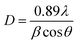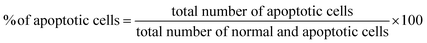Fabrication of a pH responsive DOX conjugated PEGylated palladium nanoparticle mediated drug delivery system: an in vitro and in vivo evaluation†
Krishnamurthy Shanthia,
Karuppaiya Vimalaa,
Dhanaraj Gopic and
Soundarapandian Kannan*b
aDepartment of Zoology, Bharathiar University, Coimbatore-641 046, Tamil Nadu, India
bProteomics and Molecular Cell Physiology Laboratory, Department of Zoology, Periyar University, Salem-636 011, Tamil Nadu, India. E-mail: skperiyaruniv@gmail.com; sk_protein@buc.edu.in; Fax: +91 2345124; Tel: +91 427 2345766 ext. 335
cDepartment of Chemistry, Periyar University, Salem-636 011, Tamil Nadu, India
First published on 12th May 2015
Abstract
Efficient delivery of therapeutics into tumor cells to increase the intracellular drug concentration is one of the key issues in cancer therapy. In this work, we designed pH responsive PEGylated palladium nanoparticles (PdNPs) as an anticancer drug nanocarrier system for effective drug delivery. The synthesis of the nanocarrier involved conjugation of doxorubicin (DOX) to the surface of PEGylated PdNPs via a hydrazone interaction. The nanoparticles were characterized by UV-spectroscopy, Transmission Electron Microscopy (TEM), Dynamic Light Scattering (DLS), zeta potential, Fourier transmission Infrared spectroscopy (FT-IR), X-ray diffraction (XRD) and Nuclear Magnetic Resonance (NMR). The drug release behavior was subsequently studied at different pH conditions. The results showed a sustained release of DOX preferentially at the desired endosomal pH (5.5). The biological activity of the DOX conjugated PEGylated PdNPs was studied by an MTT assay, fluorescence microscopy, and apoptosis. Intracellular-uptake studies revealed preferential uptake of these NPs into HeLa cancer cells. The in vitro apoptosis study revealed that the DOX conjugated PEGylated PdNPs caused significant death to the HeLa cells. Further, blank PEGylated PdNPs displayed low toxicity and good biocompatibility. DOX conjugated PEGylated PdNPs had the strongest anti-tumor efficacy against HeLa tumor xenograft models in vivo. These findings demonstrated that PEGylated PdNPs were deemed as a potential drug nanocarrier for cancer therapy.
1. Introduction
Over the past few decades, the growth of nanotechnology has developed to such an extent that it is feasible to synthesize, characterize and modify the functional properties of nanoparticles for biomedical applications.1 This has led to an enormous interest in the wide-scale production of nanoparticles (for example quantum dots, and metallic, magnetic, and polymeric nanoparticles) with different shapes and sizes for diverse biomedical applications in drug delivery, disease diagnostics, and medical imaging.2 The limitations of conventional chemotherapy include general systemic distribution of the drug, lack of drug specificity to the tumor site, insufficient local drug concentration in the tumor and poor control over drug release. The general systemic distribution of chemotherapeutic agents results in toxic side effects since the drug attacks normal, healthy cells together with the tumor cells.3 Therefore, it is very crucial to selectively target chemotherapeutic agents to the tumor. This need has aggravated a search for methods of drug delivery which can tackle this limitation and provide more effective cancer therapy. Delivery of chemotherapeutic agents within or conjugated to nanoparticles is a promising alternative to evade the problems associated with conventional chemotherapy.4 In many cases, these nanoparticles often require surface modification to guarantee their biocompatibility and/or improve the bioavailability. Generally metal nanoparticles are coated with surfactants5,6 or polymers7 to provide stabilization of nanoparticles in biological suspension, functionalization at the surface with drugs and to increase circulation time by reducing immediate clearance of the carriers by reticuloendothelial system (RES). Nanoparticle-based drug delivery systems have shown a high degree of efficacy in cancer treatments due to their improved pharmacokinetics and biodistribution profiles by means of the enhanced permeability and retention (EPR) effect.8 Although the EPR is effective in enhancing the accumulation of nanoparticles (NPs) within tumor tissues, the poor cellular internalization and insufficient drug release limits the dosages of anticancer drugs to levels below the optimum therapeutic value, thereby adversely affecting the efficacy of the chemotherapy treatment of cancer.9 In order to address these issues, stimuli-responsive delivery systems have been explored to improve bioavailability of a drug.10 As well, among the stimuli, pH-responsiveness is the most frequently investigated since pH values vary quite significantly in different tissues and cellular compartments.11–13 The extracellular environment of a tumor has a lower pH (∼6.8) than blood and normal tissues (pH 7.4)14–17 whereas those of late endosome and lysosome are even lower (∼5.0–5.5).18 Thus pH-sensitive delivery systems are of particular interest in controlled drug-delivery as evident from the literature.19 In this context, we aimed to design a potentially capable pH-responsive nanoparticle drug delivery system specifically to cancer cells.Among the metal nanoparticles, PdNPs belonging to the platinum group metals have been widely investigated because they exhibit unusual optical, electronic, and chemical properties, depending on their size and shape, thus opening many possibilities with respect to technological applications. The high surface to volume ratio of nanomaterials makes them highly efficient as potential catalysts. Surface plasma resonance (SPR) is another important feature in palladium nanoparticles which is useful in sensing, chemo-optical transducers, plasmonic wave guiding.20–22 Owing to these unique properties, Pd has been used as a catalyst to manufacture pharmaceuticals,23 degrade harmful environmental pollutants,24 and as sensors for the detection of various analytes.25–27 Additionally, Pd and Pd2+ ions also play a fundamental role in several biotechnological processes. More recently, PdNPs have been found to have antimicrobial properties.28 Notably, PdNPs show very broad absorption through the UV-vis-NIR region. This broad absorption nature has attracted extensive interest in using PdNPs in photothermal therapy. Xiao et al. prepared PdNPs with porous structure, which exhibit superior performance in photothermal therapy compared to solid Pd nanocubes in HeLa cells.29 In addition, a few literatures authenticated the anticancer activity of Pd complexes along with Pt(II) complex.30,31 Furthermore, the anti-invasive property of Pd complex was also witnessed in earlier studies.32–34 While the advantages and uses of Pd are extensive, advances are yet to be uncovered as the metals are reduced to the nanoscale. Moreover, despite the myriad applications of PdNPs in various fields, relatively no studies have been conducted to determine the therapeutic applications of PdNPs. Thus for PdNPs, there is much room for exploring their potential properties in the fields of drug delivery and it is also important to determine a baseline of toxicity for PdNPs as well as examine their potential for therapeutic applications.
Herein we introduce a novel PdNPs hybrid system to investigate their biological responses against cervical cancer. The critical part of the hybrid system is the functionalization of PdNPs surface properties by poly(ethylene)glycol (PEG)-hydrazide polymers and conjugating DOX onto the PEG-hydrazide (PEGylated) PdNPs. Hydrophilic polymer PEG was chosen as surface modifiers of PdNPs because of its biocompatibility.35 PEG coated nanoparticle surfaces when dispersed reduce nonspecific protein adsorption and clearance by macrophages, and render the nanoparticles capable of crossing the cell membrane.36,37 DOX was chosen as a model anti-cancer agent owing to its high therapeutic index and better activity against a wide spectrum of tumors.38 In the present study, our goal is to conjugate DOX with PEGylated PdNPs nanocarriers and study the drug loading and controlled release profile from these nanocarriers. Additionally, the targeting activity of the nanoparticles in HeLa cancer cells was evaluated by cellular uptake and cytotoxicity in vitro. The targeting characteristics of DOX conjugated PEGylated PdNPs were further investigated by anti-tumor efficacy studies in vivo. The process involved in the fabrication of PEGylated PdNPs, can be used to conjugate with DOX via hydrazone bond. When the as-fabricated nanoparticles targeted the HeLa cells, the nanoparticles enter into the cytoplasm mediated by endocytosis effects. DOX is then released from nanoparticles at low pH which degrades polymer and resulting in inhibition of tumor cell growth through apoptosis.
2. Experimental
2.1. Materials
The human cervical cancer cell line (HeLa) was procured from National Center for Cell Science (NCCS, Pune), palladium(II) chloride (PdCl2) as the source of palladium ions, methoxy poly(ethylene glycol)-succinimidyl (5k) (PEG-SCM), triethylamine (TEA), 3-(4,5-dimethylthiazolyl-2)-2,5-diphenyltetrazolium bromide (MTT), 4′-6-diamidino-2-phenylindole (DAPI), acridine orange/ethidium bromide (Ao/EtBr), tetrahydrofuran (THF), sodium borohydrate (NaBH4), sodium dodecyl sulphate (SDS) and hydrazine was purchased from Sigma-Aldrich (Bangalore). Analytical grade reagents were purchased from Sigma-Aldrich (Bangalore). All the samples were prepared in Milli-Q water.2.2. Synthesis of DOX conjugated PEGylated PdNPs via hydrazone bonds
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 10 min, washed several times with water, and then dried at 60 °C for 5 h in a vacuum dryer or re-dispersed in water to produce a colloidal suspension for further characterizations.
000 rpm for 10 min, washed several times with water, and then dried at 60 °C for 5 h in a vacuum dryer or re-dispersed in water to produce a colloidal suspension for further characterizations.| DLC (wt%) = (weight of loaded drug/weight of drug − loaded carrier) × 100% |
2.3. Characterization of nanoparticles
The average grain size of the PEGylated PdNPs and the DOX conjugated PEGylated PdNPs is calculated using Scherrer formula:
2.4. In vitro drug release response
The drug release response from polymer modified nanoparticles was investigated at the physiological temperature of 37 °C and pH of 7.4, 6.8 and 5.5. The medium of pH 7.4 corresponds the physiological pH while pH 6.8 simulates the pH of tumor tissue and pH of 5.5 corresponds to the mature endosomes of tumor cells. The release profiles of DOX from PEGylated PdNPs were studied using a dialysis bag (MWCO-3500). Briefly DOX conjugated PEGylated PdNPs was dispersed in 10 mL of the respective PBS buffer, allowed to stabilize for 30 min, and then placed in a dialysis bag. The dialysis bag was immersed in 50 mL of PBS solution (pH 7.4, 6.8, or 5.5) in a beaker and then placed in a 37 °C water bath shaker at 110 rpm. At predetermined time intervals, 3 mL of the release medium was collected to measure the released drug concentration and then was replaced with the same fresh PBS. For the measurement of released DOX concentration, the absorbance of the release medium at 480 nm was recorded on a Shimadzu UV-vis absorption spectrophotometer. Experiments for all samples were performed three times at each pH value.2.5. In vitro cytotoxicity study
Cytotoxicity of DOX-free PEGylated PdNPs and DOX-conjugated PEGylated PdNPs against HeLa cells was assessed in vitro by MTT assay. This assay is based on the ability of live cells to convert the MTT (solution of yellow color) into blue formazan salts. Briefly, HeLa cells were seeded into a 96-well plate at a density of 5.0 × 103 cells per well in 100 μL of complete DMEM containing 10% FBS. The cells were cultured for 1 day at 37 °C in 5% CO2 atmosphere. Afterwards, the cells were incubated with PEGylated PdNPs, DOX conjugated PEGylated PdNPs, or free DOX for 48 h at 37 °C. The concentrations of DOX-free PEGylated PdNPs ranged from 3.125 μg mL−1 to 200 μg mL−1. DOX conjugated PEGylated PdNPs or free DOX were diluted in complete DMEM with final DOX concentrations from 0.125 μg mL−1 to 8 μg mL−1. After incubation, MTT stock solution (5 mg mL−1 in PBS, 20 μL) was added to each well and incubated for 4 h. The media were completely removed and 150 μL of DMSO was added to each well to dissolve the formazan blue crystal and the absorbance was monitored using a microplate reader (Bio-TekELx800) at the wavelength of 490 nm. The cytotoxicity was expressed as the percentage of cell viability compared to untreated control cells. All experiments were repeated three times.2.6. Cellular uptake of DOX conjugated PEGylated PdNPs
CLSM was used to examine the intracellular distribution of DOX. HeLa cells were seeded on slides on a 24-well plate at a density of 5.0 × 104 cells per well in 1 mL of complete DMEM containing 10% FBS. The cells were cultured for 1 day at 37 °C in 5% CO2 atmosphere. The cells were then incubated with DOX conjugated PEGylated PdNPs at a final DOX concentration of 2 μg mL−1 in DMEM for 2 or 24 h at 37 °C. At each predetermined time, the culture media were subsequently removed and the cells were washed with PBS (1 min × 3) to remove DOX-loaded micelles that were not ingested by the cells. Thereafter, the cells were fixed with 4% (w/v) paraformaldehyde aqueous solution for 10 min at room temperature. The slides were then rinsed with PBS (5 min × 3). Finally, the cells were stained with DAPI (5 mg mL−1 in PBS) at 37 °C for 8 min, and the slides were rinsed with PBS (5 min × 3). The prepared slides were examined by CLSM (Nikon, TE2000, EZ-C1, Japan).For fluorescence microscopic study, HeLa cells were seeded in a 24-well plate and incubated for 24 hours to allow the cells to attach. A certain amount of free DOX and DOX conjugated NPs (DOX concentration 2 μg mL−1) were added, and the cells were further incubated for 48 hours. After washing the cells with PBS (pH 7.4) three times, the cellular uptake was observed by fluorescence microscope (DMIL; Leica Microsystems Ltd, Wetzlar, Germany).
2.7. AO/EtBr staining assay
Approximately 1 μL of a dye mixture (100 mg mL−1 acridine orange (AO) and 100 mg mL−1 ethidium bromide (EtBr) in distilled water) was mixed with 9 mL of cell suspension (1 × 105 cells per mL) on clean microscope cover slips. The cancer cells were collected, washed with phosphate buffered saline (PBS) (pH 7.2) and stained with 1 mL of AO/EtBr. After incubation for 2 min, the cells were washed twice with PBS (5 min each) and visualized under a fluorescence microscope (Nikon Eclipse, Inc., Japan) at 400× magnification with an excitation filter at 480 nm.2.8. DAPI staining for nuclear morphology study
For visualization of HeLa cells, the nuclei of the cells were stained with DAPI. The efficiency of DOX conjugated PEGylated PdNP was tested through apoptosis study. For this purpose, HeLa cells were treated with nanoparticles at their different concentration for 24 h at 37 °C. Then, cells were fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.1% Triton X-100, and stained with 1 mg mL−1 DAPI for 10 min. The cells were then rinsed with PBS and morphological changes were analyzed under fluorescence microscopy (Nikon Eclipse, Inc., Japan) at 400× magnification with excitation filter at 510–590 nm. The percentage of total apoptotic cells was determined by the following formula:2.9. Semi-quantitative RT-PCR analysis
RT-PCR analysis was used to examine the expression levels of Bcl-2, cytochrome c and caspase 3 and 9 mRNA. RT-PCR for β-actin was also independently performed as an internal control. The total RNA was isolated using the TRIzol reagent (Sigma-Aldrich) according to the manufacturer's instructions and reverse transcribed. Briefly, the cDNA was amplified in a 50 μL reaction containing primer pairs (1.0 μL)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10× buffer (5.0 μL), cDNA (2.0 μg), and 25 mmol L−1 MgCl (3.0 μL), 10 mmol L−1 dNTPs (1.0 μL), and Taq polymerase (2.5 U). Semi-quantitative RT-PCR amplification cycles consisted of denaturation at 94 °C for 1 min, primer annealing at 57 °C for 45 s and extension at 72 °C for 45 s, for a total of 30 cycles followed by final extension at 72 °C for 10 min. The PCR product was separated on 1.5% agarose gels. The primer sequences used for PCR were:
10× buffer (5.0 μL), cDNA (2.0 μg), and 25 mmol L−1 MgCl (3.0 μL), 10 mmol L−1 dNTPs (1.0 μL), and Taq polymerase (2.5 U). Semi-quantitative RT-PCR amplification cycles consisted of denaturation at 94 °C for 1 min, primer annealing at 57 °C for 45 s and extension at 72 °C for 45 s, for a total of 30 cycles followed by final extension at 72 °C for 10 min. The PCR product was separated on 1.5% agarose gels. The primer sequences used for PCR were:
| Gene product | Primers sequences | Product size (bp) |
|---|---|---|
| Bcl-2 | Sense 5′-CCAAGCTGAGCGAGTGTC-3′ and anti-sense 5′-ACAAAGATGGTCACGGTCTGCC-3′ | 415 |
| Caspase 3 | Sense 5′-GACAACAACGAAACCTCCGT-3′ and anti-sense 5′-GACTTCGTATTTCAGGGCCA-3′ | 382 |
| Caspase 9 | Sense 5′-TGTGGTGGTCATCCTCTCTCA-3′ and anti-sense 5′-GTCACTGGGGGTAGGCAAACT-3′ | 282 |
| Cyto c | Sense 5′-GGAGGCAAGCATAAGACTGG-3′ and anti-sense 5′-GTCTGCCCTTTCTCCCTTCT-3′ | 170 |
| β-Actin | Sense 5′AACCGCGAGAAGATGACCCAGATCATGTTT-3′ and anti-sense 5′-AGCAGCC GTGGCCATCTCTTGCTCGAAGTC-3′ | 350 |
2.10. Hemolytic assay
Ethylenediamine tetraacetic acid (EDTA)-stabilized human blood samples were freshly collected and used within 3 h of being drawn. A 4 mL sample of whole blood was added to 8 mL of phosphate-buffered saline (PBS). The PBS solution was formulated to the following composition (mM): 0.14NaCl, 2.7KCl, 10Na2HPO4, and 1.8KH2PO4. NaOH was added to the PBS solution to adjust the pH to 7.4 as measured by a calibrated, pH meter (320, Corning Inc., Corning, NY). The RBCs were isolated from serum by centrifugation at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 016g for 5 min. The RBCs were further washed five times with sterile PBS solution. Following the last wash, the RBCs were diluted to 40 mL of PBS. Then 0.2 mL of diluted RBC suspension was added to PEGylated PdNP and DOX conjugated PdNP solutions at systematically varied concentrations and mixed by vortexing. All the sample tubes were kept in static condition at room temperature for 3 h. Finally, the mixtures were centrifuged at 10
016g for 5 min. The RBCs were further washed five times with sterile PBS solution. Following the last wash, the RBCs were diluted to 40 mL of PBS. Then 0.2 mL of diluted RBC suspension was added to PEGylated PdNP and DOX conjugated PdNP solutions at systematically varied concentrations and mixed by vortexing. All the sample tubes were kept in static condition at room temperature for 3 h. Finally, the mixtures were centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 016g for 3 min, and 100 μL of supernatant of all samples was taken, and its absorbance was recorded on a spectrophotometer (Shimadzu UV-vis Spectrophotometer) at 545 nm. The percentage hemolysis was calculated using the following relationship.
016g for 3 min, and 100 μL of supernatant of all samples was taken, and its absorbance was recorded on a spectrophotometer (Shimadzu UV-vis Spectrophotometer) at 545 nm. The percentage hemolysis was calculated using the following relationship.herein, RBC incubation with D.I. water and PBS were used as the positive and negative controls, respectively.
2.11. Assessment of anti-tumor activity in vivo
The anti-tumor efficiency of DOX conjugated PEGylated PdNPs was assessed in tumor-induced mice. Briefly, the subcutaneous dorsa of BALB/c female nude mice were inoculated with HeLa cells (1 × 107) in 100 μL of normal saline. When the volume of the xeno-graft tumor reached approximately 70–100 mm3 the mice were randomly divided into four groups with six mice in each group: group A, normal saline; group B, free DOX; group C, PEGylated PdNPs; group D, DOX conjugated PEGylated PdNPs. Various DOX formulations with the drug concentration of 5 mg kg−1 was injected intravenously every 2 days, and the mice were then observed for 18 days. The tumor diameters were measured every 3 days interval for each group. The tumor volumes (V) and body weight were calculated using the formula V = [length × (width)2]/2. For the assessment of toxicity, organs such as, liver, heart, kidney, lung, and spinal cord were collected, fixed in 4% paraformaldehyde solution and made into 4 mm sections which were stained with hematoxylin and eosin (H&E) and observed under a microscope. All experiments were performed in compliance with the relevant laws and institutional guidelines (Animal Ethical Committee, Periyar University, Salem) and this work has been approved by the IAEC (Institutional Animal Ethical Committee) constituted as per the Rules and Regulations of Ministry of Animal Husbandry, Government of India.3. Statistical analysis
All the measurements were made in triplicate and all values were expressed as the mean ± standard error. The results were subjected to an analysis by Student's t-test. The results were considered statistically significant if the p-value was ≤0.05.4. Results and discussion
4.1. Synthesis and characterization of DOX conjugated PEGylated PdNPs
Hydrazone bond was employed in this work to connect DOX, as it is more sensitive to tumor microenvironments than other covalent linkers. The negatively charged group of oxygen in PEG-hydrazide could react with H2PdCl4, which were reduced to become Pd atoms by adding NaBH4 (reducing agent). The accumulating Pd atoms leads to the formation of PdNP, thus PEG assembled to be absorbed to the surface PdNPs, leaving the hydrazine group outward. Meanwhile DOX was conjugated onto the as-synthesized PEGylated PdNPs through hydrazine linkage formed between the carbonyl groups of DOX and hydrazine side groups of PEGylated PdNPs. The reaction scheme for the synthesis of DOX conjugated PEGylated PdNPs is shown in Scheme 1.The UV-vis spectra for bare DOX and DOX conjugated PEGylated PdNPs is presented in Fig. 1b. The bare DOX shows peaks at 232, 262, 290 and 490 nm.47 The conjugation of DOX onto the PEGylated PdNPs was evident from the spectrum of DOX conjugated PEGylated PdNPs solution, which clearly shows the characteristic absorption peaks of DOX. Moreover, these peaks showed red-shifts. For example, the peaks of DOX at 232, 262 and 490 nm shifted to 234, 265 and 501 nm after interaction with PEGylated PdNPs.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibrations, at 1401 cm−1 attributable to –NH3+ and at 1104 cm−1 corresponding to C–O–C are observed in PEGylated PdNPs, confirming the attachment of PEG-hydrazide onto PdNPs. FTIR was further extended to study the conjugation of DOX with the PEGylated PdNPs. FTIR spectrum of pure DOX shows peaks at 3450 cm−1 due to N–H stretching vibrations for the primary amine structure and at 3330 cm−1 due to O–H stretching vibrations. However, in case of DOX conjugated PEGylated PdNPs, peak due to N–H stretching vibrations and O–H stretching vibrations overlap, are broadened and shifted to the lower frequency range (∼3265 cm−1).1,58 Further when compared to the IR spectrum of DOX, the peaks at 1734 cm−1 corresponding to C
O stretching vibrations, at 1401 cm−1 attributable to –NH3+ and at 1104 cm−1 corresponding to C–O–C are observed in PEGylated PdNPs, confirming the attachment of PEG-hydrazide onto PdNPs. FTIR was further extended to study the conjugation of DOX with the PEGylated PdNPs. FTIR spectrum of pure DOX shows peaks at 3450 cm−1 due to N–H stretching vibrations for the primary amine structure and at 3330 cm−1 due to O–H stretching vibrations. However, in case of DOX conjugated PEGylated PdNPs, peak due to N–H stretching vibrations and O–H stretching vibrations overlap, are broadened and shifted to the lower frequency range (∼3265 cm−1).1,58 Further when compared to the IR spectrum of DOX, the peaks at 1734 cm−1 corresponding to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O disappears and C
O disappears and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond showed up at 1636 cm−1 indicative of hydrazone bond. The characteristic absorbance bands of 681 cm−1 relates to DOX. From this FTIR result, it can be interpreted that the attachment of DOX to PEGylated PdNPs occurs via the formation of hydrazone bond between the hydrazide groups of PEGylated PdNPs and the carbonyl groups of DOX.
N bond showed up at 1636 cm−1 indicative of hydrazone bond. The characteristic absorbance bands of 681 cm−1 relates to DOX. From this FTIR result, it can be interpreted that the attachment of DOX to PEGylated PdNPs occurs via the formation of hydrazone bond between the hydrazide groups of PEGylated PdNPs and the carbonyl groups of DOX.
The NMR spectra depicted in Fig. 5a authenticate the presence of PEGylated PdNPs. The respective chemical shifts peaks had been noticed at 1.98, 2.40, 3.31–3.95 minutes. Interestingly the DOX conjugated PEGylated PdNPs accentuated the characteristic peaks at 3.80, 3.47, 2.49, 1.20 minutes respectively. This signal frequency in the form of chemical shift as detected by NMR spectroscopy is proportional to the magnetic field applied to the nucleus of the DOX conjugated PEGylated PdNPs (Fig. 5b). In addition the occurrence of PEGylated PdNPs had been confirmed by showing the chemical shift peaks at 7.89, 7.61, 1.11, 1.02 minutes. Thus the Fig. 5b reveals the firm conjugation of DOX onto PEGylated PdNPs.
4.2. Doxorubicin (DOX) drug loading profile
To assess the feasibility of using PEGylated PdNPs as an anticancer drug carrier, we performed loading and in vitro DOX release studies using PEGylated PdNPs. DOX loading was attributed to the conjugation of carbonyl group in DOX to the surface active hydrazide group in PEGylated PdNPs. Before loading DOX onto the NPs, DOX·HCl was stirred with twice the number of moles of TEA in DMSO to detach HCl and render the drug hydrophobic. The characteristics of DOX conjugated PEGylated PdNPs, including DLC, particle size, PDI and zeta potential are summarized in Table S1.† The theoretical DLC was set at 10 wt%, and the results showed that the DLC of DOX conjugated PEGylated PdNPs were 8.79 wt%, implying that DOX was effectively conjugated onto the NPs. After DOX loading, PEGylated PdNPs had a larger size (approximately 40 nm) than DOX-free PEGylated PdNPs (approximately 17 nm).4.3. In vitro drug release profile
Macromolecules and particles are uptaken by cells via endocytosis mechanism. Endocytic pathway involves acidic membrane-vesicles (endosomes and lysosomes). Besides, some tumor sites have slightly acidic extracellular environment.59 In view of that, we aimed to synthesize a pH sensitive PEGylated DDS for the antitumor drug, DOX. The drug release response of PEGylated PdNPs was evaluated under different pH conditions, pH 7.4 (corresponds to the environment of blood), pH 6.8 (the pH of tumor tissue), and pH 5.5 (simulates the pH in mature endosomes of tumor cells) at a temperature of 37 °C. The temperature of 37 °C was selected for drug release response because it is close to the physiological temperature. As shown in Fig. 6 PEGylated PdNPs exhibited obvious pH-related release behavior. The result shows that at pH 7.4 the drug release was slow and sustained with release ratio at about 19 ± 0.23% in 48 h. The hydrazone linkage enables DOX conjugated PEGylated PdNPs to remain stable for a considerable period of time during circulation in the blood at pH 7.4 and thereby eliminates the premature burst release. Such stability, to a large extent, can reduce the side effects of the drug on normal cells. At pH 6.8, the drug release ratio was higher than that at pH 7.4; about 68 ± 0.27% loaded drug was released due to the slight protonation. However at pH 5.5 DOX was released more rapidly with approximately 89 ± 0.34% within the same period indicating the sensitivity of DOX conjugated PEGylated PdNP to endosomal pH. At lower pH 5.5 the degradation of hydrazone bonds of DOX conjugated PEGylated PdNPs contributes to the facile release of the conjugated drug.4.4. In vitro cytotoxicity
To evaluate the cytotoxicity of DOX conjugated PEGylated PdNP, in vitro cytotoxicity tests of free DOX, and DOX conjugated PEGylated PdNP against human against HeLa cell line were conducted for 24 h and 48 h by MTT assay at different DOX concentrations. It is well known that the biocompatibility of nanoparticles is most important for biomedical applications. To ensure that the toxicity of PEGylated PdNPs (DOX-free), in vitro cytotoxicity test for PEGylated PdNPs against HeLa cells was also conducted. No appreciable deduction in cell viability was observed for both 24 h and 48 h incubation, indicating that PEGylated PdNPs is highly biocompatible (Fig. S3†). As can be seen from Fig. 7, free DOX and DOX conjugated PEGylated PdNPs showed dose-dependent toxicity for both 24 h and 48 h. As expected, DOX delivery mediated by PEGylated PdNPs could cause substantially enhanced cytotoxicity to HeLa cells. The results (Fig. 7) indicate a significant cytotoxic effect after 24 h of incubation when free DOX at 2 μg mL−1 is used. In particular, after 24 h of incubation with free drug, the HeLa cells were 50% viable, in agreement with previous literature results60 and on further incubation for 48 h, 50% viability was seen at even lower concentration of 1 μg mL−1. Accordingly the half-maximal inhibitory concentration (IC50) value of free DOX was 2 ± 0.1 μg mL−1 and 1.0 ± 0.2 μg mL−1 for 24 h and 48 h respectively. In the case of DOX conjugated PEGylated PdNP the cytotoxic response was significantly higher when compared to free DOX. The IC50 values of DOX conjugated PEGylated PdNPs were 1.0 ± 0.3 μg mL−1 and 0.5 ± 0.1 μg mL−1 for 24 h and 48 h respectively. These results demonstrated that DOX conjugated PEGylated PdNPs showed increased toxicity compared to free DOX (*p < 0.05) and exhibited significant in vitro antitumor activity. This result is considered to be due to the enhanced loading of drug onto the PEGylated PdNPs and efficient internalization mediated by endocytosis. Thus DOX conjugated PEGylated PdNPs effectively decreased the in vitro cancer cell viability, which could imply the potential targeting effects of these nanocarriers.4.5. Cellular uptake of DOX conjugated PEGylated PdNPs
To verify intracellular drug release, DOX conjugated PEGylated PdNPs were monitored using confocal microscopy after treatment with HeLa cells for 24 and 48 h. As shown in Fig. 8, the distribution of DOX in cells was different at 24 and 48 h. Indeed, in 24 h, the red fluorescence of DOX conjugated PEGylated PdNPs was observed to accumulate around the nucleus inside the cells (though there was some red fluorescence diffused through nucleus) indicating that the DOX conjugated PEGylated PdNPs were initially located within the endosomal intracellular compartments, releasing cleaved DOX in the cytosol region in a sustained manner. With further incubation for 48 h, maximum DOX fluorescence could be detected in the nucleus. Full drug cleavage from the PEGylated PdNPs prodrug had taken place and almost all the drug molecules had migrated into the nucleus, over a prolonged incubation period (48 h) and they were eventually located in the nucleus. This suggested that the DOX conjugated PEGylated PdNPs entered into the cells via endocytosis. The same accreditation was made by Nguyen-Van Cuong61 in MCF-7 cells incubated with DOX-loaded micelle.The cellular uptake was further observed by fluorescence microscope after the cells were incubated with free DOX and DOX conjugated PEGylated PdNPs for 48 h. The negative control was HeLa cells without any treatment. Fig. S4a† shows the fluorescence images of DOX against HeLa cells with incubation times of 48 h. It was clear that at the same incubation time, the DOX fluorescence intensity in HeLa cells was enhanced when DOX was conjugated to the PEGylated PdNPs. On the basis of these fluorescence images, it can be concluded that DOX conjugated to the PEGylated PdNPs can enter into the cells and distribute throughout the nucleus more efficiently than free DOX by 48 h. An earlier report indicated that DOX fluorescence is observed only when DOX is released because of the self-quenching effect of DOX in NPs.62 In the current work, the image indicated the release of DOX from PEGylated PdNPs and its localization in nuclei. The observation can be attributed to the cleavage of hydrazone bonds in response to the intracellular pH level of endosome. Consequently, the DOX burst release from the disruptive NPs and can be readily diffused into the nuclei. This result was consistent with the cytotoxicity of pH-sensitive DOX conjugated PEGylated PdNPs showed higher cytotoxicity because of the accumulation of DOX in its intracellular active site (nuclei), which enhanced its effect. Fig. S4b† displays the mean DOX fluorescence intensity of the HeLa cells on the basis of the flow cytometry analysis. Compared with the cells incubated with free DOX, the DOX conjugated PEGylated PdNPs displays stronger fluorescence intensity. It can be seen that the fluorescence intensity of HeLa cells cultured with DOX conjugated to the PEGylated PdNPs was about 2.5 times of cells cultured with free DOX. The results clearly demonstrated that DOX conjugated to the PEGylated PdNPs enhanced the intracellular release of DOX compared with free DOX. The fluorescence signals are associated with the DOX release quantity from PEGylated PdNPs. Therefore, the enhanced intracellular DOX fluorescence in cells treated with DOX conjugated PEGylated PdNPs was due to the rapid and complete intracellular DOX release. This result was in accordance with our expectation that hydrazone bonds would be cleaved in the intracellular environment. Hence, PEGylated PdNPs was a suitable anticancer drug carrier.
To investigate quantitatively the delivery of DOX conjugated PEGylated PdNPs to the nucleus, cell fractionation experiments were done. Cellular and nuclear uptake kinetics of free DOX and DOX conjugated to the PEGylated PdNPs was examined for HeLa cells at elapsed time points, shown in Fig. S5.† Cellular and nuclear uptakes of DOX conjugated to the PEGylated PdNPs were time dependent. In the case of free DOX most of the drug was found in the nuclear fraction already after 2 h of incubation. The intense DOX accumulation in the nucleus for free DOX occurred because intracellular DOX molecules in the cytosol could transport rapidly to the nucleus and intensely bound to the chromosomal DNA.63 In fact, the drug concentration in cells treated with free DOX did not increase at all when incubation was prolonged from 2 h to 48 h. This might be due to the fact that free DOX without drug carrier releases rapidly and the release is almost completed within 8 h.64 In contrast to free DOX, DOX conjugated PEGylated PdNPs were mainly distributed in the cytoplasm without exhibiting much accumulation in the nuclear fraction. With further incubation for 12 h, the DOX molecules inside the cells increased and the accumulation of DOX in the cytoplasmic fraction were later observed to slowly migrate into the nucleus by 24 h. Over a more prolonged incubation period (48 h) DOX molecules were eventually located in the nuclear fraction and became more evident than 24 h of incubation. Almost only little DOX was observed in the cytoplasmic fraction, indicating that maximum DOX molecules from PEGylated PdNPs solely located in nuclear fraction at 48 h incubation. Thus the nuclear drug concentration obtained with DOX conjugated PEGylated PdNPs clearly surpasses the concentration obtained with free DOX, especially after 48 h of incubation. The enhanced uptake of DOX conjugated PEGylated PdNPs can be attributed to their facilitated endocytotic transport, relative to passive diffusion of free DOX through the cell membrane.
4.6. Fluorescence microscopic studies
4.7. Effect of DOX conjugated PEGylated PdNPs on intrinsic apoptosis
Cell death via apoptosis is an important event involved in a number of immunological processes. As most of the anticancer drugs are believed to trigger apoptosis via mitochondria-mediated pathway,67,68 we here hypothesize that, as a new hybrid system, DOX conjugated PEGylated PdNPs might also initiate the apoptosis via mitochondria-mediated pathway. Under this premise, we studied the changes of the levels of the mitochondrial-dependent apoptotic proteins, including the caspase-3, the most important effector caspase, caspase-9, Bcl-2 and cytochrome c. Previous studies demonstrated that down regulation of anti-apoptotic protein Bcl-2 leads to release of cytochrome c from the mitochondria to cytosol, which is an essential step in the induction of apoptosis. Cytochrome c release from mitochondria to cytosol in turn leads to the activation of the caspase cascade such as caspase-3 and 9 which is critical in executing apoptosis, as it is either partially or totally responsible for the proteolytic cleavage of many key proteins.69 Thus it is remarkable to speculate the analysis of Bcl-2, cytochrome c, and caspases-3 and 9 gene expressions. The results (Fig. 10) revealed a significant decrease in the expression of Bcl-2 and with an significant increase in the expression of cytosolic cytochrome c and caspase-3 and 9 in cells treated with DOX conjugated PEGylated PdNPs compared to untreated control.4.8. Hemolysis activity
Determination of hemolytic properties is one of the most common tests in studies of nanoparticle interactions with blood components.70 Erythrocyte interaction with nanoparticles is particularly important in the application of nanoparticles for biological applications.71 Two different methods were used to assess the hemolytic potential of PEGylated PdNP and DOX conjugated PEGylated PdNP; hemoglobin release analysis and cell morphology analysis. The RBCs were exposed to each NP sample for 3 h. Hemoglobin release analysis (Table S2†) shows the hemolytic activity of control, PEGylated PdNP and DOX conjugated PEGylated PdNP. When water is added to RBCs, hemolysis takes place and the released haemoglobin is measured. This serves as a positive control and represents absorbance to be 3.14 ± 0.062 (100% haemolysis). When, PEGylated PdNP and DOX conjugated PdNP was added, hemolysis was found to be less than 5% and are comparable to that suspended in PBS with absorbance of 0.02 ± 0.003 (0% hemolysis) which acts as a negative control. Fig. 11a shows photographs of the hemolytic test on both the nanoparticle samples. The supernatant from PEGylated PdNP and DOX conjugated PdNP at different concentrations is achromatic, implying that no significant hemolysis occurred. It has been reported that up to 5% hemolysis is permissible for biomaterials.72 Thus, both PEGylated PdNP and DOX conjugated PEGylated PdNP at the tested concentration exhibited no significant hemolysis. The largest percentage hemolysis obtained was 1.44 ± 0.027% for 200 μg mL−1 PEGylated PdNP and 1.57 ± 0.054% for DOX conjugated PEGylated PdNP at 8 μg mL−1 DOX concentration. Since this is much lower than 5%, it indicates that both PEGylated PdNP and DOX conjugated PEGylated PdNP are hemocompatible for drug delivery applications. The cell morphology analysis (Fig. 11b) corroborated the hemoglobin release analysis results. The cell morphology analysis indicated that incubation of RBCs with 200 μg mL−1 PEGylated PdNP and DOX conjugated PEGylated PdNP (8 μg mL−1 DOX concentration) did not result in hemolysis or change in morphology of red blood cells when compared to control. Yu-Shen Lin et al.73 showed the influence of PEG surface coating on hemolytic activity of mesoporous silica nanoparticle (MS NPs). The authors report that contrary to bare MS NPs, no apparent hemolysis was observed for PEG-coated MS NPs after 3 h blood incubation. In our study, the absence of hemolysis maybe due to biocompatible polymer PEG coating which prevented the adhesion of both the NPs to red blood cell membrane. Thus this simple surface modification stratagem is critical to ensure the safety of DOX conjugated PEGylated PdNP in biomedical applications.4.9. In vivo anti-tumor activity
In order to conform the feasibility of DOX-conjugated NPs for cancer therapy in vivo, the normal saline, DOX conjugated PEGylated PdNPs, PEGylated PdNPs and free DOX were injected at a dose of 5 mg DOX/kg body weight through the tail vein of the mice bearing cervical carcinoma HeLa cells and the anti-tumor activity of NPs was assessed. Here, the tumor volume was recorded for a period of 18 days after the start of treatment, and mice injected with saline were treated as the control group. Fig. 12a shows the tumor growth curves after 18th day of treatment. The tumor volume of mice receiving normal saline, PEGylated PdNPs, rapidly increased at the end of 18th day and was observed to be 1102.27 ± 0.12 mm3 and 1050.03 ± 1.67 mm3, respectively. There were no significant differences of the tumor inhibition rate among the groups treated with normal saline solution and PEGylated PdNPs. It was clearly indicated that the growth of tumor was significantly (*p < 0.05) suppressed by the treatment of DOX conjugated PEGylated PdNPs and free DOX group when compared to the respective normal saline group. These results indicated that DOX conjugated PEGylated PdNPs could improve DOX delivery into the HeLa tumor by passive targetting. Moreover, the tumor growth suppression volume of the DOX conjugated PEGylated PdNPs group was 164.74 ± 0.17 mm3 than that of the free DOX group 500.02 ± 1.34 mm3, indicating the enhanced anti-tumor activity of DOX conjugated PEGylated PdNPs. The high antitumor activity of the DOX conjugated PEGylated PdNPs can be attributed to a higher accumulation in cancer cells, a controlled release feature and a decreased influence of MDR effect, as suggested earlier. Thus we inferred that DOX conjugated PEGylated PdNPs are uptaken by the tumor cells via an endocytic process. Once entering the cells, DOX is released from the DOX conjugated PEGylated PdNPs triggered by the acidic endocytic environment thereby greatly enhancing the anti-tumor activity. It is reported that the delivery of targeted NPs facilitates gradual accumulation of NPs in tumor tissue via endocytosis resulting in a potent anti-tumor activity. However, non-conjugated NPs remain in extracellular matrix of the tumor tissue and undergo degradation or phagocytosis, resulting in release of the drug. The results are nearly similar as those reported elsewhere.74 As summarized in Fig. 12a, although the tumor growth was inhibited after DOX conjugated PEGylated PdNPs treatment, it is critical to evaluate the mice body weight loss. The fluctuation in animal body weight is recognized as a useful indicator to assess in vivo toxicity of drug delivery systems. Mice administrated with saline showed a steadily increasing body weight. Similar to the control group, mice administered with blank PEGylated PdNPs and DOX conjugated PEGylated PdNPs exhibited no decline in body weight, indicating the nontoxicity of the NPs. In contrast, the body weight of DOX treated group of animals sharply decreased, compared to control and DOX conjugated PEGylated PdNPs groups, (Fig. 12b), suggesting that toxic side effects were induced by DOX at the given dose. Further, histological analysis of mice treated with normal saline, PEGylated PdNPs and DOX conjugated PEGylated PdNPs revealed no significant signal of damage from H&E stained organ slices including liver, heart, kidney, lung, and spinal cord (Fig. 12c). However, for group treated with DOX, acute inflammatory cell infiltration with obvious organ damage of necrosis were apparent in heart and kidney tissues compared with the muscle fibers and organ structure from control mice. These findings indicated that free DOX was delivered not only in tumor cells but also to other normal cells and caused side effects; whereas DOX conjugated PEGylated PdNPs can reduce assorted side effects. It is expected that most DOX conjugated with PEGylated PdNPs will remain on the particle surface via a pH sensitive hydrazone bond for a considerable length of time in the blood at normal physiological conditions (pH 7.4), which greatly reduces the exposure of DOX to normal tissues and thus decreased the toxicity and adverse side effect of DOX while effectively enhancing its anticancer activity.755. Conclusion
To our knowledge, this is the first report on using PEGylated PdNPs as drug delivery system to deliver a chemotherapeutic agent into cancer cells. In conclusion, we have developed as a proof of concept, a novel pH-responsive DOX conjugated PEGylated PdNPs as a new delivery vehicle. For the first time, that the use of PEG as caps on the surface of palladium nanocarrier provides a suitable method for the design of delivery system able to selectively release conjugated cargos in responsive to cancer cells. The uniqueness of this drug delivery NP system is that the DOX was linked through pH-sensitive hydrazone bond to the hydrazide moiety of PEGylated PdNPs. These NPs had well-controlled DOX loading yield, enhanced cellular uptake properties and showed excellent pH responsive drug release kinetics, leading to enhanced in vitro and in vivo cytotoxicity against HeLa cells as compared to free DOX. As an environmentally sensitive drug delivery vehicle, these NPs can potentially minimize the drug loss during their circulation in the blood, where the pH value is neutral, and trigger rapid intracellular drug release when the NPs are endocytosed by the target cells. This characteristic drug release kinetics may suppress cancer cell chemoresistance and improve the therapeutic efficacy of the drug payload. In summary, the proposed new family of DOX conjugated PEGylated PdNP mediated drug delivery system has potential for tumor targeting and controlled release.Conflict of interest
No conflict of interest was reported by the author of this article.Acknowledgements
This research work was partially supported by UGC – Basic Science for Research (BSR) – RFSMS (Ref. G2/3142/UGC – BSR – RFSMS/2013) and DST-Nano-mission Project, Department of Science and Technology, Nano-mission division, New Delhi (Ref. SR/NM/NS-60/2010 dt. 08-07-2011).References
- S. Kayal and R. V. Ramanujan, Mater. Sci. Eng., C, 2010, 30, 484–490 CrossRef CAS PubMed.
- H. Yang, S. Y. Fung and M. Liu, Angew. Chem., Int. Ed., 2011, 50, 9643–9646 CrossRef CAS PubMed.
- D. A. Gewirtz, M. L. Bristol and J. C. Yalowich, Curr. Opin. Invest. Drugs, 2010, 11, 612–614 CAS.
- M. Ferrari, Nat. Rev. Cancer, 2005, 5, 161–171 CrossRef CAS PubMed.
- W. Von Rybinski and M. J. Schwuger, Adsorption and wetting, in Nonionic Surfactants: Physical Chemistry, Surfactant Sci. Ser., ed. M. J. Schick, Marcel Dekker, New York, 1987, vol. 23 Search PubMed.
- K. E. Bremmell, S. R. Biggs and G. J. Jameson, Colloids Surf., A, 1999, 146, 75–87 CrossRef CAS.
- K. Sathish kumar, J. Madhusudhanan, R. A. Thanigaivel and V. A. Veni, Res. J. BioTechnol., 2013, 8, 70–77 Search PubMed.
- J. Z. Du, X. J. Du, C. Q. Mao and J. Wang, J. Am. Chem. Soc., 2011, 133, 17560–17563 CrossRef CAS PubMed.
- T. Zhou, C. Xiao, J. Fan, S. Chen, J. Shen, W. Wu and S. Zhou, Acta Biomater., 2013, 9, 4546–4557 CrossRef CAS PubMed.
- Y. Zhu, J. Shi, W. Shen, X. Dong, J. Feng, M. Ruan and Y. Li, Angew. Chem., Int. Ed., 2005, 117, 5213–5217 CrossRef PubMed.
- W. H. Chiang, V. T. Ho, W. C. Huang, Y. F. Huang, C. S. Chern and H. C. Chiu, Langmuir, 2012, 42, 15056–15064 CrossRef PubMed.
- D. Hua, J. Jiang, L. Kuang, J. Jiang, W. Zheng and H. Liang, Macromolecules, 2011, 44, 1298–1302 CrossRef CAS.
- R. Liu, P. Liao, J. Liu and P. Feng, Langmuir, 2011, 27, 3095–3099 CrossRef CAS PubMed.
- L. E. Gerweck and K. Seetharaman, Cancer Res., 1996, 56, 1194–1198 CAS.
- H. Sun, B. Guo, X. Li, R. Cheng, F. Meng, H. Liu and Z. Zhong, Biomacromolecules, 2010, 4, 848–854 CrossRef PubMed.
- W. Gao, J. M. Chan and O. C. Farokhzad, Mol. Pharmaceutics, 2010, 6, 1913–1920 CrossRef PubMed.
- D. Wang, X. Huan, L. Zhu, J. Liu, F. Qiu, D. Yan and X. Zhu, RSC Adv., 2012, 2, 11953–11962 RSC.
- X. Hu, X. Hao, Y. Wu, J. Zhang, X. Zhang, P. C. Wang, G. Zou and X. J. Liang, J. Mater. Chem. B, 2013, 1, 1109–1118 RSC.
- A. Popat, J. Liu, G. Q. M. Lu and S. Z. J. Qiao, J. Mater. Chem., 2012, 22, 11173–11178 RSC.
- P. Tobiska, O. Hugon, A. Trouillet and H. Gagnaire, Sens. Actuators, B, 2001, 74, 168–172 CrossRef CAS.
- Z. H. Chen, J. S. Jie, L. B. Luo, H. Wang, C. S. Lee and S. T. Lee, Nanotechnology, 2007, 34, 345502 CrossRef.
- P. R. West, S. Ishii, G. V. Naik, N. K. Emani, V. M. Shalaev and A. Boltasseva, Laser Photonics Rev., 2010, 6, 795–808 CrossRef PubMed.
- J. L. Malleron, J. C. Fiaud and J. Y. Legros, Handbook of palladium-catalyzed organic reactions, Academic, London, 2000 Search PubMed.
- M. O. Nutt, J. B. Hughes and S. W. Michael, Environ. Sci. Technol., 2005, 39, 1346–1353 CrossRef CAS.
- F. Favier, E. C. Walter, M. P. Zach, T. Benter and R. M. Penner, Science, 2001, 293, 2227–2231 CrossRef CAS PubMed.
- S. Yu, U. Welp, L. Z. Hua, A. Rydh and W. K. Kwok, et al., Chem. Mater., 2005, 17, 3445–3450 CrossRef CAS.
- Z. Chang, H. Fan, K. Zhao, M. Chen and P. He, et al., Electroanalysis, 2008, 20, 131–136 CrossRef CAS PubMed.
- C. P. Adams, K. A. Walker, S. O. Obare and K. M. Docherty, PLoS One, 2014, 9(1), e85981, DOI:10.1371/journal.pone.0085981.
- J. W. Xiao, S. X. Fan, F. Wang, L. D. Sun, X. Y. Zheng and C. H. Yan, Nanoscale, 2014, 6, 4345 RSC.
- I. Eryazici, C. N. Moorefield and G. R. Newkome, Chem. Rev., 2008, 108, 1834–1895 CrossRef CAS PubMed.
- S. D. Cummings, Coord. Chem. Rev., 2009, 253, 1495–1516 CrossRef CAS PubMed.
- G. Tamasi, M. Casolaro, A. Magnani, A. Sega, L. Chiasserini, L. Messori, C. Gabbiani, S. M. Valiahdi, M. A. Jakupec, B. K. Keppler, M. B. Hursthouse and R. Cini, J. Inorg. Biochem., 2010, 104, 799–814 CrossRef CAS PubMed.
- A. Divsalar, A. A. Saboury, H. Mansoori-Torshizi and F. Ahmad, J. Phys. Chem. B, 2010, 114, 3639–3647 CrossRef CAS PubMed.
- E. Ulukaya, F. Ari, K. Dimas, E. I. Ikitimur, E. Guney and V. T. Yilmaz, Eur. J. Med. Chem., 2011, 46, 4957–4963 CrossRef CAS PubMed.
- A. Abuchowski, T. Van Es, N. C. Palczuk and F. F. Davis, J. Biol. Chem., 1977, 252, 3578–3581 CAS.
- L. T. M. Hoa, T. T. Dung, T. M. Danh, N. H. Duc and D. M. J. Chien, J. Phys.: Conf. Ser., 2009, 187, 012048 CrossRef.
- Y. Zhang, N. Kohler and M. Zhang, Biomaterials, 2002, 7, 1553–1561 CrossRef.
- A. Di Marco, M. Gaetani and B. Scarpinato, Cancer Chemother. Rep., 1969, 53, 33–37 CAS.
- T. Teranishi and M. Miyake, Chem. Mater., 1998, 10, 594–600 CrossRef CAS.
- X. Yang, Q. Li, H. Wang, J. Huang, L. Lin, W. Wang, D. Sun, Y. Su, J. B. Opiyo, L. Hong, Y. Wang, N. He and L. Jia, J. Nanopart. Res., 2010, 12, 1589–1598 CrossRef CAS.
- P. Mulvaney, Langmuir, 1996, 12, 788–800 CrossRef CAS.
- C. C. Luo, Y. H. Zhang and Y. G. Wang, J. Mol. Catal. A: Chem., 2005, 229, 7–12 CrossRef CAS PubMed.
- K. Okitsu, H. Bandow, Y. Maeda and Y. Nagata, Chem. Mater., 1996, 8, 315–317 CrossRef CAS.
- T. Yonezawa, K. Imamura and N. Kimizuka, Langmuir, 2001, 17, 4701–4703 CrossRef CAS.
- P. F. Ho and K. M. Chi, Nanotechnology, 2004, 15, 1059–1064 CrossRef CAS.
- S. Sudeshna, S. Dipankar, D. Piyali, L. Rima and S. Amitabha, Tetrahedron, 2009, 65, 4367–4374 CrossRef PubMed.
- H. Huang, Q. Yuan, J. S. Shah and R. D. K. Misra, Adv. Drug Delivery Rev., 2011, 63, 1332–1339 CrossRef CAS PubMed.
- C. Wang, L. Feng, H. Yang, G. Xin, W. Li, J. Zheng, W. Tian and X. Li, Phys. Chem. Chem. Phys., 2012, 38, 3233–13238 Search PubMed.
- D. C. Litzinger, A. M. J. Buiting, N. Rooijen and L. Huang, Biochim. Biophys. Acta, 1994, 1190, 99–107 CrossRef CAS.
- H. Maeda and Y. Matsumura, Crit. Rev. Ther. Drug Carrier Syst., 1989, 6, 193–210 CAS.
- C. Li, Adv. Drug Delivery Rev., 2002, 54, 695–713 CrossRef CAS.
- A. Mandal, V. Meda, W. J. Zhang, K. M. Farhan and A. Gnanamani, Colloids Surf., B, 2012, 90, 191–196 CrossRef CAS PubMed.
- E. Illés, E. Tombácz, M. Szekeres, I. Y. Tóth, Á. Szabó and B. Iván, J. Magn. Magn. Mater., 2015, 380, 132–139 CrossRef PubMed.
- C. Barrera, A. Herrera, Y. Zayas and C. Rinaldi, J. Magn. Magn. Mater., 2009, 321, 1397–1399 CrossRef CAS PubMed.
- C. He, Y. Hu, L. Yin, C. Tang and C. Yin, Biomaterials, 2010, 31, 3657–3666 CrossRef CAS PubMed.
- F. Alexis, E. Pridgen, L. K. Molnar and O. C. Farokhzad, Mol. Pharm., 2008, 5, 505–515 CrossRef CAS PubMed.
- P. Aggarwal, J. B. Hall, C. B. McLeland, M. A. Dobrovolskaia and S. E. McNeil, Adv. Drug Delivery Rev., 2009, 61, 428–437 CrossRef CAS PubMed.
- S. Kayal and R. V. Ramanujan, J. Nanosci. Nanotechnol., 2010, 10, 1–13 CrossRef PubMed.
- L. Wong, M. Kavallaris and V. Bulmus, Polym. Chem., 2011, 2, 385–393 RSC.
- J. Bénard, J. Da Silva, M. C. De Blois, P. Boyer, P. Duvillard, E. Chiric and G. Riou, Cancer Res., 1985, 45, 4970–4979 Search PubMed.
- N. V. Cuong, J. L. Jiang, Y. L. Li, J. R. Chen, S. C. Jwo and M. F. Hsieh, Cancers, 2011, 3, 61–78 CrossRef CAS PubMed.
- Y. Bae, S. Fukushima, A. Harada and K. Kataoka, Angew. Chem., Int. Ed., 2003, 42, 4640–4643 CrossRef CAS PubMed.
- E. R. Gillies and J. M. J. Frchet, Bioconjugate Chem., 2005, 16, 361–368 CrossRef CAS PubMed.
- L. Jing, X. Liang, X. Li, Y. Yang and Z. Dai, Acta Biomater., 2013, 9, 9434–9441 CrossRef CAS PubMed.
- R. Vivek, R. Thangam, K. Muthuchelian, P. Gunasekaran, K. Kaveri and S. Kannan, Process Biochem., 2012, 47, 2405–2410 CrossRef CAS PubMed.
- S. Banalata, K. Sanjana, P. Devi, B. Rakesh, K. M. Tapas, P. Panchanan and D. Dibakar, ACS Appl. Mater. Interfaces, 2013, 5, 3884–3893 Search PubMed.
- A. S. Don and P. J. Hogg, Trends Mol. Med., 2004, 10, 372–378 CrossRef CAS PubMed.
- J. M. Grad, E. Cepero and L. H. Boise, Drug Resist. Updates, 2001, 4, 85–91 CrossRef CAS PubMed.
- R. Vivek, R. Thangam, V. NipunBabu, C. Rejeeth, S. Sivasubramanian, P. Gunasekaran, K. Muthuchelian and S. Kannan, ACS Appl. Mater. Interfaces, 2014, 6, 6469–6480 CAS.
- F. Y. Cheng, C. H. Su, Y. S. Yang, C. S. Yeh, C. Y. Tsai, C. L. Wu, M. T. Wu and D. B. Shieh, Biomaterials, 2005, 26, 729–738 CrossRef CAS PubMed.
- R. K. Kainthan, M. Gnanamani, M. Ganguli, T. Ghosh, D. E. Brooks, S. Maiti and J. N. Kizhakkedathu, Biomaterials, 2006, 27, 5377 CrossRef CAS PubMed.
- J. P. Singhal and A. R. Ray, Biomaterials, 2002, 23, 1139–1145 CrossRef CAS.
- Y. S. Lin and C. L. Haynes, J. Am. Chem. Soc., 2010, 132, 4834–4842 CrossRef CAS PubMed.
- S. Aryal, J. J. Grailer, S. Pilla, D. A. Steeber and S. Gong, J. Mater. Chem., 2009, 19, 7879–7884 RSC.
- K. T. Oh, H. Yin, E. S. Lee and Y. H. Bae, J. Mater. Chem., 2007, 17, 3987–4001 RSC.
Footnote |
| † Electronic supplementary information (ESI) available: Details on the hydrodynamic diameters and surface zeta potential value of PEGylated PdNPs and DOX conjugated PEGylated PdNPs, the cytotoxicity of PEGylated PdNPs against HeLa cells as determined by MTT assay, cellular uptake and intracellular localization of free DOX and DOX conjugated PEGylated PdNPs in HeLa cells observed by fluorescence microscopy. The cellular and nuclear uptakes of free DOX and DOX conjugated PEGylated PdNPs in HeLa cells. Tables showing the characteristics of PEGylated PdNPs and DOX conjugated PEGylated PdNPs and percent haemolysis for positive control, negative control, PEGylated PdNPs and DOX conjugated PEGylated PdNPs at different concentrations for 3 h are given. See DOI: 10.1039/c5ra05803a |
| This journal is © The Royal Society of Chemistry 2015 |