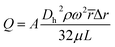Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Isothermal solid-phase recombinase polymerase amplification on microfluidic digital versatile discs (DVDs)†
Luis A.
Tortajada-Genaro
*a,
Sara
Santiago-Felipe
a,
Mary
Amasia
b,
Aman
Russom
b and
Ángel
Maquieira
a
aDepartamento de Química, Instituto Interunversitario de Reconocimiento Molecular y Desarrollo Tecnológico (IDM), Universidad Politécnica de Valencia, Spain. E-mail: luitorge@qim.upv.es
bDiv. of Nanobiotechnology, KTH Royal Institute of Technology, Stockholm, Sweden
First published on 27th March 2015
Abstract
A new advancement in massive DNA-based screening in limited-resource settings is demonstrated through the incorporation of easy-to-fabricate microfluidic chambers on digital versatile discs (DVDs) to perform isothermal recombinase polymerase amplification (RPA) in a microarray format. Standard un-modified DVD discs and commercial drives are used for the low-cost detection method. DNA primers were printed in a microarray format on the polycarbonate surfaces of DVDs with integrated control spots to guarantee the absence of false-negatives and false-positives. The solid-phase amplification assay, including the washing protocols and development reaction, was performed by the dispensation of solutions through the inlet and by controlling the flow-movement by DVD drive centrifugation. The final disc with reaction products was inserted into a DVD player and microarray images were captured and automatically processed. This simple approach was applied for the screening of genetically modified organisms (GMOs) in food samples. The limit of detection was 7 μg g−1, which is well below the EU regulation limit for GMOs in food products. Therefore, the only required materials for food safety monitoring were standard store-bought DVDs, plastic chambers, tips, pipettes, an oven, and a standard DVD drive. The proposed strategy allows an integrated microarray system with low manipulation, reduced sample volume, and portability, which are beneficial for low-resource settings.
Introduction
The development of DNA biosensors is related to the adequate selection and integration of support, probes, assay format, and transduction phenomena that are needed to perform and detect a biorecognition. Unlike silicon chips, plastic polymers, as analytical platforms, offer the advantages of being transformed easily and cheaply into devices that combine operations of sample treatment, fluid management, and detection.1 However, in some cases, the proposed platforms are not useful for real applications because the systems for fluid management and signal detection are not easily adaptable to a wide range of scenarios.Several research groups and companies have been working on the development of biosensors based on the use of compact discs or ‘lab-on-a-CD’ systems. There are two main approaches depending on the nature of the disc used.2 The first one involves the use of plastic substrates that are several mm thick with circular shapes, which are integrated in a microfluidic system (such as, microchannels, valves, and chambers).3,4 Some of them, known as micro-total analysis systems (μTAS), integrate all the analytical steps that are required for genomic assays.5 Other lab-on-a-CD devices present a lower integration level, and the procedures involve some handling steps.6 In both the cases, the measurement is generally performed with instruments, such as colorimeters or expensive static detectors such as fluorescence microscopes or other complex non-integrated systems.
The second approach directly uses audio-video compact discs as the support for carrying out bioassays, and the detection is based on the scanning of a focused laser that is present in conventional disc drives.7 The main advantage of these technologies (namely, CD, DVD, and Blu-Ray) is that they are mass-produced for the consumer electronic market with high-quality standards and cost-effective prices. Our group has demonstrated that it is possible to use a CD player/writer as a detector, using low-reflective discs (transmission/reflection mode) or conventional discs (reflection mode); moreover, the chemical modification of the surfaces can be incorporated. These systems show a higher sensitivity and working capacity (e.g. multiplexing), allowing the implementation of extremely inexpensive optical devices for biological applications.8–10
The development of DNA hybridization assays has been addressed using centrifugal disc platforms. The procedures include flow hybridization in various types of reservoirs, such as double-spiral,11 channel,12,13 or chamber,14 and all of them were combined with fluorescence detection. An interesting approach is the integration of isothermal amplification and fluorescent real-time detection with commercially available centrifugal disc and analyzer.6 Moreover, hybridization assays on a microarray format have been performed based on DVD9 and BD10 technology (e.g., disc and detector). The experimental steps are similar to those when glass or other solid supports are used. A distinct advantage is that the use of expensive and bulky scanners, which typically image the spots, is avoided, paving the way for the widespread use of the microarray technology. Nevertheless, integrated approaches are required to reduce the number of steps and the manipulation of samples.
In a recent study, Santiago-Felipe et al.15 demonstrated the advantages of isothermal recombinase polymerase amplification (RPA) combined with DVD hybridization and detection using a technique called as solid-phase amplification. In this approach, one primer is attached onto the polycarbonate surface of a DVD (bottom layer), while the other amplification components are retained in the liquid phase. The polymerase extension of the primer produces a tethered and detectable amplification product. The results have shown that this can be a new strategy to integrate amplification and hybridization into a same platform at a constant low temperature, avoiding the use of devices with technologically complex heating/cooling systems. On the other hand, Russom et al.16 have shown how merging optical discs and microfluidics is a new step towards the low-cost point-of-care applications. Low reflectivity DVDs were fabricated from 0.6 mm DVD substrates, including a spiral groove of 0.74 mm track pitch, which was coated with a 10 nm thick layer of SiO2. The microfluidic layers, containing microchannels and other fluidic reservoirs, were incorporated over the metallic layer of the discs (top layer). Then, the integration of assay development, controlled by spinning rate, and reading (transmission mode) in only one platform was achieved. The system was applied to low-cost HIV diagnostics by counting CD4+ cells isolated from whole blood. However, this approach requires some disc modifications and incorporation of a planar photodiode into the DVD drive to detect the transmitted light.
In this study, a semi-automated DNA assay in microarray format is proposed based on the integration of a simple adhesive microfluidic layer onto the polycarbonate surface of conventional DVDs (bottom layer). Primers are pre-printed onto the DVD surface, and lyophilized RPA reagents are stored within a spinning platform. The rest of the reagents are hand-dispensed and the solid-phase amplification is performed, resulting in the attachment of the amplified products onto the DVD surface. After the development of the amplification products, the microfluidic layer is removed and the disc is read by reflection mode (i.e., conventional DVD drive). The presence of the amplification product modifies the light intensity of the scanning laser of the DVD drive (reflection mode), and using a data acquisition software, a microarray image is generated. As proof of concept, the method has been applied for the low-cost, reliable, rapid screening of genetically modified organisms (GMOs).
Methods
Target genes
GMO testing was based on the determination of several genetic elements. Screening elements were the two most common transgenic genes (namely, 35S-promoter from cauliflower mosaic virus, or p35S, and nopaline synthase terminator, or tNOS), allowing the detection of most of the authorized or unauthorized lines.17,18 Taxon-specific elements detect genes that are specific for plants, such as lectin (Le1) for Glycine max (soybean), alcohol dehydrogenase 1 (adh1) for Zea mays (maize), and LAT52 protein (LAT52) for Solanum lycopersicum (tomato). These elements increase the characterization of involved transgenic ingredients, allowing increased selectivity for certain GMOs.19 Construction-specific elements are included for the complete identification of GMO events such as Bt-11 construction, which involves a junction region between the intron 6 (IVS6) from a maize alcohol dehydrogenase 1 gene (adh1-1S) and a synthetic cryIA(b) gene.20Integrated DVD system design
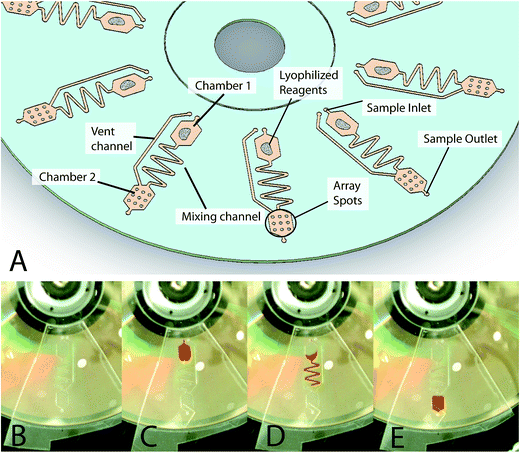
The DVD-based bioanalytical platform consists of two disc substrates (namely, optical layer and microfluidic layer) bonded together (see ESI†). The optical layer substrate used is a standard store-bought DVD-ROM disc purchased from MPO Iberica (Spain). According to the DVD specifications, a standard blank disc is composed of two 0.6 mm thick polycarbonate substrates with a middle layer of highly reflective metallic material (thickness 1.000–1.500 Å). The polycarbonate layer at the bottom had an injection molded spiral microguide (track pitch of 0.74 mm) to guide the subsystems of the detector laser (λ = 650 nm) along the data track.The microfluidic substrate was fabricated using a 0.2 mm pressure-sensitive adhesive (PSA) (adhesive transfer tape 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 022, 3 M, USA) bonded to a disc-shaped piece of polycarbonate plastic (thickness = 0.6 mm) with drilled access through-holes (diameter = 1 mm). Ten identical fluidic structures, containing microfluidic components, including channels and reservoirs, were radially arrayed to enable multiplexed assays on a single disc using a CO2 laser cutter (Hylax Hypertronics). A scheme of this device is shown in Fig. 1 with details of the two chambers and channel locations. Each structure has a chamber for pre-amplification mixing (Fig. 1A, Chamber 1) and a chamber for solid-phase amplification and detection (Fig. 1A, Chamber 2). The dimensions of the chambers are 5.5 mm in length, 5.5 mm in width and 0.2 mm in height, and therefore can contain a sample volume of 6 μL. The two chambers are connected by a 0.6 mm wide mixing channel as the hydrophobic valve. The disc was designed to enable these simple fluidic steps at low spinning rates (<1500 rpm) that are achievable in commercial DVD drives.
022, 3 M, USA) bonded to a disc-shaped piece of polycarbonate plastic (thickness = 0.6 mm) with drilled access through-holes (diameter = 1 mm). Ten identical fluidic structures, containing microfluidic components, including channels and reservoirs, were radially arrayed to enable multiplexed assays on a single disc using a CO2 laser cutter (Hylax Hypertronics). A scheme of this device is shown in Fig. 1 with details of the two chambers and channel locations. Each structure has a chamber for pre-amplification mixing (Fig. 1A, Chamber 1) and a chamber for solid-phase amplification and detection (Fig. 1A, Chamber 2). The dimensions of the chambers are 5.5 mm in length, 5.5 mm in width and 0.2 mm in height, and therefore can contain a sample volume of 6 μL. The two chambers are connected by a 0.6 mm wide mixing channel as the hydrophobic valve. The disc was designed to enable these simple fluidic steps at low spinning rates (<1500 rpm) that are achievable in commercial DVD drives.
Standard commercial DVD-ROMs were firstly conditioned by gentle ethanol washing, water rinsing, and drying by centrifugation. Biotinylated primers (Table SI.1†) were immobilized on passively adsorbed streptavidin. For immobilization, each mixture of streptavidin (5 mg L−1) and biotinylated-labelled primer (100 nM) in 50 mM carbonate buffer, pH 9.6, and 1% glycerol (v/v) was printed on a polycarbonate disc surface (50 nL) with a non-contact printer (AD 1500 BioDot Inc., CA). Working temperature and relative humidity were adjusted at 25 °C and 90%, respectively. Because this arrayer is traditionally used for printing on standard glass slides, a custom printing-layout was developed to print multiple arrays. The printing area for each chamber was 4 × 4 mm2 with an allowance for minor misalignment with the printer. In a single run, 10 arrays of 9 spots (3 × 3) for primers, negative controls, and positive controls were spotted in the Chamber 2 region with a 1 mm track pitch. Pre-stored lyophilized reagents (0.8 mg) for amplification were dispensed into Chamber 1 with a spatula. The reagent mixture composed of 2 mM DTT, 5% Carbowax 20 M, 200 μM dNTPs, 3 mM ATP, 50 mM phosphocreatine, 100 ng μL−1 creatine kinase, 30 ng μL−1 Bsu, and recombinase proteins (namely, 900 ng μL−1 gp32, 120 ng μL−1 uxsX, and 30 ng μL−1 uvsY) was supplied by TwistDx (UK). Finally, the microfluidic layer was aligned with the DVD-ROM surface and affixed to enable a bubble-free flat disc. The outer holes were sealed with PCR sealer tape (Corning, USA), and the disc was stored at −20 °C until use.
DVD drive
The assay performed on the disc was controlled and measured by an adapted DVD drive from LG Electronics Inc. (Englewood Cliffs, USA). The device incorporated a data acquisition board (model DT9832A-02-OEM, Data Translation, Germany) with a sample rate of up to 2 megasamples per second. A standard DVD drive has a motor to rotate the disc, an optical system with a laser (λ = 650 nm), and a servo focus/tracking system to centre and focus the beam on the spiral track. For the acquisition of data stored on the track, the laser scans the entire disc surface and reads the reflected intensity. Our device takes advantage of these components in two ways. First, the DVD drive is used as a centrifuge, which controls the spinning rate of the disc. Second, the drive is used as a detector, which captures the signal variation during surface scanning due to the presence of biochemical interactions (solid products).The performances of the optical disc drive were controlled by a customized software, written in Visual C++, running on a laptop connected to it through a universal serial bus (USB 2.0) interface. During the disk scanning, only the signals coming from selected areas are processed for digitization and stored in the computer (5 MB size file). The signals for each track and microarray are deconvoluted into an image. The image analysis of the microarray was also performed by the software.
Samples and extraction of genomic DNA
The certified reference materials (CRMs) were purchased from the Institute for Reference Material and Measurements (Belgium). Food products were bought from local stores. For genomic DNA extraction, aliquots of 5–20 g of the homogenized sample were extracted using a GMO-extraction kit based on column purification according to the manufacturers' instructions (Applied Biosystems, Spain). The extracted DNA was quantified by spectrophotometry (NanoDrop 2000/2000c, Thermo Scientific Inc., USA), and stored at −20 °C until analysis.Amplification and development
Each sample was analyzed in quintuplicate, including positive and negative amplification controls. Solid phase RPA was performed on the disc (10 reactions per disc, see ESI†). Reaction mixtures (6 μL) contained 480 nM of each 5′-digoxigenin labelled primer (Table SI.1†), 5 ng of genomic DNA, 14 mM of Mg-acetate, and 1× rehydration buffer. Denhardt's reagent (2.5×, Life Technologies, Spain) was also added to reduce the non-specific background. Subsequently, the mixtures were dispensed with a micropipette onto the corresponding microfluidic structure through the inlet holes. The outer hole was sealed with PCR sealer tape (Corning, USA). Then, the disc was inserted into the DVD player and a slow spin was performed (<600 rpm for 10 s) to lead the reaction mixture to Chamber 1. After a proper reconstitution and mixing of the reagents, the spin was increased to 1000 rpm to fully move the sample into Chamber 2, covering the pre-printed array. Subsequently, the disc was introduced into a container (standard DVD plastic box) in a water-saturated atmosphere, and the solid-phase amplification reactions were carried out at 37 °C for 40 min in an oven (model UF30, Memmert, Germany). After removing the sealer tape, the chamber was emptied by spinning at 1000 rpm for 10 s. The dispensation of solutions and reagents were carried out through the inlet with the movement of solutions controlled by centrifugation as described above. The array was washed by dispensation of 0.1× washing solution (SSC, 1× saline sodium citrate, containing 150 mM of NaCl and 15 mM of sodium citrate, pH 7) and water through the inlet holes (6 μL). The chamber was emptied by spinning at 1000 rpm for 10 s. The detection was carried out using a mixture of anti-digoxigenin antibody produced in sheep (1/4000) and anti-sheep conjugated with horseradish peroxidase (1/500) in PBS-T (phosphate buffered saline and 0.05% (v/v) Tween 20, pH 7.4) (6 μL). The developing reagent was 3,3′,5,5′-tetramethylbenzidine (TMB) (6 μL). The array was washed with PBS-T plus deionised water as described above. Finally, the fluidic layer was removed and the disc was inserted into the DVD player.DVD surface scanning
Because the inner structure of the optical disc remained unaltered, the microarrays on the surface were correctly read. The disc was scanned at a rotation speed of 4× ≡ 13.46 m s−1 and the signal was acquired at 26 dB gain and 1700 M samples per s with a reading time lower than 10 min. Then, 10 microarray images were created (tagged image file format, grey-scale with 16 bit-colour depth, scale 0–65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 535). Optical intensity signals of each spot, corresponding to the amount of reaction product, were quantified. In the absence of a solid biorecognition product, the reflection properties of the disc surface were unchanged and the beam intensity collected by the DVD drive was at the maximum, corresponding to the background signal of the image. However, when the laser hit the deposit of TMB product, the intensity of the laser beam that reached the photodiode decreased, corresponding to a signal for the microarray spot. In this configuration, the mean intensity of each spot was 1963 pixels (spot diameter = 500 μM). After subtracting the local background, any spot displaying a signal-to-noise ratio higher than three was considered to be positive. The reading and image processing (i.e., feature gridding, addressing, segmentation, and quality assurance) was automatically performed in less than 15 min by the disc. Used discs were discarded following the same laboratory safety guidelines as those for ELISA plates.
535). Optical intensity signals of each spot, corresponding to the amount of reaction product, were quantified. In the absence of a solid biorecognition product, the reflection properties of the disc surface were unchanged and the beam intensity collected by the DVD drive was at the maximum, corresponding to the background signal of the image. However, when the laser hit the deposit of TMB product, the intensity of the laser beam that reached the photodiode decreased, corresponding to a signal for the microarray spot. In this configuration, the mean intensity of each spot was 1963 pixels (spot diameter = 500 μM). After subtracting the local background, any spot displaying a signal-to-noise ratio higher than three was considered to be positive. The reading and image processing (i.e., feature gridding, addressing, segmentation, and quality assurance) was automatically performed in less than 15 min by the disc. Used discs were discarded following the same laboratory safety guidelines as those for ELISA plates.
Complementary measurements
Digital images of the fluidic process, shown in Fig. 1B–E, were obtained using a custom visualization stand composed of a motor (EC-Max 40 Series DC) and controller (ESCON 5/50) supplied by Maxon (USA) and a CMOS camera (uEYE camera model UI-3360CP) supplied by IDS (Germany).Amplification products recovered from the reaction chamber were checked by electrophoresis on a 3% (w/v) agarose gel at 110 V and room temperature. Gels were stained for 30 min with 0.5× TBE buffer (Tris/Borate/EDTA) containing the fluorophore RealSafe (Real Laboratories, Spain) at 0.01% (v/v), and bands were visualized with a UV transilluminator. Product size was determined by comparison with a 50 bp ladder (Fermentas, Lithuania).
Results and discussion
Microfluidic DVD characterization
A simple system composed of a reaction/detection chamber (microarray) connected by a channel to a loading chamber for the dispensation of reagents was designed (Fig. 1). Fluid propulsion on the DVD disc platform was achieved through controlling the centrifugally induced pressure on the fluid as the disc spins.21 The flow rate was dependent mainly on the rotational speed of the disc, the location of the fluid, the geometry of the fluidic channels, and the specific fluidic properties (namely, density, viscosity, and surface energy). Through the utilization of combinations of different channel geometries and spin speeds, precise flow rates ranging from nanoliters to millilitres per second were modulated. The fluidic valves were designed for use in the 0–1500 rpm range to allow for easy automation within a standard DVD drive.Process monitoring was performed with a custom visualization stand for the optimization of the process. Regarding the reagent loading, the images showed that the fluidic samples were perfectly introduced into the loading chamber (Chamber 1 of Fig. 1C) and held in place via a capillary valve. A critical challenge studied in the development of an integrated system was the storage of amplification reagents and their release. The microfluidic structure included a hydrophobic valve to allow the rehydration of reagents after sample loading, and the active life of the in-disc reagents was verified for at least two weeks.
After the initial set of tests, a serpentine mixing channel was designed to ensure the complete mixing of the lyophilized reagents with the inserted sample before they reached the reaction chamber. The selected dimensions were a width of 600 μm and a total linear length of 38 mm containing six elbows. With this configuration, the capillary valve was open when the disc was spun at 2000 rpm for 20 seconds, and the sample was fully transferred into the reaction chamber (Chamber 2 of Fig. 1D and E). The entire reaction chamber was completely covered with solutions. No formation of bubbles or other artefacts were observed.
In addition, the microfluidic structure was designed for washing the reaction chamber by controlling the disc rotation. To characterize the optimum washing speeds, the flow rate through the structures was modelled using the governing equation [Kellogg] as follows:
![[r with combining macron]](https://www.rsc.org/images/entities/i_char_0072_0304.gif) is the average distance of the liquid element from the CD centre, Δr is the radial extent of the fluid, μ is the fluid viscosity, and L is the length of the liquid in the channel. The hydraulic diameter of the channel, Dh is defined as 4A/P, where A is the cross-sectional area of a rectangular channel and P is the wetted perimeter of the channel. In repeated trials using a 600 μm serpentine channel, the washing liquid was fully transferred in 6 seconds at 500 rpm, corresponding to a flow rate of 1 μL per second. This result closely matches the calculated theoretical flow rates when the channel design specifications are known.
is the average distance of the liquid element from the CD centre, Δr is the radial extent of the fluid, μ is the fluid viscosity, and L is the length of the liquid in the channel. The hydraulic diameter of the channel, Dh is defined as 4A/P, where A is the cross-sectional area of a rectangular channel and P is the wetted perimeter of the channel. In repeated trials using a 600 μm serpentine channel, the washing liquid was fully transferred in 6 seconds at 500 rpm, corresponding to a flow rate of 1 μL per second. This result closely matches the calculated theoretical flow rates when the channel design specifications are known.
Optimization of the solid-phase RPA format
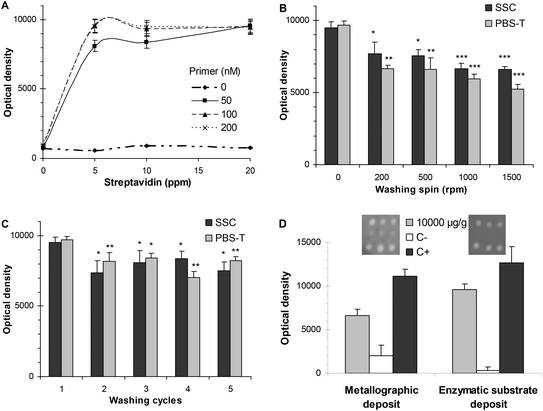
The isothermal solid-phase amplification was addressed according to the assay that was recently published based on a heminested mechanism.15 It consisted of immobilising primers, following a microarray layout, on the support of the reaction chamber. Other amplification components in the liquid phase were dispensed from the loading chamber. After the reaction chamber was filled, the amplification was produced in the static mode, i.e. rotation was not carried out during the reaction. As unbound primers were added to the reaction mixture, the amplification was produced in both the phases (in liquid and on the surface). Then, centrifugal microfluidics was just used for the semi-automated reconstitution of lyophilized reagents, for mixing them with samples and their dispensation, minimizing the contamination and increasing the reproducibility of assays. Nevertheless, merging DNA assays on microfluidic discs required the selection of robust chemistry that is able to resist the solution flow. Three processes—sensitive to flow action—were studied: (i) immobilization of primers, (ii) washing protocols, and (iii) developing reaction.In regards to the anchoring of primer, an indirect adsorption mode was chosen based on streptavidin/biotin recognition. The main advantage was the simplicity of the process because the microarraying of biotinylated primers and streptavidin could be directly performed on bulk discs without previous surface treatment of DVD or blocking steps. Under these working conditions, immobilized primers were resistant to a flow up to 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 nL s−1 (at 2000 rpm). Moreover, the amplification process on solid supports depended on the immobilization density of the primer. To that end, coating conditions were optimized by varying the streptavidin concentration from 5 to 20 ppm and the primer concentration from 50 to 200 nM. The highest signal was obtained for a primer concentration of 100 nM (0.06 fmol mm−2) (Fig. 2A).
200 nL s−1 (at 2000 rpm). Moreover, the amplification process on solid supports depended on the immobilization density of the primer. To that end, coating conditions were optimized by varying the streptavidin concentration from 5 to 20 ppm and the primer concentration from 50 to 200 nM. The highest signal was obtained for a primer concentration of 100 nM (0.06 fmol mm−2) (Fig. 2A).
The set-up of post-amplification steps was important for the integration of the assay in the microfluidic device. Compared to PCR mixtures, an RPA mixture is a highly viscous solution with a higher number of components, making the washing protocol more crucial. After the solid-phase amplification, the rest of the reaction components must be effectively removed without the release of the products that are immobilized on the disc surface of the reaction chamber. The composition of washing solutions was optimized in conventional DVDs—without microfluidic structures—by controlling parameters such as pH, ionic strength and astringency. SSC buffer and PBS-T buffer were selected for washing after the amplification process and after the incubation of antibodies, respectively. In the microfluidic discs, two protocols were assayed for both the post-amplification and the post-antibody incubation steps. In the static protocol, referred as 0 rpm, the washing method was based on completely filling the array chamber, incubation, and fast removal of the liquid. In the in-flow washing method, the washing buffers were passed continuously through the array (disc spin 0–1500 rpm). Because the in-flow washing protocol significantly decreased the signal, the static protocol was chosen (Fig. 2B). Then, the washing cycles (1–5) were studied for both the washings (namely, post-amplification and post-antibody incubation). Fig. 2C shows that a higher number of washing cycles produced a significant decrease in the signal due to the release of the product.
Two signal enhancement reactions were compared for developing the digoxigenin-labelled products that were immobilized in the reaction chamber. The first was a metallographic reaction based on the dispensation of anti-digoxigenin antibody produced in a rabbit (1/7500 dilution) and anti-rabbit antibody conjugated with gold (1/100 dilution) using silver as the developing reagent. The second approach consisted of an enzymatic reaction that dispensed anti-digoxigenin antibody produced in sheep (1/4000 dilution) and anti-sheep conjugated to horseradish peroxidase (1/500 dilution), followed by the addition of TMB as the developing reagent.
Both reaction sequences (namely, metallographic and enzymatic approaches) produced a detectable precipitate that modified the laser intensity of the DVD drive (λ = 650 nm) during the disc scanning (Fig. 2D).
However, the metallographic reaction provided high and variable background signals, which resulted in low and irreproducible signal/noise ratios and false-positives, which increased the number of washing cycles as well. The unusually high background signal observed during metallographic developing may be due to an incomplete washing of components present in the RPA mix (such as carbowax and proteins) or developing reagents, i.e. antibodies, which could enhance the reduction of silver, leading to nonspecific depositions. Because the enhancement reaction using TMB was less sensitive to the rest of the RPA components (absence of nonspecific signal), the enzymatic reaction was selected.
Analytical performances for GMO detection
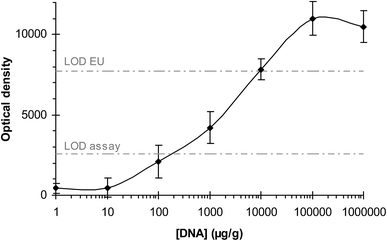
The method was applied for the screening of GMOs including screening, plant-specific, and construction-specific elements. The increasing production of transgenic crops and the concern regarding the safety of derived foods has led to the extensive monitoring of foodstuff that could contain GMOs. Therefore, the development of low-cost, reliable, rapid analytical methods for their detection and quantification through the entire production chain is of great importance.The sensitivities of the assays were determined in two ways: by analyzing serially diluted genomic DNA (10-fold dilutions) and by analyzing samples with different concentrations of transgenic ingredient. For the first approach, the concentration of DNA dispensed into the inlet chamber was varied from 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 μg g−1 (10%) to 0 μg g−1 (0%). An excellent correlation between the concentration of the transgenic template and the optical intensity measured by the DVD detector was found (R2 = 0.976) (Fig. 3).
000 μg g−1 (10%) to 0 μg g−1 (0%). An excellent correlation between the concentration of the transgenic template and the optical intensity measured by the DVD detector was found (R2 = 0.976) (Fig. 3).
For the second approach, the sensitivity of the method was assessed by simultaneously determining samples with decreasing concentrations of transgenic foods in relation to non-GMOs foods from 0% (w/w) to 10% (w/w). A t-test revealed that there was no significant difference between the slopes of the calibration curves (p-value = 0.806 > 0.05).
The detection limits (LODs) were calculated as the lowest amount of DNA that is able to produce a signal that could be distinguished from the blanks (NTC: control solution without template or food without transgenic ingredient). All the target genetic elements were detected at a concentration of 110–460 μg g−1 (0.011–0.046%).
The estimation of the sensitivity of the method in terms of copy numbers can be made theoretically as described in ref. 20. Taking into account the genome size of the plants and considering them in their haploid form, 100 ng of DNA would contain 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 630 copies of the genome for the maize and 88
630 copies of the genome for the maize and 88![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 496 copies for the soybean. The limit of detection of 0.01% GMO in that amount of genomic DNA (100 ng) would then correspond to about 3.7–8.8 haploid copies of target sequences for the different plant species. These results are similar to or better than others obtained by RT-PCR (1–16 copies)22 or by a PCR-microarray (37–88 copies).20 Thus, the system can reliably comply with the legal requirement of 0.9% limit of detection set by the EU.
496 copies for the soybean. The limit of detection of 0.01% GMO in that amount of genomic DNA (100 ng) would then correspond to about 3.7–8.8 haploid copies of target sequences for the different plant species. These results are similar to or better than others obtained by RT-PCR (1–16 copies)22 or by a PCR-microarray (37–88 copies).20 Thus, the system can reliably comply with the legal requirement of 0.9% limit of detection set by the EU.
Assay reproducibility, expressed as relative standard deviation (RSD), was determined from the optical density of spots from samples analyzed in triplicate, each in three DVDs. The intra-day RSD varied from 3.0% to 6.7%, whereas the inter-day RSD varied from 5.4% to 11.4%.
Detection of GMOs in certified reference material and food samples
The capabilities of the method were evaluated by detecting GMOs in 7 certified reference materials: maize Bt176 (ERM-BF411), maize Bt11 (ERM-BF412b and ERM-BF412f), maize GA21 (ERM-BF414b), maize MON810 (ERM-BF413ck), maize Bt176 (ERM-BF411) and Roundup ReadyTM Soya (ERM-BF410dk). All the elements of the different GMOs were detected, and they corresponded to the expected patterns (Table 1). The four lines of transgenic maize provided positive responses for the maize-specific gene (namely, adh1 gene) and negative for the rest of the taxon probes. The transgenic line of soybean (RRS) provided positive responses for the soybean-specific gene (Le1 gene) and negative for the rest of the taxon probes. Screening elements and construct-specific elements were correctly detected in all the cases and no cross-contamination between adjacent chambers was observed (n = 50). A sample was considered positive when the optical response was higher than the cut-off value (namely, an optical density of 2550).| Sample | Certified/declared GMO content | DVD detection | |||||
|---|---|---|---|---|---|---|---|
| Taxon | Screening | Construction | |||||
| Le1 | adh1 | LAT52 | p35S | tNOS | Bt-11 | ||
| CRM Bt-176 maize | 2.00% ± 0.11% | − | + | − | + | − | − |
| CRM Bt-11 maize | 0.98% ± 0.29% | − | + | − | + | + | + |
| CRM Bt-11 maize | 4.89% ± 0.21% | − | + | − | + | + | + |
| CRM GA21 maize | 0.10% ± 0.08% | − | + | − | − | + | − |
| CRM MON810 maize | 5.00% ± 0.11% | − | + | − | + | − | − |
| CRM RRS soya | 10.00% ± 0.10% | + | − | − | + | + | − |
| Tomato NahG | Presence | − | − | + | ++ | ++ | − |
| Feed corn | Presence | − | + | − | ++ | ++ | ++ |
| Cookies | Not declared | + | + | − | + | − | − |
| Ketchup | Not declared | − | + | + | − | − | − |
| Soy sauce | Not declared | + | − | − | − | − | − |
| Soy sauce + RRS | 0.5% | + | − | − | + | + | − |
| Sweet corn | Not declared | − | + | − | − | − | − |
| Sweet corn + MON810 | 0.1% | − | + | − | + | − | − |
| Cereal baby food | Not declared | − | + | − | − | − | − |
| Noodles | Not declared | − | − | − | − | − | − |
The method was also applied for the detection of food samples. Fig. 4 shows an example of the optical signals registered by the DVD drive. As can be seen, samples containing GMO ingredients were detected because positive responses were observed in the corresponding spots. Table 1 proves that positive results were observed in all the cases for the analytes declared, even at trace levels, or in spiked samples. Negative results were found in most of the samples declared to be analyte-free. The only exception was for cookie samples, in which despite not having any declared GMOs, positive results for p35S were obtained. This can be explained because its concentration might be lower than 0.9% (EU regulation).
The reliable and sensitive results achieved indicate that the proposed method is useful for the detection GMO in routine food-safety monitoring.
Conclusions
Screening protocols require analytical platforms with properties such as high working capacity, sensitivity, and reliability. This study is aligned with recent investigations that are focused on merging optical discs and centrifugal microfluidics to create a new step towards increased automatization, reduced sample consumption and low cost diagnostics. The proposed system integrates microfluidic chambers on digital versatile discs (DVDs) to perform an isothermal DNA solid-phase amplification in a microarray format. The novelty is that microfluidics is in the bottom layer of the DVD and the measurement is performed by reflection using a standard DVD player (which has small dimensions, light weight, and connectability to internet or telephone network device). Developed discs, due to their properties such as high mechanical resistance, good thermal stability and hydrophobicity, have demonstrated to be an excellent option as a bioanalytical platform. The prospective costs of the system (<2€ per disc and <500€ per reader) are below that of the state-of-the-art equipment, i.e. qPCr plates and fluorescence-thermocyclers. In addition, the method is easy to operate by locally trained staff and requires inexpensive and unspecific equipment (namely, extraction columns, pipettes, oven, DVD drive, and laptop). The properties of the proposed system make it suitable to be applied in a wide-range of ambits such as low resource settings, satellite/decentralized laboratories and production plants.As proof-of-concept, the device was applied in the detection of GMOs because their reliable identification is an important issue as their labelling is legally regulated. Screening methods are especially needed due to the high number of samples and genes to be controlled. This low-cost technology for semi-quantitative analyses has shown excellent analytical performances (namely, selectivity, sensitivity, reproducibility, and high throughput). The integration of the amplification and hybridisation steps into a one-pot reaction allows the processing of the samples in less than 90 min, reducing manipulation, reagent consumption, and risks of cross-contamination. Despite the simplicity of the approach, the results demonstrate that this screening assay can be applied without compromising analytical performance and that it suits routine genomic analysis.
Acknowledgements
This research has been funded through the projects FP7 Digital sequencing (European Commission), PAID-00-12 (UPV, GVA), FEDER PrometeoII/2014/040 (GVA) and CTQ/2013/45875 R (MINECO). The Spanish Ministry of Education and Science provided S.S.F. with a grant for her PhD studies.References
- S. Haeberle, D. Mark, F. von Stetten and R. Zengerle, Microsystems and Nanotechnology, Springer Berlin Heidelberg, 2012, pp. 853–895 Search PubMed.
- D. Nolte, Rev. Sci. Instrum., 2009, 80, 101101 CrossRef PubMed.
- M. Vázquez, D. Brabazon, F. Shang, J. O. Omamogho, J. D. Glennon and B. Paull, Trends Anal. Chem., 2011, 30, 1575–1586 CrossRef.
- J. Ducree, S. Haeberle, S. Lutz, S. Pausch, F. von Stetten and R. Zengerle, J. Micromech. Microeng., 2007, 17, S103–S115 CrossRef CAS.
- E. Roy, G. Stewart, M. Mounier, L. Malic, R. Peytavi, L. Clime, M. Madou, M. Bossinot, M. G. Bergeron and T. Veres, Lab Chip, 2015, 15, 406–416 RSC.
- S. Lutz, P. Weber, M. Focke, B. Faltin, J. Hoffmann, C. Müller, D. Mark, G. Roth, P. Munday, N. Armes, O. Piepenburg, R. Zengerle and F. von Stetten, Lab Chip, 2010, 10, 887–893 RSC.
- Y. Hua-Zhong, L. Yunchao and L. M. L. Ou, Acc. Chem. Res., 2013, 46, 258–268 CrossRef PubMed.
- S. Morais, L. A. Tortajada-Genaro, T. Arnandis-Chover, R. Puchades and A. Maquieira, Anal. Chem., 2009, 81, 5646–5654 CrossRef CAS PubMed.
- L. A. Tortajada-Genaro, S. Santiago-Felipe, S. Morais, J. A. Gabaldón, R. Puchades and A. Maquieira, J. Agric. Food Chem., 2012, 60, 36–43 CrossRef CAS PubMed.
- T. Arnandis-Chover, S. Morais, M. A. González-Martínez, R. Puchades and A. Maquieira, Biosens. Bioelectron., 2014, 51, 109–114 CrossRef CAS PubMed.
- X. Y. Peng, P. C. H. Li, H. Yu, M. Parameswaran and W. L. Chou, Sens. Actuators, B, 2007, 128, 64–69 CrossRef CAS.
- C. Li, X. Dong, J. Qin and B. Lin, Anal. Chim. Acta, 2009, 640, 93–99 CrossRef CAS PubMed.
- G. Jia, K. Ma, J. Kim, J. V. Zoval, R. Peytavi, M. G. Bergeron and M. J. Madou, Sens. Actuators, B, 2006, 114, 173–181 CrossRef CAS.
- J. Rupp, M. Schmidt, S. Münch, M. Cavalar, U. Steller, J. Steigert, M. Stumber, C. Dorrer, P. Rothacher, R. Zengerle and M. Daub, Lab Chip, 2012, 12, 1384–1388 RSC.
- S. Santiago-Felipe, L. A. Tortajada-Genaro, S. Morais, R. Puchades and A. Maquieira, Sens. Actuators, B, 2014, 204, 273–281 CrossRef CAS.
- H. Ramachandraiah, M. Amasia, J. Cole, P. Sheard, S. Pickhaver, C. Walker, V. Wirta, P. Lexow, R. Lione and A. Russom, Lab Chip, 2013, 13, 1578–1585 RSC.
- D. S. Elenis, D. P. Kalogianni, K. Glynou, P. C. Ioannou and T. K. Christopoulos, Anal. Bioanal. Chem., 2008, 392, 347–354 CrossRef CAS PubMed.
- M. Querci, M. van den Bulcke, J. Žel, G. van den Eede and H. Broll, Anal. Bioanal. Chem., 2010, 396, 1991–2002 CrossRef CAS PubMed.
- N. Marmiroli, E. Maestri, M. Gullì, A. Malcevschi, C. Peano, R. Bordoni and G. De Bellis, Anal. Bioanal. Chem., 2008, 392, 369–384 CrossRef CAS PubMed.
- S. Hamels, T. Glouden, K. Gillard, M. Mazzara, F. Debode, N. Foti, M. Sneyers, T. Esteve Nuez, M. Pla, G. Berben, W. Moens, Y. Bertheau, C. Audéon, G. Van den Eede and J. Remacle, Eur. Food Res. Technol., 2009, 228, 531–541 CrossRef CAS.
- R. Gorkin, J. Park, J. Siegrist, M. Amasia, B. S. Lee, J. M. Park, J. Kim, H. Kim, M. Madou and Y. K. Cho, Lab Chip, 2010, 10, 1758–1773 RSC.
- G. Cottenet, C. Blancpain, V. Sonnard and P. F. Chuah, Anal. Bioanal. Chem., 2013, 405, 6831–6844 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra02778k |
| This journal is © The Royal Society of Chemistry 2015 |