A total of 69 PhGs, including 17 new PhGs, were rapidly discovered and characterized using multiple data-mining methods such as accurate parent mass search, parent mass modification search and fragment ion search, allowing a comprehensive revelation of PhGs in Cistanche species for the first time. Based on the peak areas of the 69 PhGs identified, multivariate statistical analysis such as PCA, loading analysis and OPLS-DA were employed in herbomics research to screen and validate the PhGs that could be utilized to discriminate these three Cistanche species. Eight PhGs were finally screened and chosen as chemical markers for the species discrimination. In conclusion, this new established strategy could be exemplary for future studies on the discovery and identification of important chemical constituents, and differentiation of genuine specie or genus from adulterants. Meanwhile, it may propose a novel idea for analyzing a specific class of active chemical constituents, and is promising for quality control and evaluation of TCMs.
1. Introduction
Cistanche (Roucongrong in Chinese), well-known as “ginseng of the desert”, is a worldwide genus of holoparasitic desert plant, and mainly distributed in the arid and warm deserts in northwest China.1 As an important tonic agent in traditional Chinese medicines (TCMs), the dried succulent stems of Cistanche deserticola Y. C. Ma and C. tubulosa (Schrenk) Wig as well as C. sinensis (C. A. Mey.) G. Beck have been used for a long time to treat kidney deficiency, female infertility, morbid leucorrhea, neurasthenia, and senile constipation due to colonic inertia. However, the three plant origins of Herba Cistanches are different in terms of the pharmacological activities and chemical components.2–4 As for the clinical application and market circulation, C. tubulosa has been traditionally used as a blood circulation-promoting agent and in the treatment of impotence, sterility, lumbago, and body weakness.5,6 Consequently, it is crucial to establish a method to discriminate three different plant origins of Cistanches Herba to guarantee the quality control and clinical application.Many common analytical methods, including microscopy, ultraviolet, infrared detection, and inter simple sequence repeats method have been developed to classify the genus of Cistanches.7–13 Meanwhile, several high-performance liquid chromatography (HPLC) and HPLC-MS methods have been recently reported on the analysis of phenylethanoid glycosides (PhGs) in different Herba Cistanches, but the markers and/or samples and/or species used in the experiment are apparent and insufficient.14–17 We established a method to perform quantitative analysis of ten major phenylethanoid glycosides using HPLC coupled with diode array detection (DAD) and chemometrics methods, which could lead to successful classification of three Cistanches Herba in accordance with their origins.18 However, it mainly emphasized the differences among the limited marker compounds, thus could not provide a comprehensive comparison among various species. Moreover, although the separation and identification of PhGs contained in Cistanches Herba with phytochemistry methods have been developed,19 the previous LC-MS analytical results demonstrated that numerous known and unknown PhGs have not been comprehensively screened and characterized yet.15,16
Recently, a new hybrid LTQ-Orbitrap high-resolution (HR) MS analytical platform is applied to the analysis of small molecules in biological and TCM samples.20–22 It consists of a linear ion trap and an Orbitrap, and allows HRMS and multi-stage MS data to be acquired simultaneously. The Orbitrap mass spectrometer, otherwise defined as an electrostatic Fourier transform mass spectrometer, provides a much higher mass resolution and accuracy (<3 ppm) than any other mass spectrometers.23 This advantage facilitates the identification of the TCM constituents especially the unknown microconstituents.
In this study, a novel HPLC-LTQ-Orbitrap-based strategy was established for targeted class constituents discovery, identification, and herbomics research. Three species of Cistanches Herba, with PhGs as the specific class, were used as an example to verify the effectiveness of this strategy.
2. Experimental
2.1. Standards and reagents
Ten PhGs standards, including acteoside, isoacteoside, echinacoside, cistanoside A, poliumoside, cistanoside C, isocistanoside C, tubuloside B, 2′-O-acetylacteoside, and 2′-O-acetylpoliumoside, were isolated from Cistanche plants by silica gel column chromatography eluting in our laboratory. Their structures (shown in Fig. 1) were determined by NMR and MS.24,25 The purities of all compounds were determined to be no less than 98% according to the normalization of the peak area detected by HPLC-DAD method.HPLC-grade acetonitrile and methanol were purchased from Fisher Scientific (Fair Lawn, NJ, USA). Formic acid was purchased from Sigma Aldrich (St. Louis, MO, USA). Deionized water used throughout the experiment was purified by a Milli-Q Gradient A 10 System (Millipore, Billerica, MA, USA). The 0.22 μm membranes were purchased from Xinjinhua Co. (Shanghai, China).
2.2. Crude plant material and sample preparation
Seventeen batches of Cistanches Herba (10 C. deserticola and 7 C. tubulosa) were collected by the authors from Xinjiang and Inner Mongolia provinces, the indigenous cultivating regions of Cistanches Herba; the other nineteen batches (4 C. deserticola, 3 C. tubulosa, and 12 C. sinensis) were purchased from different pharmacies in Guangdong, Anhui, and Shandong provinces of China (illustrated in Table 1). All the identities of Cistanches Herba samples were respectively authenticated to be dried succulent stems of C. deserticola, C. tubulosa, and C. sinensis by morphological and histological methods by Dr Zhi-Guo Ma. The voucher specimens were deposited at Center of Scientific Experiment, Beijing University of Chinese Medicine, Beijing city, China.| Sample | Codea | Sources | Collection date | Sample | Codea | Sources | Collection date |
|---|---|---|---|---|---|---|---|
| a CD, C. deserticola; CT, C. tubulosa; CS, C. sinensis.b The analysts are the same as in Fig. 3 and Table 2.c The analysts are selected at random for the verification test. | |||||||
| 1b | CD-1 | Changji, Xinjiang | November 2011 | 19 | CT-5 | South Xinjiang | November 2009 |
| 2 | CD-2 | Changji, Xinjiang | November 2011 | 20 | CT-6 | South Xinjiang | November 2009 |
| 3 | CD-3 | Changji, Xinjiang | March 2012 | 21 | CT-7 | Changji, Xinjiang | May 2010 |
| 4 | CD-4 | Changji, Xinjiang | November 2011 | 22 | CT-8 | Yantai, Shandong | February 2010 |
| 5 | CD-5 | Changji, Xinjiang | April 2012 | 23c | CT-9 | Jinan, Shandong | June 2010 |
| 6c | CD-6 | Changji, Xinjiang | November 2011 | 24 | CT-10 | Guangzhou, Guangdong | June 2010 |
| 7 | CD-7 | Alashan, Inner Mogolia | February 2012 | 25b | CS-1 | Guangzhou, Guangdong | June 2010 |
| 8 | CD-8 | Bozhou, Anhui | March 2010 | 26 | CS-2 | Guangzhou, Guangdong | June 2010 |
| 9 | CD-9 | Changji, Xinjiang | May 2011 | 27 | CS-3 | Guangzhou, Guangdong | June 2010 |
| 10c | CD-10 | Alashan, Inner Mogolia | November 2012 | 28 | CS-4 | Guangzhou, Guangdong | June 2010 |
| 11 | CD-11 | Guangzhou, Guangdong | October 2009 | 29 | CS-5 | Guangzhou, Guangdong | June 2010 |
| 12 | CD-12 | Tacheng, Xinjiang | May 2011 | 30 | CS-6 | Guangzhou, Guangdong | June 2010 |
| 13 | CD-13 | Guangzhou, Guangdong | May 2011 | 31 | CS-7 | Guangzhou, Guangdong | June 2010 |
| 14c | CD-14 | Guangzhou, Guangdong | May 2011 | 32c | CS-8 | Guangzhou, Guangdong | June 2010 |
| 15b,c | CT-1 | South Xinjiang | November 2009 | 33 | CS-9 | Guangzhou, Guangdong | June 2010 |
| 16 | CT-2 | South Xinjiang | November 2009 | 34 | CS-10 | Guangzhou, Guangdong | June 2010 |
| 17 | CT-3 | South Xinjiang | November 2009 | 35 | CS-11 | Guangzhou, Guangdong | June 2010 |
| 18 | CT-4 | South Xinjiang | November 2009 | 36c | CS-12 | Guangzhou, Guangdong | June 2010 |
The dried powders of Cistanches Herba samples were grounded and sieved through a no. 65 mesh sieve. After accurately weighing and soaking for 30 min, an amount of 1.0 g was extracted with 50 mL of methanol/water (75![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25, v/v) in an ultrasonic bath (40 kHz, Eima Ultrasonics Corp., Germany) for 30 min at room temperature, and then the same solvent was added to compensate for the lost weight during the extraction. After centrifugation (10
25, v/v) in an ultrasonic bath (40 kHz, Eima Ultrasonics Corp., Germany) for 30 min at room temperature, and then the same solvent was added to compensate for the lost weight during the extraction. After centrifugation (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm, 10 min), the methanol solution was filtered through a 0.22 μm microporous membrane before injection to LC-MS system for analysis.
000 rpm, 10 min), the methanol solution was filtered through a 0.22 μm microporous membrane before injection to LC-MS system for analysis.
2.3. Standard solutions preparation
The appropriate amount of each standard was weighed and dissolved in methanol to make ten individual stock solutions. Then, each stock solution was mixed with methanol to prepare a final mixed standard solution.2.4. HPLC-LTQ-Orbitrap analysis
Thermo Scientific Accela 600 HPLC system used in the experiment equipped with a binary pump and an autosampler. An Agilent Zorbax Extended C18 (250 × 4.6 mm i.d., 5 μm) was used for separation of the PhGs at room temperature. 0.1% formic acid aqueous solution (solvent A) and acetonitrile (solvent B) were used as mobile phase. The flow rate was 1.0 mL min−1 and elution conditions at room temperature applied with a linear gradient as follows: 0–20 min, 10–35% B; 20–32 min, 35% B; 32–55 min, 35–42% B; 55–75 min, 42–48% B with a 10 minute post run.HRMS and MS/MS spectral analysis were performed on an LTQ-Orbitrap mass spectrometer (Thermo Scientific, Bremen, Germany). The mass spectrometer was connected to the HPLC instrument via an ESI interface in a post-column splitting ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4. Samples were analyzed in the negative ion mode with a tune method set as follows: sheath gas (nitrogen) flow rate of 30 arb, aux gas (nitrogen) flow rate of 5 arb, spray voltage of 4.0 kV, capillary temperature of 350 °C, capillary voltage of 25 V, tube lens voltage of 110 V. Accurate mass analysis were calibrated using a standard solution mixture of caffeine, sodium dodecyl sulfate, sodium taurocholate, the tetrapeptide MRFA acetate salt, and Ultramark. The measured masses were within 3 ppm of the theoretical masses. Centroided mass spectra were acquired in the mass range of m/z 100–1000.
4. Samples were analyzed in the negative ion mode with a tune method set as follows: sheath gas (nitrogen) flow rate of 30 arb, aux gas (nitrogen) flow rate of 5 arb, spray voltage of 4.0 kV, capillary temperature of 350 °C, capillary voltage of 25 V, tube lens voltage of 110 V. Accurate mass analysis were calibrated using a standard solution mixture of caffeine, sodium dodecyl sulfate, sodium taurocholate, the tetrapeptide MRFA acetate salt, and Ultramark. The measured masses were within 3 ppm of the theoretical masses. Centroided mass spectra were acquired in the mass range of m/z 100–1000.
In the full scan experiment, resolution of the Orbitrap mass analyzer was set at 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. Data-dependent MS/MS scanning was performed to minimize total analysis time as it can trigger fragmentation reactions of target ions. The collision energy for CID was adjusted to 40% of maximum, and the isolation width of precursor ions was m/z 2.0 Da. The dynamic exclusion to prevent repetition was enabled, and the repeat count was set at 5 with the dynamic repeat time at 30 s and dynamic exclusion duration at 60 s.
000. Data-dependent MS/MS scanning was performed to minimize total analysis time as it can trigger fragmentation reactions of target ions. The collision energy for CID was adjusted to 40% of maximum, and the isolation width of precursor ions was m/z 2.0 Da. The dynamic exclusion to prevent repetition was enabled, and the repeat count was set at 5 with the dynamic repeat time at 30 s and dynamic exclusion duration at 60 s.
2.5. Peak selections and data processing
Thermo Xcalibur 2.1 workstation was used for the data acquiring and processing. In order to obtain as many fragments as possible, the peaks detected with intensity over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 were selected for identifications. The chemical formulas for all parent and fragment ions of the selected peaks were calculated from the accurate mass using a formula predictor by setting the parameters as follows: C [0–30], H [0–50], O [0–20], and Ring Double Bond (RDB) equivalent value [0–15]. Other elements such as N, P, S, Cl, and Br were not considered as they are rarely present in this traditional herb. A serial of data-processing software, including Tracefinder, MetWorks and Mass Frontier (Thermo Scientific, Bremen, Germany) were used for peak area extraction, data-filtering, and identification in the experiment.
000 were selected for identifications. The chemical formulas for all parent and fragment ions of the selected peaks were calculated from the accurate mass using a formula predictor by setting the parameters as follows: C [0–30], H [0–50], O [0–20], and Ring Double Bond (RDB) equivalent value [0–15]. Other elements such as N, P, S, Cl, and Br were not considered as they are rarely present in this traditional herb. A serial of data-processing software, including Tracefinder, MetWorks and Mass Frontier (Thermo Scientific, Bremen, Germany) were used for peak area extraction, data-filtering, and identification in the experiment.
2.6. Method validation and statistical analysis
Method repeatability and precision were assessed by analysis of the ten known PhGs in three independently prepared samples (no. 1, 15, and 25) six times over 48 hours, respectively. The relative standard deviation (RSD) of peak area was taken as the indicator for method validation. As a multivariate analysis technique,26,27 unsupervised PCA, supervised OPLS-DA, and Hierarchical clustering analysis (HCA) methods were respectively displayed by SPSS statistics 17.0 (SPSS Inc., IL, USA) and SIMCA-P 13.0 (Umetrics AB, Umea, Sweden) to bring insight into the chemical differences among three different Cistanches plants.3. Results and discussion
3.1. Analytical strategy for PhGs discovery, identification, and herbomics
A novel strategy was proposed for the systematic discovery, characterization of PhGs, and herbomics research of three Cistanches plant origins using multiple data acquisition and processing techniques.Firstly, the data-dependent acquisition was performed on HPLC-LTQ-Orbitrap, HRMS and MS3 data sets of high quality in negative mode were obtained. Then a high throughput screening assays was developed for quickly and accurately filtering the known and unknown PhGs candidate peaks by TraceFinder, MetWorks and Mass Frontier software. For the known PhGs, a chemical formula database of the reported PhGs was built in an Excel file and imported to the TraceFinder software. Then the known PhGs candidates were searched out by the TraceFinder software according to the accurate mass of [M − H]− ion. For the unknown PhGs, the candidates were screened out from MS1 to MSn level by MetWorks and Mass Frontier software. Constituents belonging to one specific class usually share the same parent structure, thus the targeted class constituents could be discovered based on parent structure and substituent modifications (MS1 level), or fragments of parent structure (MSn level). For the MS1 level search, a custom modification list including common modifications such as hydroxylation and methylation was built in MetWorks software. Then the extracted ion chromatograms (EICs) of each PhG candidate were generated automatically based on the calculated accurate mass of modified parent structure. For the MSn level search, diagnostic product ions of PhG parent structure were input into Mass Frontier software. The eligible peaks were subsequently screened using fragment ion search (FISh) function. All these steps allow a comprehensive revelation of PhGs in Cistanches Herba. For the structure characterization, manual elucidation was confirmed by the Fragments and Mechanisms function of Mass Frontier software, which based on a fragmentation library containing abundant literature-proposed fragmentation mechanisms, giving more precise identification results.
For the herbomics research, the LC-MS data was first processed to extract information of the true analytical peaks in both HPLC and MS domains. A list of peak intensities (characterized chromatographic PhG peak areas) was then generated for the first chromatogram using the tR–m/z data pairs as identifiers using TraceFinder software. To determine the potential discriminatory PhGs markers among three Cistanches plant origins, an approach of chromatographic profiling analysis in combination with PCA data statistics method was exploited. Meanwhile, a supervised OPLS-DA statistical model was constructed with training sample and the PhGs markers were validated with the test sample, which were most likely considered to be the discriminatory markers attributing to sample classification. The general procedures of our strategy and approach are summarized into a diagram as illustrated in Fig. 2.
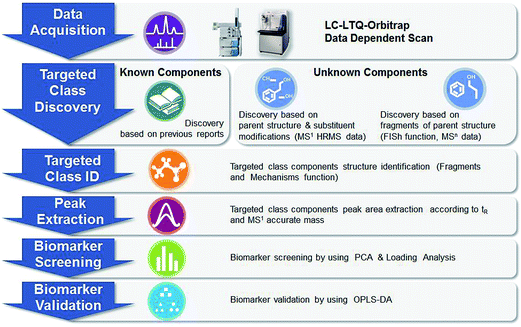 | ||
| Fig. 2 Summary diagram of presently developed analytical strategy and methodology for TCM target class discovery, identification, and herbomics research. | ||
3.2. Optimum conditions for HPLC-LTQ-Orbitrap analysis
In order to achieve satisfactory extraction efficiency for all the PhGs, the extraction conditions, including extraction methods (ultrasonication and refluxing), extraction solvents (50%, 75%, and 100% methanol), and extraction time (15, 30, and 45 min) were assessed based on orthogonal experiments. The best extraction efficiency was obtained by ultrasonication extraction with 75% methanol for 30 min.18 The different HPLC parameters including mobile phases (methanol/water and acetonitrile/water), the concentration of formic acid in water (0.1% and 0.3%), category of RP-ODS columns (Agilent Zorbax TC-C18 column, 250 × 4.6 mm i.d., 5 μm and Agilent Zorbax Eclipse XDB-C18 column, 250 × 4.6 mm i.d., 5 μm) were compared in parallel. The results presented that methanol was much more suitable for the chromatographic separation as mobile phase than acetonitrile; 0.1% formic acid was advantageous to obtain the best resolution of adjacent peaks and restrain the peak tailing of PhGs; TC-C18 column exhibited better peak separation than that of XDB-C18 column.PhGs under investigation are composed of saccharide and phenylethanol groups, which are the aglycons of these constituents. The reported PhGs almost include some substituents such as caffeoyl, coumaroyl, or feruloyl group, which usually replace the hydroxyl groups at 4- or 6-carbon of the central saccharide. These characteristics make it predestined for ESI in negative ion mode. Different MS conditions were optimized on LTQ-Orbitrap MS instrument using the standard solution of acteoside (3.62 μg mL−1). To achieve the optimized collision energy that generates adequate fragment ions for structural elucidation and characterization, a series of ESI-MS/MS experiments were carried out at different collision energy (CE, 10–100%). By gradually increasing the CE, the intensity of product ions was first increased to maximum and then gradually decreased. Even though the optimum CE might vary for different PhGs, the result demonstrated that 40% CE was adequate to yield abundant fragment ions for the structural elucidation.
3.3. Rapid discovery of targeted PhGs class
In previous report, PhGs are usually obtained by repeated column chromatography from Tubiflorae containing Oleaceae, Orobanchaceae, Verbenaceae, Lamiaceae, Scrophulariaceae, and Plantaginaceae. Although the phytochemical approach has provided a quantity of useful information to discovery new PhGs from various plants, it is time-consuming and laborious, and minor constituents are often neglected during the separation process. Here, LC-HRMS was adopted to screen and identify the PhGs in three different Herba Cistanches. After the acquisition of HRMS datasets, TraceFinder software providing a comprehensive system incorporating built-in method was adopted to discover the commonly known PhGs by searching the accurate mass of targeted parent ions. Meanwhile, since the multiple constituents contained in a certain traditional herb are usually derived from one or more certain biosynthetic pathways, which indicates these constituents could be structurally classified into several chemical families with same carbon skeletons and various substituents.28,29 MetWorks software, initially developed to automate and simplify the identification of drug metabolites, is adopted to discover PhGs, especially the unknown PhGs by defining parent nucleus and potential substitutes in this study. PhGs have regularity in elemental composition with the basic aglycone structure and substituents such as caffeoyl (162 Da, C9H6O3), feruloyl (176 Da, C10H8O3), coumaroyl (146 Da, C9H6O2), glucoside residue (162 Da, C6H10O5), rhamnose residue (146 Da, C6H10O4), and acetyl (42 Da, C2H2O). Accordingly, molecular weights of the basic aglycone structures are 284 Da (C14H20O6), which are increased by 162, 146, 176, or 42 when those substituents were attached, respectively. Hence, the screening table could be designed by arranging substituents at the molecular weights 284 Da from one to seven positions using the built-in software. As a result, more than 80 PhGs candidates were automatically screened from the large mass dataset using the Tracefinder and MetWorks software according the accurate mass of [M − H]− ions.3.4. Fragmentation behaviors of PhGs in negative ion mode
Mass Frontier software can predict and display comprehensive fragmentation pathways based on a set of general ionization, fragmentation, and rearrangement rules and by automatically extracting a decomposition mechanism for each fragmentation reaction in the fragmentation library. To facilitate the structural identification and differentiation of PhGs candidates, it was applied to propose the fragmentation behaviors of ten reference compounds representing the major structural types of the PhGs in Cistanches Herba. Taking isoacteoside for instance (shown in Fig. 3), it exhibited the [M − H]− ion at m/z 623.1968 (C29H34O15, error −0.32 ppm) in its ESI-MS spectrum. In the ESI-MS2 spectrum, it yielded [M − H − 162]− base peak ion at m/z 461 by neutral loss of a caffeoyl unit from the precursor ion according to the bibliography data.16 Meanwhile, the minor [M − H − rha]− ion at m/z 477 was also observed. In its ESI-MS3 spectrum, the base peak at m/z 315 was generated from the precursor ion m/z 461 by the neutral loss of rhamnose residue. Moreover, the minor ions including [461 − rha − H2O]− ion at m/z 297, [caffeic acid − H]− ion at m/z 179, [caffeic acid − H − H2O]− ion at m/z 161, and [caffeic acid − H − CO2]− ion at m/z 135 were also detected. In the following ESI-MS/MS experiments, the other nine PhGs reference compounds produced the similar patterns of MS/MS fragmentation. The neutral loss at m/z 162, 146, 60, or 42, were related to caffeic acid or glucose, rhamnose, acetic acid, or acetyl, while the elimination of a molecule of H2O or CO2 occasionally occurred in the fragmentations. Identical product ions at m/z 179, 161, and 135 were frequently observed and could serve as evidences for the existence of caffeoyl, anhydroglucose, anhydrorhamnose, and anhydrophenethanol.30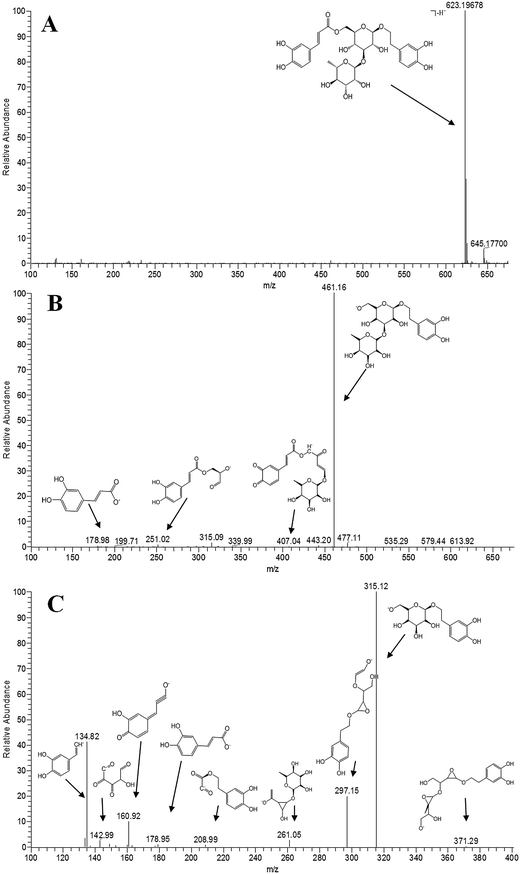 | ||
| Fig. 3 ESI-MSn spectra of isoacteoside in negative ion mode: (A) MS spectrum; (B) MS2 spectrum (precursor-ion was m/z 623); (C) MS3 spectrum (precursor-ion was m/z 461). | ||
3.5. Rapid assignment of PhGs chromatographic peaks in negative ion mode
The total ion current chromatograms of all samples were demonstrated in Fig. 1S.† Among them, no. 1, 15, and 25 were selected as the typical chromatography profiling for three different plant origins, respectively, as shown in Fig. 4. According to the results from fragmentation behaviors deduced, m/z 179, 161, and 135 were adopted as the criterion to further screen and identify candidate peaks for PhGs using the FISh technology built-in Mass Frontier software. Meanwhile, these DPIs generated from the neutral loss from parent ions including [M − H]−, [M − H − caffeoyl/feruloyl/coumaroyl]−, [M − H − glc/rha]−, [M − H − caffeoyl/feruloyl/coumaroyl − glc/rha]−, [M − H − CH3COOH]−, and [M − H − CH2CO]−, etc., were used as the diagnostic fragmentation pathways of PhGs, which were adopted to rapidly determine and validate the results for structure skeletons and substitution patterns of PhGs in the complex matrices.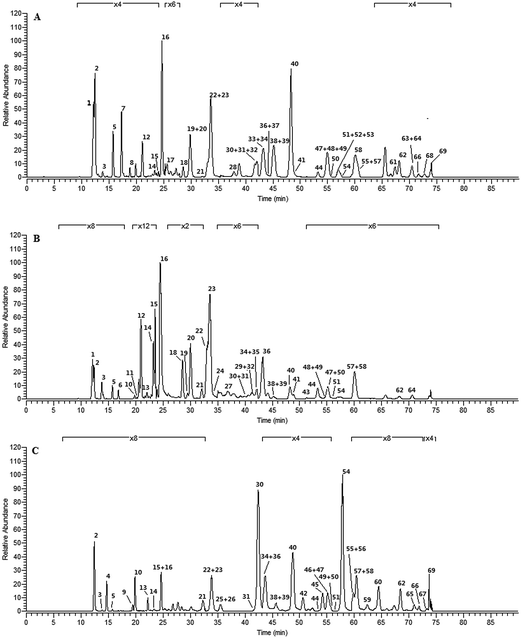 | ||
| Fig. 4 The representative total ion current chromatogram of three different Cistanche species: (A) CD; (B) CT; (C) CS. The names of these compounds correspond to those given in Table 2. | ||
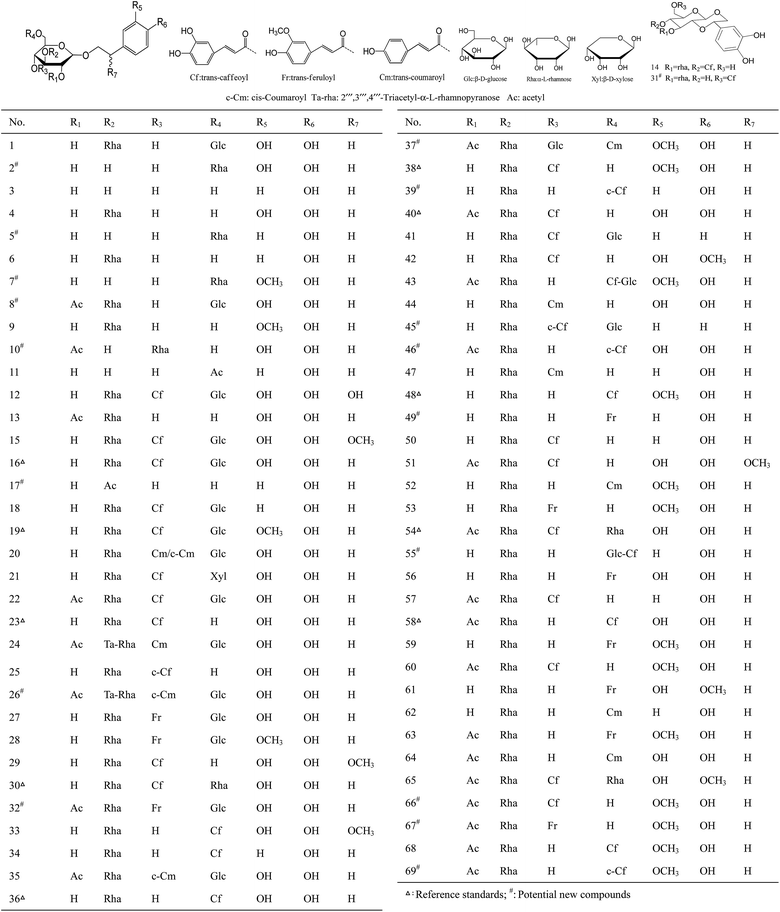
Finally, based on on-line LC-MSn data acquisition and off-line data processing methods, 69 PhGs were validated and identified from three different Herba Cistanches (47 from C. deserticola, 42 from C. salsa, and 42 from C. tubulosa). Among them, 10 PhGs were unambiguously confirmed as echinacoside (16), cistanoside A (19), acteoside (23), poliumoside (30), isoacteoside (36), cistanoside C (38), 2′-O-acetylacteoside (40), isocistanoside C (48), 2′-O-acetylpoliumoside (54), and tubuloside B (58) by comparison the retention time, accurate mass of [M − H]− ions, and ESI-MS/MS spectra with those of reference substances. Meanwhile, the other 59 compounds were tentatively characterized to be 42 known and 17 new PhGs by comparing their MS data with those reported in the literatures, fragmentation pathways deduced, and C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values (n-octanol/water partition coefficient, a parameter for determining the elution order31–33). For example, peak 25 elute at 34.32 min yielded the same [M − H]− ion at m/z 623.1969 (C29H35O15, error −0.16 ppm) as those of peak 23 (acteoside) and 36 (isocateoside). In its MS/MS spectra, the main fragment ions at m/z 461 and 477 were generated from elimination of caffeoyl unit and rhamnose residue from the parent ion. The fragment ion at m/z 443 came from the successive neutral loss of H2O from m/z 461 owing to deoxidation from side chain hydroxyl. The fragment ion at m/z 315 was generated from the successive elimination of caffeoyl and rhamnose residue from the parent ion. The fragment ion at m/z 135 was from the dehydrating residues of anhydrophenethanpl after loss of all the side chains. These characteristic fragmentation pathways were similar with those of acteoside (C
P values (n-octanol/water partition coefficient, a parameter for determining the elution order31–33). For example, peak 25 elute at 34.32 min yielded the same [M − H]− ion at m/z 623.1969 (C29H35O15, error −0.16 ppm) as those of peak 23 (acteoside) and 36 (isocateoside). In its MS/MS spectra, the main fragment ions at m/z 461 and 477 were generated from elimination of caffeoyl unit and rhamnose residue from the parent ion. The fragment ion at m/z 443 came from the successive neutral loss of H2O from m/z 461 owing to deoxidation from side chain hydroxyl. The fragment ion at m/z 315 was generated from the successive elimination of caffeoyl and rhamnose residue from the parent ion. The fragment ion at m/z 135 was from the dehydrating residues of anhydrophenethanpl after loss of all the side chains. These characteristic fragmentation pathways were similar with those of acteoside (C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P: −0.890) and isoacteoside (C
P: −0.890) and isoacteoside (C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P: −0.057). Because the retention time of 25 was almost same as that of acteoside, and thus it was tentatively characterized as cisacteoside (C
P: −0.057). Because the retention time of 25 was almost same as that of acteoside, and thus it was tentatively characterized as cisacteoside (C![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P: −0.890). Likely, structures of the other chromatographic peaks were tentatively elucidated. All the results were demonstrated in Fig. 1 and Table 2.
P: −0.890). Likely, structures of the other chromatographic peaks were tentatively elucidated. All the results were demonstrated in Fig. 1 and Table 2.
| Peak | tR (min) | Formula | Theoretical mass (m/z) | Experimental mass (m/z) | Mass error (ppm) | ESI-MS2 (m/z) | ESI-MS3 (m/z) | Structural identification | Origins | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| CD | CT | CS | |||||||||
| a PhGs identified by comparison with reference standards.b New PhGs never reported in the bibliography data.c Being detected in the sample. | |||||||||||
| 1 | 12.16 | C26H39O17 | 623.2182 | 623.2173 | −1.44 | 477(100), 461(77.4), 459(22.0) | 315(100), 221(12.6) | Kankanoside F | c | c | |
| 2b | 12.45 | C20H29O12 | 461.1654 | 461.1657 | 0.65 | 315(100), 297(10.9), 161(11.8) | 135(100), 113(3.6) | Decaffeoylacteoside isomer | c | c | c |
| 3 | 13.79 | C14H19O7 | 299.1125 | 299.1130 | 1.67 | 119(100), 137(91.2) | — | Salidroside | c | c | c |
| 4 | 14.81 | C20H29O12 | 461.1653 | 461.1664 | 2.39 | 315(100), 135(74.9), 161(28.8) | 135(100) | Decaffeoylacteoside | c | ||
| 5b | 15.71 | C20H29O11 | 445.1704 | 445.1710 | 1.35 | 299(100), 161(27.7) | 179(100), 143(97.0), 113(81.0), 161(41.1), 119(80.7) | Cistanoside G isomer | c | c | c |
| 6 | 16.74 | C20H29O11 | 445.1704 | 445.1711 | 1.57 | 299(100), 161(22.2), 205(19.1) | 179(100), 161(66.2), 143(33.5) | Cistanoside G | c | ||
| 7b | 17.25 | C21H31O12 | 475.1810 | 475.1815 | 1.05 | 329(100), 161(32.6), 143(10.5) | 161(100), 179(86.2), 149(77.1) | Cistanoside E isomer | c | ||
| 8b | 18.76 | C28H41O18 | 665.2287 | 665.2285 | −0.30 | 623(100), 605(8.6), 461(1.9) | 477(100), 461(96.0), 459(18.1), 315(13.1), 179(1.0) | Decaffeoyltubuloside A | c | ||
| 9 | 19.57 | C21H31O12 | 475.1810 | 475.1821 | 2.31 | 329(100), 311(22.9), 161(34.3) | 161(100), 179(85.4), 149(78.3) | Cistanoside E | c | ||
| 10b | 19.91 | C22H31O13 | 503.1759 | 503.1768 | 1.79 | 461(100), 443(3.5) | 315(100), 135(35.0), 297(12.0) | Cistanoside H isomer | c | c | |
| 11 | 20.13 | C16H21O8 | 341.1231 | 341.1235 | 1.17 | 281(100), 179(24.7), 161(14.0), 119(12.1) | — | 6′-Acetylsalidroside | c | ||
| 12 | 21.06 | C35H45O21 | 801.2448 | 801.2442 | −0.75 | 783(100), 621(29.0), 639(11.3) | 621(100) | Cistantubuloside C1/C2 | c | c | |
| 13 | 22.17 | C22H31O13 | 503.1759 | 503.1768 | 1.79 | 443(100), 461(69.0) | 261(100), 297(77.5), 215(25.0), 177(25.0) | Cistanoside H | c | c | |
| 14 | 23.24 | C29H33O15 | 621.1814 | 621.1822 | 1.29 | 475(100), 295(6.0) | 295(100), 269(76.9), 267(55.3) | Crenatoside | c | c | c |
| 15 | 23.85 | C36H47O21 | 815.2604 | 815.2587 | −2.09 | 653(100), 797(27.6), 635(10.3) | 491(100), 507(34.6), 329(10.3) | Kankanosides K1/K2 | c | c | c |
| 16a | 24.38 | C35H45O20 | 785.2499 | 785.2489 | −1.27 | 623(100), 477(1.0) | 477(100), 461(87.9), 459(22.1) | Echinacoside | c | c | c |
| 17b | 25.25 | C16H21O8 | 341.1231 | 341.1233 | 0.59 | 281(100), 119(11.5), 179(9.7), | 164(100) | 4′-Acetylsalidroside | c | ||
| 18 | 28.06 | C35H45O19 | 769.2550 | 769.2543 | −0.91 | 607(100), 461(13.9), 445(11.2) | 445(100), 461(81.1), 443(31.2) | Cistantubuloside A | c | c | |
| 19a | 29.25 | C36H47O20 | 799.2655 | 799.2647 | −1.00 | 637(100), 623(18.9), 491(10.2) | 491(100), 475(48.3), 473(34.7) | Cistanoside A | c | c | |
| 20 | 29.43 | C35H45O19 | 769.2550 | 769.2543 | −0.91 | 623(100), 605(6.8) | 477(100), 461(92.2), 459(20.9) | Cistantubuloside B1/B2 | c | c | |
| 21 | 32.02 | C34H43O19 | 755.2393 | 755.2384 | −1.19 | 593(100), 623(2.3) | 447(100), 461(98.4), 429(23.3) | Arenarioside | c | c | c |
| 22 | 32.22 | C37H47O21 | 827.2604 | 827.2592 | −1.45 | 665(100), 623(63.2), 785(8.2) | 623(100), 605(69.4), 519(10.7), 503(7.8) | Tubuloside A | c | c | c |
| 23a | 32.88 | C29H35O15 | 623.1970 | 623.1960 | −1.60 | 461(100), 315(41.9), 443(10.6), 477(8.9) | 315(100), 297(26.1), 135(17.0) | Acteoside | c | c | c |
| 24 | 33.97 | C43H53O23 | 937.2972 | 937.2973 | 0.11 | 811(100), 607(43.1) | — | Tubuloside D | c | ||
| 25 | 34.32 | C29H35O15 | 623.1970 | 623.1969 | −0.16 | 461(100), 443(12.6), 477(6.9) | 315(100), 135(77.0), 205(36.7) | Cisacteoside | c | ||
| 26b | 35.06 | C43H53O23 | 937.2972 | 937.2968 | −0.43 | 811(100), 607(77.9) | — | Cistubuloside D | c | ||
| 27 | 36.32 | C36H47O20 | 799.2655 | 799.2651 | −0.50 | 637(100), 653(16.0), 623(4.9) | 461(100) | Wiedemanninoside C | c | ||
| 28 | 37.67 | C37H49O20 | 813.2811 | 813.2802 | −1.11 | 619(100), 473(19.1), 491(13.7) | 473(100), 457(38.8), 443(16.8) | Cistanoside B | c | ||
| 29 | 41.04 | C30H37O16 | 653.2076 | 653.2073 | −0.46 | 491(100), 476(4.1) | 476(100), 345(38.9), 150(10.0) | Campneoside I | c | ||
| 30a | 40.81 | C35H45O19 | 769.2550 | 769.2543 | −0.91 | 607(100), 623(3.4) | 461(100), 443(14.5) | Poliumoside | c | c | c |
| 31b | 40.41 | C29H33O15 | 621.1813 | 621.1822 | 1.45 | 459(100), 179(19.2), 487(15.7) | 151(100), 277(39.8) | Crenatoside isomer | c | c | c |
| 32b | 40.69 | C38H49O21 | 841.2761 | 841.2757 | −0.48 | 665(100), 623(33.5), 799(34.5) | 623(100), 605(8.4) | Cistanoside N isomer | c | c | |
| 33 | 41.93 | C30H37O16 | 653.2076 | 653.2075 | −0.15 | 491(100), 638(9.3), 507(6.1) | 476(100), 345(38.9) | Isocampneoside I | c | ||
| 34 | 41.97 | C29H35O14 | 607.2021 | 607.2017 | −0.66 | 445(100), 461(11.2), 427(11.2) | 299(100), 161(15.0), 179(11.2) | Kankanoside G | c | c | c |
| 35 | 42.87 | C37H47O20 | 811.2655 | 811.2654 | −0.12 | 665(100), 649(27.1), 769(12.6) | 623(100), 605(7.8) | Kankanoside H1/H2 | c | ||
| 36a | 42.62 | C29H34O15 | 623.1970 | 623.1968 | −0.32 | 461(100), 315(11.3), 477(7.4) | 315(100), 297(42.6), 135(16.4) | Isoacteoside | c | c | c |
| 37b | 43.76 | C38H49O21 | 841.2761 | 841.2757 | −0.48 | 799(100), 637(49.8), 619(19.0) | — | Cistanoside N isomer | c | ||
| 38a | 43.86 | C30H37O15 | 637.2127 | 637.2117 | −1.57 | 475 (100), 461(45.9), 491(14.8), 457(12.2) | 329(100), 161(30.2), 315(12.5) | Cistanoside C | c | c | c |
| 39b | 44.06 | C29H35O14 | 607.2021 | 607.2017 | −0.66 | 461(100), 443(1.4), 445(100), 461(11.2), 427(11.2) | 315(100), 297(17.0) | cis-Kankanoside G | c | c | c |
| 40a | 47.04 | C31H37O16 | 665.2076 | 665.2072 | −0.60 | 461(100), 503(38.9), 443(5.7) | 315(100), 297(19.6) | 2′-O-Acetylacteoside | c | c | c |
| 41 | 47.58 | C35H45O18 | 753.2600 | 753.2593 | −0.93 | 591(100), 607(11.3) | 445(100), 427(34.2) | Kankanoside I | c | c | |
| 42 | 49.45 | C30H37O15 | 637.2127 | 637.2129 | 0.31 | 475(100), 461(31.1), 457(15.7) | 329(100), 161(35.3) | Jionoside D | c | ||
| 43 | 50.36 | C38H49O21 | 841.2761 | 841.2754 | −0.83 | 695(100), 653(39.9), 799(23.8) | — | Cistanoside N | c | ||
| 44 | 52.34 | C29H35O14 | 607.2021 | 607.2017 | −0.66 | 461(100), 445(49.1) | 315(100), 135(59.2), 161(18.9) | Isosyringalide-3′-α-L-rhamnose | c | c | c |
| 45b | 52.78 | C35H45O18 | 753.2593 | 753.2606 | 1.73 | 607(100), 589(4.5) | 461(100), 443(15.2) | Ciskankanoside I | c | ||
| 46b | 53.77 | C31H37O16 | 665.2076 | 665.2078 | 0.30 | 503(100), 605(48.6), 443(39.1), 623(34.0), 461(23.0), 519(4.0) | 443(100), 461(70.2), 357(36.7) | Cistubuloside B | c | ||
| 47 | 53.52 | C29H35O13 | 591.2072 | 591.2068 | −0.68 | 445(100), 427(28.6) | 299(100), 145(24.7), 163(23.5), 161(22.9) | Osmanthuside B | c | c | c |
| 48a | 53.75 | C30H37O15 | 637.2127 | 637.2117 | −1.57 | 461(100), 491(36.9), 475(17.3) | 315(100), 135(67.6), 297(17.6) | Isocistanoside C | c | c | |
| 49b | 53.83 | C30H37O14 | 621.2178 | 621.218 | 0.32 | 445(100), 427(44.7), 175(40.7) | — | Cistanoside M ismor | c | c | c |
| 50 | 54.18 | C29H35O14 | 607.2021 | 607.2025 | 0.66 | 461(100), 443(2.39) | 315(100), 135(46.5), 161(16.0) | Syringalide A-3′-α-L-rhamnose | c | c | c |
| 51 | 56.04 | C32H39O17 | 695.2182 | 695.2174 | −1.15 | 491(100), 653(34.3), 533.2(23.4) | 345(100) | Kankanosides J1/J2 | c | c | c |
| 52 | 55.35 | C30H37O14 | 621.2178 | 621.218 | 0.32 | 475(100), 457(42.8), 443(11.6) | 329(100), 161(45.3) | Cistanoside M | c | ||
| 53 | 55.55 | C31H39O15 | 651.2283 | 651.2282 | −0.15 | 475(100), 193(24.3), 457(22.7) | 329(100), 161(62.6), 311(17.6) | Cistanoside D | c | ||
| 54a | 56.14 | C37H47O20 | 811.2655 | 811.2655 | 0 | 607(100), 649(66.4), 769(22.8) | 461(100), 443(18.5) | 2′-O-Acetylpoliumoside | c | c | c |
| 55b | 58.99 | C32H39O16 | 679.2232 | 679.2227 | −0.74 | 637(100), 475(16.6), 461(8.3) | 461(100), 491(11.6), 475(6.3) | Isocistansinenside A | c | c | |
| 56 | 58.71 | C30H37O15 | 637.2126 | 637.2129 | 0.47 | 475(100), 461(88.3), 491(74.7) | 329(100), 161(37.2) | Plantainoside C | c | ||
| 57 | 58.61 | C31H37O15 | 649.2127 | 649.2131 | 0.62 | 607(100), 503(39.4), 461(356.6) | 461(100), 443(2.6) | Salsaside D | c | c | c |
| 58a | 58.73 | C31H37O16 | 665.2076 | 665.2072 | −0.60 | 461(100), 503(53.9), 623(47.2) | 315(100), 135(46.1), 297(19.3), 161(14.3) | Tubuloside B | c | c | c |
| 59 | 60.80 | C31H39O15 | 651.2283 | 651.2289 | 0.92 | 475(100), 457(49.6), 505(31.3) | 329(100), 161(45.3), 311(10.3) | Epimeridinoside A | c | ||
| 60 | 62.81 | C32H39O16 | 679.2232 | 679.2239 | 1.03 | 637(100), 475(25.7) | 475(100), 461(58.3), 491(36.6) | Salsaside E | c | ||
| 61 | 66.07 | C31H39O15 | 651.2283 | 651.2282 | −0.15 | 505(100), 475(52.4), 487(27.8) | 161(100), 193(48.1), 297(37.3) | Isomartynoside | c | ||
| 62 | 66.38 | C29H35O13 | 591.2072 | 591.2068 | −0.68 | 445(100), 427(13.9) | 145(100), 299(47.8), 163(42.4), 265(30.6) | Osmanthuside B6 (Z/E) | c | c | c |
| 63 | 68.55 | C33H41O16 | 693.2389 | 693.2384 | −0.72 | 651(100), 633(21.8), 505(14.7) | 505(100), 475(46.5), 487(27.1) | Cistanoside J | c | ||
| 64 | 69.12 | C31H37O15 | 649.2126 | 649.2131 | 0.77 | 607(100), 461(36.4), 503(34.5) | 461(100), 443(2.7) | Salsaside F | c | c | |
| 65 | 69.16 | C38H49O20 | 825.2811 | 825.2817 | 0.73 | 783(100), 621(56.2), 603(27.0), 663(22.6) | 637(100) | Cistansinenside B | c | ||
| 66b | 70.28 | C32H41O16 | 679.2232 | 679.2227 | −0.74 | 637(100), 503(14.5), 461(11.0) | 461(100), 491(8.4) | Isocistanoside K | c | c | |
| 67b | 71.71 | C33H41O16 | 693.2389 | 693.2396 | 1.01 | 651(100), 633(21.1) | 475(100), 505(94.3), 193(29.5) | Isocistanoside J | c | ||
| 68 | 71.26 | C32H39O16 | 679.2232 | 679.2227 | −0.74 | 637(100), 461(9.6), 619(8.7) | 461(100), 491(39.9), 475(11.2) | Cistanoside K | c | ||
| 69b | 73.65 | C32H39O16 | 679.2232 | 679.2239 | 1.03 | 637(100), 475(8.3), 619(8.3), 461(4.7) | 461(100), 491(51.0), 475(46.8) | Ciscistanoside K | c | c | |
| 47 | 42 | 42 | |||||||||
3.6. Multivariate data analysis with PCA and OPLS-DA methods
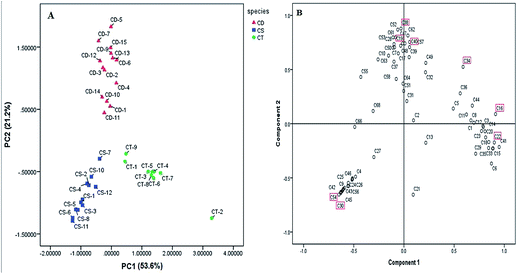
The repeatability and precision of ten obvious peaks in three samples was measured and analyzed. The RSD values calculated using Microsoft Excel 2011 were all less than 7.0%, indicating the conditions for the targeted profiling analysis were reliable and applicable.In order to determine the chemical differences among three different plant origins collected from various resources, an approach of chromatographic profiling for the characterized 69 PhGs in combination with PCA multivariate data analysis approach was exploited. Calculation was performed with the standard-projection plot of 1–2 principal component sized correlation matrix. The principal factorial plane summarized 74.80% of the whole variability, and two PCA axes explained 53.64% and 21.26% of the variance, respectively. As shown in the distribution plot, 36 batches of Herba Cistanches could be explicitly clustered into three groups (Fig. 5A). Interestingly, these three groups were consistent with the three species very well, which demonstrated that chemical differences among them are great to some extent. The score plot of CT-2 was far away from the cluster of C. tubulosa owing to the extremely high contents of echinacoside (16), acteoside (23), and isoacteoside (36). At the same time, the clustering of C. deserticola was not very tight as several individual samples were sparsely distributed, indicating that the quality of C. deserticola is not well consistent and easily affected by the exterior factors, for example, the habitats, the hosts, harvested time, and processing methods, etc. In addition, according to the correlation plot (Fig. 5B), we found that eight PhGs contributing to the data sets separation including echinacoside (16), cistanoside A (19), tubuloside A (22), poliumoside (30), isosyringalide-3′-α-L-rhamnose (34), cistanoside C (38), 2′-O-acetylacteoside (40), and 2′-O-acetylpoliumoside (54), were much more statistically significant in chemotaxonomy than the other PhGs identified. And these eight common compounds might be the most important chemical markers for discrimination of the internal quality of Cistanches Herba samples, indicating that their contents could be regarded as the criterion to evaluate the quality of Cistanches samples from different sources.
Moreover, a supervised OPLS-DA discriminant model based on the eight potential PhGs markers was constructed to validate their reliability markers and determine which class a new sample belongs to. This is simply performed by projecting a new sample onto the eigenvectors space and by selecting the nearest class using OPLS-DA, a statistical method for supervised classification of data to provide good or bad, qualified or unqualified results. The method requires a training data set consisting of samples with a set of attributes and their class memberships. Training samples of the class are in the acceptable region after being used to build a model, and other samples are located outside the acceptable region. A test sample is projected onto the model to validate the model. If the sample is in the acceptable region, it belongs to the class. If not, the sample is unknown.
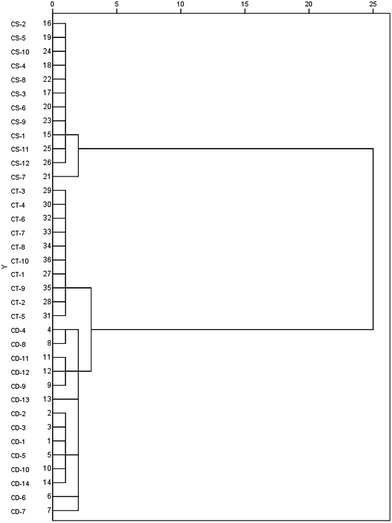
As a result, the OPLS-DA model constructed in this experiment resulted in (2 + 3 + 0) component with Q2 of 87.35% and R2(Y) of 92.24%, which demonstrated good quality of the model. Three sets of different samples were better separated in the eigenvectors space. In order to further validate the discriminatory efficiency, test samples including 3 C. deserticola, 2 C. tubulosa, and 2 C. sinensis remained at random were projected onto the built model using OPLS-DA method. Fig. 6A demonstrated that the model's checking ability was high. The remaining validation C. deserticola, C. tubulosa, and C. sinensis samples were correctly projected onto their respective groups. The recognition values and validated results are also illustrated in Table 3, indicating that these eight discriminatory PhGs were the most significant markers and had more influence on the sample discrimination than the other PhGs. Moreover, the results of HCA analysis based on 8 investigated markers demonstrated that the chemical variation is obvious among the three different species of Cistanches Herba (shown in Fig. 7), which further confirmed the deduced results from PCA and OPLS-DA analysis. In the primary work, the anti-inflammatory and neuroprotective activities of PhGs have been intensively investigated. For example, PhGs from the stems of C. deserticola indicated NO radical-scavenging activity, which possibly contributed to their anti-inflammatory effects;34 tubuloside B and echinacoside showed neuroprotective effect on tumor necrosis factor-alpha (TNF-α)-induced apoptosis in human neuroblastoma (SHSY5Y) cells,35,36 which were be closely associated with inflammation related neuronal degenerative diseases including Alzheimer's disease (AD) and Parkinson's diseases (PD). Therefore, these eight PhGs could be regarded as the indicators to perform the quality control of the three different Cistanches species.
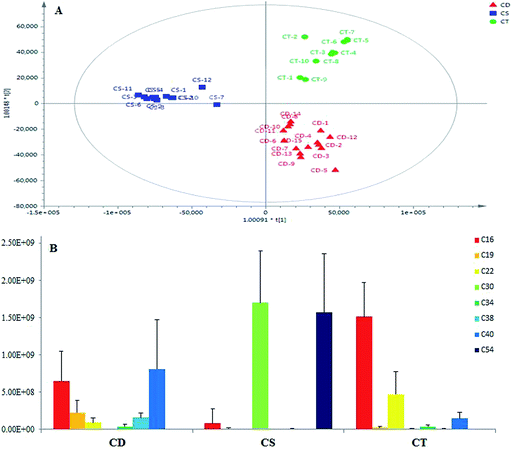 | ||
| Fig. 6 The OPLS-DA score plot obtained from the groups of CD, CT, and CS (A) and column plot of variable average intensity of the eight discriminatory PhGs markers (B). | ||
| Samples | Recognition value CD | Recognition value CT | Recognition value CS |
|---|---|---|---|
| CD-6 | 1.0089 | −0.1005 | 0.0916 |
| CD-10 | 0.7158 | −0.8682 | 0.1974 |
| CD-14 | 1.0351 | −0.0373 | 0.0023 |
| CT-1 | 0.1160 | 0.7510 | 0.0427 |
| CT-9 | 0.2059 | 0.7513 | 0.1330 |
| CS-8 | −0.0009 | −0.0029 | 1.0038 |
| CS-12 | −0.0849 | 0.0185 | 1.0663 |
The above studies compared the chemical constituents at a global level rather than based on a few markers, and thus led to more reliable results. This is the first systematic chemical analysis for PhGs in three main species of Cistanches plants. Importantly, the chemical difference among species was comprehensively elucidated for the first time. Moreover, each level of this method, i.e. HPLC analysis, LC-MS coupled with PCA analysis, and LC-MS coupled with OPLS-DA multivariate statistical analysis, could be used for quality control and species authentication.
4. Conclusions
This study developed and validated an HPLC-LTQ-Orbitrap-based strategy for TCM targeted class components discovery, identification, and herbomics research using phenylethanoid glycosides (PhGs) in Cistanche deserticola, C. sinensis, and C. tubulosa as an example. From these three different plant origins, a total of 69 PhGs were rapidly discovered and characterized by HPLC-ESI-LTQ-Orbitrap using multiple data-mining methods such as accurate parent mass search, parent mass modification search and fragment ion search. Among them, 10 PhGs were unambiguously confirmed, while the others were tentatively identified to be 42 known and 17 new PhGs. This is the first report on comprehensive chemical analysis of PhGs in Cistanche species. Based on the information of 69 PhGs identified, multivariate statistical analysis of PCA and OPLS-DA were applicated in herbomics research to conform that eight PhGs were mainly responsible for the chemical discrimination among three different plant origins. The result suggested that these eight PhGs were likely to be the chemical markers leading to their differences. In conclusion, this new developed strategy could be targets for future studies on the discovery and identification of important chemical constituents and classification of genuine specie or genus from other adulterants. Meanwhile, it may propose a novel idea for specific analysis of active chemical constituents in the same type, and is promising for quality control and evaluation of TCMs.Acknowledgements
The authors greatly appreciate the financial support from China Postdoctoral Science Foundation (No. 2013M530563) and National Foundation of Natural Sciences of China (No. 30902001).References
- H. Kobayashi, H. Oguchi, N. Takizawa, T. Miyase, A. Ueno, K. Usmanghani and M. Ahmad, Chem. Pharm. Bull., 1987, 35, 3309 CrossRef CAS.
- P. F. Tu, Z. C. Lou, S. C. Li, C. S. Wang, W. Y. Jiang and Y. M. Wang, J. Chin. Med. Mater., 1996, 19, 420 Search PubMed.
- Y. Zhang, H. Wu, S. N. Wang and H. C. Zheng, China J. Chin. Mater. Med., 1994, 19, 169 CAS.
- L. Lei, Z. H. Song and P. F. Tu, Chin. Tradit. Herb. Drugs, 2003, 34, 473 CAS.
- P. Yamada, R. Iijima, J. Han, H. Shigemori, S. Yokota and I. Hiroko, Planta Med., 2010, 14, 1512 CrossRef PubMed.
- T. Morikawa, Y. Pan, K. Ninomiya, K. Imura, H. Matsuda, M. Yoshikawa, D. Yuan and O. Muraoka, Bioorg. Med. Chem., 2010, 18, 1882 CrossRef CAS PubMed.
- Y. P. He, Z. Z. Yin, P. F. Tu and Z. C. Lou, J. Chin. Med. Mater., 1997, 20, 117 CAS.
- S. Y. Zhu, J. Chin. Med. Mater., 2001, 24, 24 CAS.
- X. Q. Xu, H. Ni, H. Y. Wei, X. G. Jia, B. G. Zhang and D. G. Qing, Lishizhen Med. Mater. Med. Res., 2013, 24, 881 Search PubMed.
- M. Zhang, X. L. Fu and S. Y. Wang, J. Chin. Med. Mater., 2004, 27, 400 Search PubMed.
- R. Xu, S. Q. Sun, Y. G. Liu, J. Chen, T. N. Chen and S. L. Liu, Spectrosc. Spectral Anal., 2010, 30, 897 CAS.
- J. Chen, S. Q. Sun, R. Xu, Y. G. Liu, J. Yu, T. N. Liu and J. Q. Li, Spectrosc. Spectral Anal., 2009, 29, 1502 CAS.
- J. P. Han, J. Y. Song, C. Liu, J. Chen, J. Qian, Y. J. Zhu, L. C. Shi, H. Yao and S. L. Chen, Acta Pharmaceutica Sinica B, 2010, 45, 126 CAS.
- H. M. Shi, J. Wang, M. Y. Wang, P. F. Tu and X. B. Li, Biol. Pharm. Bull., 2009, 32, 142 CAS.
- Y. Jiang, S. P. Li, Y. T. Wang, X. J. Chen and P. F. Tu, J. Chromatogr. A, 2009, 1216, 2156 CrossRef CAS PubMed.
- L. F. Han, M. B. Yiadom, E. W. Liu, Y. Zhang, W. Li, X. B. Song, F. H. Fu and X. M. Gao, Phytochem. Anal., 2012, 23, 668 CrossRef CAS PubMed.
- S. H Zheng, X. Jiang, L. B. Wu, Z. H. Wang and L. F. Huang, PLoS One, 2014, 9, e98061 Search PubMed.
- D. Y. Lu, J. Y. Zhang, Z. Y. Yang, H. M. Liu, S. Li, B. J. Wu and Z. G. Ma, J. Sep. Sci., 2013, 36, 1945 CrossRef CAS PubMed.
- Z. D. Nan, K. W. Zeng, S. P. Shi, M. B. Zhao, Y. Jiang and P. F. Tu, Fitoterapia, 2013, 89, 1674 CrossRef PubMed.
- M. S. Zhu, L. Ma, H. Y. Zhang and W. G. Humphreys, Anal. Chem., 2007, 79, 8333 CrossRef CAS PubMed.
- W. Xu, J. Zhang, Z. H. Huang and X. H. Qiu, Anal. Methods, 2012, 4, 1806 RSC.
- J. Y. Zhang, F. Wang, H. Zhang, J. Q. Lu and Y. Q. Qiao, Phytochem. Anal., 2014, 25, 405 CrossRef PubMed.
- A. Makarov, E. Denisov, O. Lange and S. Horning, J. Am. Soc. Mass Spectrom., 2006, 17, 977 CrossRef CAS PubMed.
- P. F. Tu, H. M. Shi, Z. H. Song, Y. Jiang and Y. Y. Zhao, J. Asian Nat. Prod. Res., 2007, 9, 79 CrossRef CAS PubMed.
- M. F. Lahloub, A. M. Zaghloul, S. A. El-Khayaat, M. S. Afifi and O. Sticher, Planta Med., 1991, 57, 481 CrossRef CAS PubMed.
- G. H. Lu, K. Chan, Y. Z. Liang, K. Leung, C. L. Chan, Z. H. Jiang and Z. Z. Zhao, J. Chromatogr. A, 2005, 1073, 383 CrossRef CAS PubMed.
- F. Gong, Y. Z. Liang, P. S. Xie and F. Chau, J. Chromatogr. A, 2003, 1002, 25 CrossRef CAS.
- J. Y. Zhang, Z. J. Wang, Q. Zhang, F. Wang, Q. Ma, Z. Z. Lin, J. Q. Lu and Y. J. Qiao, Talanta, 2014, 124, 111 CrossRef CAS PubMed.
- J. Y. Zhang, Q. Zhang, N. Li, Z. J. Wang, J. Q. Lu and Y. J. Qiao, Talanta, 2013, 104, 1 CrossRef CAS PubMed.
- M. Qi, A. Z. Xiong, F. Geng, L. Yang and Z. T. Wang, J. Sep. Sci., 2012, 35, 1470 CrossRef CAS PubMed.
- M. Y. Liu, S. H. Zhao, Z. Q. Wang, Y. F. Wang, T. Liu, S. Li, C. C. Wang, H. T. Wang and P. F. Tu, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2014, 115, 949 Search PubMed.
- J. Y. Zhang, W. Cai, Y. Zhou, Y. Liu, X. D. Wu, Y. Li, J. Q. Lu and Y. J. Qiao, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2015, 985, 91 CrossRef CAS PubMed.
- Y. Z. Zhang, F. Xu, J. Dong, J. Liang, Y. Hashi, M. Y. Shang, D. H. Yang, X. Wang and S. Q. Cai, J. Pharm. Biomed. Anal., 2012, 70, 425 CrossRef CAS PubMed.
- Q. Xiong, Y. Tezuka, T. Kaneko, H. Li, L. Q. Tran, K. Hase, T. Namba and S. Kadota, Eur. J. Pharmacol., 2000, 400, 137 CrossRef CAS.
- M. Deng, J. Y. Zhao, P. F. Tu, Y. Jiang, Z. B. Li and Y. H. Wang, Eur. J. Pharmacol., 2004, 505, 11 CrossRef CAS PubMed.
- M. Deng, J. Y. Zhao, X. D. Ju, P. F. Tu, Y. Jiang and Z. B. Li, Acta Pharmacol. Sin., 2004, 25, 1276 CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra13276b |
| This journal is © The Royal Society of Chemistry 2015 |