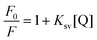Design, synthesis and biological evaluation of small molecule-based PET radioligands for the 5-hydroxytryptamine 7 receptor†
A. K. Tiwariab,
J. Yuia,
Poojab,
S. Aggarwalb,
T. Yamasakia,
L. Xiea,
N. Chadhab,
Y. Zhanga,
M. Fujinagaa,
Y. Shimodaa,
K. Kumataa,
A. K. Mishra*b,
M. Ogawaac and
M.-R. Zhang*a
aMolecular Imaging Centre, National Institute of Radiological Sciences, Inage-ku, Chiba 263-8555, Japan. E-mail: zhang@nirs.go.jp
bDivision of Cyclotron and Radiopharmaceutical Sciences, Institute of Nuclear Medicine and Allied Sciences, Brig. S. K. Mazumdar Road, Delhi-110054, India. E-mail: akmishra@inmas.drdo.in
cSHI Accelerator Service Co. Ltd, Shinagawa-ku, Tokyo 141-8686, Japan
First published on 10th February 2015
Abstract
The 5-HT7 receptor is a recently cloned G-protein-coupled receptor (GPCR) that is important in regulating sleep, depression, and circadian rhythms. However, the potential pathophysiological roles of 5-HT7 have not been fully elucidated, and no 5-HT7 positron emission tomography (PET) radioligands are available, thus limiting imaging studies of this receptor in humans. Here, we present the radiosynthesis and biological evaluation of 5-(4-([11C]methoxyphenyl)-1-methyl-4-nitro-1H-imidazole ([11C]1) as a new PET ligand for 5-HT7. Three-dimensional pharmacophore evaluation and docking studies confirmed its high affinity for 5-HT7, and in vitro binding assays showed that the binding affinity was 16.8 ± 0.9 nM. The specific activity was found to be 48 ± 29 GBq μmol−1 for [11C]1 in a synthetic time of 26 ± 3 min (n = 8), having 38 ± 7% radiochemical yield (decay-corrected) based on [11C]CO2. Ligand interactions with human serum albumin were studied by fluorescence quenching to obtain a Stern–Volmer plot, which showed a binding constant of 1.15 × 104 M−1. Whole-body biodistribution patterns were evaluated in normal mice by 1 h dynamic PET imaging; this analysis showed rapid clearance of radioactivity from the main peripheral organs, with the exception of the liver. Preliminary PET studies in rat brains showed rapid accumulation of radioactivity in the brain. The regional radioactivity reached a maximum within 0–2 min after the radioligand injection and then decreased rapidly, resulting in minimal radiation burden in the brain during the scan. In summary, this specific biaryl system has shown potential as a 5-HT7 ligand and further optimization and longitudinal studies may yield the first small molecule-based PET ligand for 5-HT7 in clinical settings.
Introduction
Serotonin (5-hydroxytryptamine, 5-HT) is a neurotransmitter that mediates its effects on the central and peripheral nervous systems via interaction with 5-HT receptors.1 The serotonin 5-HT7 receptor is the most recently identified serotonin receptor subtype.2 This seven-transmembrane domain, G-protein-coupled receptor (GPCR) was cloned in 1993 by three different groups and was found to be expressed in the central nervous system (CNS), with low expression in the periphery.3,4 Additionally, 5-HT7 was found to be positively coupled to an adenylate cyclase second messenger system. Distribution of 5-HT7 receptors has been evaluated by in situ hybridization, which showed predominant abundance in cerebral cortex, hippocampus, thalamus, hypothalamus, and amygdala of rats and guinea pigs.5 Therefore it is relevant to design a radioligand that is sensitive to changes in cerebral levels of endogenous 5-HT.6Its widespread distribution in the CNS is correlated with important functional roles in thermoregulation, circadian rhythm, endocrine regulation, sleeping, schizophrenia, depression, and memory.7,8 Therefore, in vivo imaging of the 5-HT7 receptor in the human brain is needed to assess the direct involvement of the receptor in neuropsychiatric diseases and to facilitate the development of possible therapies. Molecular imaging techniques with positron emission tomography (PET) have provided opportunities for in vivo observation in both animal models and humans; however, no radioligands have been identified to study the 5-HT7 receptor at the clinical level.
To date, many researchers have attempted to develop PET ligands for 5-HT7, including [11C]DR4446![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9 and [18F]2FP3
9 and [18F]2FP3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 derived from SB-269970 (Fig. 1), as well as many more ligands.11–13 Unfortunately, none of these ligands has shown applicability at the clinical level in humans. With recent advancements in computer-assisted drug design methods, various groups have attempted to develop useful PET ligands for 5-HT7 in the human brain. The most successful approach to date has been reported with the synthesis of a group of oxindole compounds, including [11C]Cimbi-712 (3-{4-[4-(4-[11C]methylphenyl) piperazine-1-yl]butyl}-1,3-dihydro-2H-indol-2-one) and [11C]Cimbi-717 (3-{4-[4-(3-[11C]methoxyphenyl)piperazine-1-yl]butyl}-1,3-dihydro-2H-indol-2-one).14
10 derived from SB-269970 (Fig. 1), as well as many more ligands.11–13 Unfortunately, none of these ligands has shown applicability at the clinical level in humans. With recent advancements in computer-assisted drug design methods, various groups have attempted to develop useful PET ligands for 5-HT7 in the human brain. The most successful approach to date has been reported with the synthesis of a group of oxindole compounds, including [11C]Cimbi-712 (3-{4-[4-(4-[11C]methylphenyl) piperazine-1-yl]butyl}-1,3-dihydro-2H-indol-2-one) and [11C]Cimbi-717 (3-{4-[4-(3-[11C]methoxyphenyl)piperazine-1-yl]butyl}-1,3-dihydro-2H-indol-2-one).14
In continuation of our work on the design of 5-HT7 ligands, we identified a small molecule as a lead pharmacophore for the development of new PET agents (Fig. 2). We first attempted to develop the 1-(2-diphenyl)piperazine derivative ([11C]PM20),15 which acts as a potent and selective agonist of the 5-HT7 receptor and an active metabolite of the synthetic 5-HT7 agonists LP-12 [4-(2-diphenyl)-N-(1,2,3,4-tetrahydronaphthalen-1-yl)-1-piperazinehexanamide] and LP-211 [N-(4-cyanophenylmethyl)-4-(2-diphenyl)-1-piperazinehexanamide].16 Evaluation of [11C]PM20 showed that a small molecule having a biaryl system may provide a promising lead compound for development of PET imaging agents. We further looked on other known biaryl systems AS19 and E55888, in which the biaryl system contained a substituted azole (Fig. 2).
Based on the above assumption, we designed a new biaryl derivative 5-(4-methoxyphenyl)-1-methyl-4-nitro-1H-imidazole (1), which has substituted azoles at one end, through receptor-ligand docking for 5-HT7. After computational studies and synthesis, compound 1 was evaluated in vitro for binding affinity, selectivity over 5-HT1A and for identification of permeability issues. After in vitro characterisation, compound 1 was labelled with 11C and evaluated in vivo as a 5-HT7 PET ligand.
Results and discussion
Chemical synthesis and radiosynthesis
Fig. 3 shows the synthetic pathway for the synthesis of [11C]1 and its precursor 2, along with unlabelled 1. The targeted compound was synthesized by Suzuki's coupling reaction between 5-chloro-1-methyl-4-nitroimidazole and 4-methoxyphenylboronic acid in the presence of Pd(PPh3)4. The desmethyl precursor 2 for radiosynthesis was synthesised by Suzuki's coupling reaction between the chloro compound and 4-(4,4,5,5-tetramethyl-1,3,2-dioxaborolan-2-yl)phenol in the presence of Pd(PPh3)4. The yields of both the reactions were in the range of 72% to 86%. Proton nuclear magnetic resonance (NMR) and high-resolution mass spectrometry (MS) confirmed the proposed stoichiometry and structure of both compounds 1 and 2. | ||
| Fig. 3 Chemical synthesis and radiosynthesis. Reagents and conditions: (a) Pd(PPh3)4, K2CO3, 1,4-dioxane/water (3/1), reflux, 4–6 h. (b) [11C]CH3I, NaOH, DMF, 70 °C, 5 min. | ||
[11C]1 was synthesized by reacting desmethyl precursor 2 with [11C]CH3I in the presence of NaOH. After the reaction, separation, and formulation, specific activity was found as 48 ± 29 GBq μmol−1 for [11C]1 in a time frame of 26 ± 3 min (n = 8) at the end of synthesis (EOS). The radiochemical yield for this reaction was found to be 38 ± 7% (decay-corrected) based on the cyclotron-produced [11C]CO2. The radiochemical purity of [11C]1 was higher than 99% and remained radiochemically stable for 1.5 h at room temperature. These analytical results were in compliance with our in-house quality control and assurance specifications.
Computational docking of receptor–ligand binding analysis
The 5-HT7 receptor has high sequence similarity with the 5-HT1A receptor (49%). This may result in similar binding pockets for the two subtypes of serotonin receptors, presenting a challenge for the development of compounds that are selective for each subtype. In our evaluation of ligand–protein interactions through docking analysis of molecule 1 with the 5-HT7 homology model (Fig. 4), we also compared its selectivity with that of 5-HT1A through the combined approach of docking and MD simulation with important interactive residues. The Gscore from the glide was obtained for molecule 1 (12.108 for 5-HT7 and −9.882 for 5-HT1A, respectively), which confirmed more appropriate and strong electrostatic interaction of ligand for 5-HT7 binding pocket. This gives a preliminary idea of selectivity of ligand for 5-HT7. This interactive score was higher for 1 than for best scored 1,2-substituted azole 4 (ESI Fig. 1†). Compared with known references (SB-269970 and E55888), compound 1 exhibited a Gscore on the same order (Table 1). The strong interaction between 1 and 5-HT7, as indicated by the Gscore, was observed due to the following interactions: the docked site with hydrogen bonding of piprazine-protonated NH with Asp113, the N of the imidazole group interacting with the side chain of Ser203, the π–π stacking of indole aromatic rings interacting with Phe289 and Phe290 (Fig. 4).| Ligand | XP Gscore |
|---|---|
| 1 | −12.1 |
| SB-269970 | −12.9 |
| E55888 | −11.3 |
| 4 | −10.2 |
In vitro evaluation
The computed lipophilicity at pH 7.4 (c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D) of [11C]1 was 2.1, and the corresponding experimental value of 2.3 (log
D) of [11C]1 was 2.1, and the corresponding experimental value of 2.3 (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D) was determined using the Shake Flask method reported in an earlier work15 (Table 2). The lipophilicity value of this ligand was sufficient to indicate permeation of the blood–brain barrier.
D) was determined using the Shake Flask method reported in an earlier work15 (Table 2). The lipophilicity value of this ligand was sufficient to indicate permeation of the blood–brain barrier.
| Compound | Binding affinity (Ki) | Lipophilicity | |
|---|---|---|---|
| Experimentally measured | Calculated | ||
| 5-HT7a | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Db Db |
c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Dc Dc |
|
a Binding affinities represent the mean ± standard error of the mean (SE) of triplicate samples.b log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D values were measured in octanol–phosphate buffer (pH = 7.4) using the shake flask method (n = 3; maximum range, ±5%).c c D values were measured in octanol–phosphate buffer (pH = 7.4) using the shake flask method (n = 3; maximum range, ±5%).c c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D values were calculated using Pallas 3.4 software. D values were calculated using Pallas 3.4 software. |
|||
| 1 | 16.8 nM | 2.3 | 2.1 |
| PM20 | 2.6 nM (ref. 15) | 1.7 | 3.3 |
| DR4446 | 9.7 nM (ref. 9) | Not determined | 4.5 |
| SB-269970 | 8.3 nM (ref. 27) | — | 2.4 |
Compound 1 was biologically evaluated in competition binding experiments for native serotonin 5-HT1A (rat hippocampus) and cloned human 5-HT7 (stably expressed in HEK-293 cells) receptors are shown in Fig. 1. The Ki value of 1 was found as 16.8 ± 0.9 nM, demonstrating its selectivity over that for the 5-HT1A receptor (Ki = 152 ± 3.5 nM). Selectivity in these subtypes is important because the 5-HT7 and 5-HT1A receptors exhibit similar distributions in the CNS and pharmacological properties.
The binding constant of 1 with human serum albumin (HSA) was calculated using a Stern–Volmer plot, as shown in Fig. 5. The Stern–Volmer binding constant was 1.15 × 104 M−1 for 1. This value suggested that compound 1 had the ability to bind with HSA in order to transport 1 to a specific site in the body. This value is sufficient for further use of this compound as a drug molecule for in vivo studies.
 | ||
| Fig. 5 Emission plot for ligand–HSA fluorescence quenching and Stern–Volmer plot for determination of the binding constant. | ||
In vivo analysis
Radiometabolite study was performed up to 30 min after the injection of [11C]1, only one radioactive metabolite peak (tR = 3.4 min) was recovered from the plasma on radio-HPLC, in addition to intact [11C]1 (tR = 5.4 min). Table 3 shows the percentages of intact [11C]1 in the plasma at two time point. The fraction corresponding to intact [11C]1 in the plasma very rapidly decreased to 5.83 ± 3.65%, 30 min after the injection (Table 3).
| Time after injection (min) | Percentage (%) of [11C]1 in plasma |
|---|---|
| 10 min | 23.75 ± 3.47 |
| 30 min | 5.83 ± 3.65 |
The PET study was also performed in rat brains; this analysis showed an initial high uptake, with a peak 1.34 ± 0.02 SUV after injection of [11C]1. The subsequent rapid washout of radioactivity indicated reversible tracer kinetics of [11C]1. Washout of 50% of the compound's activity in the brain was achieved within 10–12 min, after which time the activity continued to decrease very slowly (Fig. 7). Thus, the kinetic pattern exhibited a biphasic nature, with rapid changes just after injection, followed by continued, moderate changes. Blocking studies with unlabelled 1 (1 mg kg−1) and with 5-HT7-selective ligand SB-269970 (1 mg kg−1) yielded only a minimal decrease in brain radioactivity, which was non significant. In terms of SUV the decrease was 0.1 SUV or less in the time scale of 0–60 min.
Though the in vivo results were not found appropriate, further optimization is required along with the contributory factor that the low density of the 5-HT7 receptor in normal rat brains may also affect it.17 In the case of 5-HT7, the density of the receptor on the brain is quite low in comparison to other 5-HT receptors.18 In general, in the temporal cortex, the KD of the used radioligand should be approximately 350 times higher for the 5-HT1A receptor than for the 5-HT7 receptor in order to allow only 10% of the signal arising from the radioligand binding to 5-HT1A receptors.19,20 Recently, the compounds Cimbi-712 and Cimbi-717 were shown to have binding affinities in the lower nanomolar range for 5-HT7, with inhibition constants (Ki) of 1.1 and 2.6 nM, respectively. Cimbi-712 is 2191-fold more selective for 5-HT7 over 5-HT1A (the Ki for 5-HT1A is 2410 nM), whereas Cimbi-717 is 130-fold more selective (the Ki for 5-HT1A is 261 nM). This suggests that the required ligand affinity for imaging of the 5-HT7 receptor should ideally be in the subnanomolar range, with very high selectivity.
While 5-HT7 has low sequence homology with other 5-HT receptors (except 5-HT1A), 5-HT7 ligands have shown the inability to discriminate between 5-HT1A and 5-HT6 receptors.10 Even the well-known 5-HT7 ligands [3H]mesulergine and [3H]SB-269970 do not display similar binding affinities in both human and rat brain tissues;21 therefore, more elaborate studies are required to understand the binding patterns of this receptor. Further investigations in multiple animal models having increased 5-HT7 expression19,20 may provide an explanation for this observation and facilitate optimization of biaryl skeleton for clinical use.
Conclusions
In this work, we carried out rational design of a new 11C-labelled ligand ([11C]1) for 5-HT7 as a follow-up study in our previous biaryl system ([11C]PM20). In the search for a small PET ligand, we analysed another biaryl derivative after validation through ligand–receptor docking analysis. The in vitro affinity of 1 at 5-HT7 was found to be sufficient for further evaluation. PET dynamic studies validated its distribution and initial high uptake in brain regions. Pretreatment with the 5-HT7-selective ligand SB-269970 slightly decreased the binding of radiolabelled compound; this showed its inefficiency for PET evaluation. Further optimization of biaryl system may give similar results for in vitro and in vivo analysis.Experimental
Chemicals and instrumentation
All chemicals were purchased from commercial sources. Melting points (mps) were determined by using a Thomas Hoover apparatus (USA) and were not corrected. 1H NMR and 13C NMR spectra were obtained using a Bruker Avance II 400 MHz NMR system (Switzerland), and mass analysis was carried out using an Agilent 6310 ion trap mass spectrometer (mass accuracy ±0.2 u; USA). HPLC separation and analysis were performed using a JASCO HPLC system (Japan). Effluent radioactivity was monitored using an NaI (Tl) scintillation detector system, and radioactivity was measured during synthesis and animal studies using an Aloka Curiemeter.Chemical synthesis
Radiosynthesis
Computational analysis
Modelling of the receptor was performed using Prime's Homology Modelling tool (Schrödinger, LLC, New York, NY 2012) as described in our previous work.23 The docking study was initiated by joining proteins and ligands. Pharmacophore modelling studies were performed using PHASE, version 3.4, (Schrödinger).Measurement and computation of lipophilicity
The log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D value was measured by mixing [11C]1 (radiochemical purity, 100%; about 200
D value was measured by mixing [11C]1 (radiochemical purity, 100%; about 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cpm) with n-octanol (3.0 g) and sodium–phosphate buffer (PBS 3.0 g, 0.1 M, pH 7.4) in a test tube, which was vortexed for 3 min at room temperature, followed by centrifugation at 3500 rpm for 5 min. An aliquot of 1 mL phosphate-buffered saline (PBS) and 1 mL n-octanol was removed, weighed, and counted. Samples from the remaining organic layer were removed and repartitioned until consistent log
000 cpm) with n-octanol (3.0 g) and sodium–phosphate buffer (PBS 3.0 g, 0.1 M, pH 7.4) in a test tube, which was vortexed for 3 min at room temperature, followed by centrifugation at 3500 rpm for 5 min. An aliquot of 1 mL phosphate-buffered saline (PBS) and 1 mL n-octanol was removed, weighed, and counted. Samples from the remaining organic layer were removed and repartitioned until consistent log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D values were obtained. The log
D values were obtained. The log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D value was calculated from the ratio of cpm/g of n-octanol to that of PBS and expressed as log
D value was calculated from the ratio of cpm/g of n-octanol to that of PBS and expressed as log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D = log[cpm/g (n-octanol)/cpm/g (PBS)]. All assays were performed in triplicate.
D = log[cpm/g (n-octanol)/cpm/g (PBS)]. All assays were performed in triplicate.
Fluorometric studies for HSA interactions
Fluorometric experiments were carried out on a Spectra Max M2 device (CA, USA). The concentration of HSA was fixed at 5 × 10−5 M, and the concentration of the ligand was varied from 0.01 to 1 mM for 1 in PBS (pH 7.4) at 25 °C. The fluorescence spectra were recorded at a λem of 350 nm by fixing the λex at 280 nm. The intensity at 350 nm (tryptophan) was used to calculate the binding constant (K). As the concentration of the ligand increased, the relative fluorescence intensity of the HSA–ligand complex decreased at the λmax of 350 nm, which showed effective quenching by the ligand. The fluorescence quenching was usually analysed with the well-known Stern–Volmer equation:where F0 and F denote the fluorescence intensity in the absence and presence of quencher 1, respectively; Ksv is the Stern–Volmer quenching constant; and [Q] is the concentration of quencher.
In vitro binding assays
Cells were transfected with 17 μg of pcDNA3.1(+) vector containing the target 5-HT7 DNA sequence, as per standard protocols by using transfection Reagent in Opti-MEM medium without serum. Vector-expressing cells were selected using geneticin (G418). After transfection, cells were cultured in normal growth medium. One day later, cells were detached and replated into growth medium containing geneticin (800 μg mL−1) and cultured for 20 days. Surviving cell clones were picked and propagated separately in 60 mm culture dishes in the same medium, with 800 μg mL−1 geneticin to maintain selection.
Radioligand binding assays for 5-HT1A and 5-HT7 receptors
Binding affinity of 1 for 5-HT1A and 5-HT7 receptors were evaluated as per previous literature.24,25 This was performed by displacement of [3H]-8-OH-DPAT from rat hippocampus homogenate for 5-HT1A, and [3H]-5-CT from cloned human 5-HT7 stably expressed in HEK293 cells. Frozen pellets were thawed and suspended in 50 mM Tris–HCl (pH 7.4), 4 mM MgCl2, 0.1% ascorbic Acid and 10 μM pargyline hydrochloride. Assay samples were incubated in a total volume of 1 mL at 37 °C for 20 min for 5-HT1A and in 0.2 mL in 96-well microtitre plates for 60 min for 5-HT7. The process of equilibration was terminated by rapid filtration for 5-HT1A or through unifilter plates with a 96-well cell harvester in the case of 5-HT7. For displacement studies the assay samples contained 1 nM [3H]-8-OH-DPAT for 5-HT1A and 0.6 nM [3H]-5-CT for 5-HT7. Both assays used 10 mM of 5-HT for non-specific binding determination. Each compound was tested in triplicate at five different concentrations (0.1 nm to 10 μM). The inhibition constants (Ki) were calculated from the Cheng–Prusoff equation.26 Results were expressed as means of at least three separate experiments.Animals
ddY mice and male Sprague-Dawley (SD) rats were purchased from Japan SLC and were maintained in an experimental animal rearing room kept under optimal conditions with a 12/12 h dark/light cycle. Animals were handled in accordance with the recommendations of the National Institute of Health and institutional guidelines of the National Institute of Radiological Sciences (NIRS). The Animal Ethics Committee of the NIRS approved the experiments conducted at the NIRS.PET studies and image analyses in mice and rats
A small-animal PET scanner (Siemens Medical Solutions) was used for imaging. Normal mice were anesthetised during the scan, and the body temperatures of the animals were maintained with a 40 °C water circulation system (T/Pump TP401, Gaymar Industries). Emission scans were acquired at different time intervals after injection of [11C]1 (5.18 ± 0.29 MBq/0.02–0.04 nmol) through the tail vein. All image frames were summed, and regions of interest (ROIs) were drawn over the brain, heart, liver, spleen, kidneys, stomach, and bone of each mouse in four time intervals 1–2, 3–4, 6–8, 25–30, and 55–60 min. TACs for the brain, heart, blood, kidney, lung, and liver were generated from the dynamic PET data in order to parameterise radioactivity uptake, clearance, and distribution in healthy mice.Brain regional PET studies were performed in normal rats anesthetized with 1–2% isoflurane during the scan at an optimal body temperature of 40 °C. An emission scan was immediately acquired for 60 min after injection of [11C]1 (37 MBq/0.4–0.6 nmol in 100 μL) through the tail vein. Each experiment was performed in triplicate.
Radioactivity (% ID/g) was estimated as the ratio of the regional activity concentration normalised according to the injected dose and the weight of the animal, yielding the PET-generated biodistribution patterns of various critical organs.
Data modelling for PET scans was performed using three-dimensional sinograms, which were changed into two-dimensional sinograms by Fourier rebinning. Dynamic image reconstruction was carried out by filtered back-projection using Hanning's filter with a Nyquist cut-off frequency of 0.5 cycles per pixel. PET images were analysed using ASIPro VM™ (Analysis Tools and System Setup/Diagnostics Tool; Siemens Medical Solutions).
Radiolabelled metabolites analysis
The ddY mice were intravenously injected with [11C]1 (37–54 MBq per mouse). The animals were sacrificed by cervical dislocation at 10 and 30 min (n = 3 for each point). Blood (0.7–1.0 mL) samples were centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 1 min at 4 °C to separate the plasma. The supernatant (0.5 mL) was then collected in a test tube containing MeCN (0.5 mL), and the resulting mixture was vortexed for 15 s and centrifuged at 20
000 rpm for 1 min at 4 °C to separate the plasma. The supernatant (0.5 mL) was then collected in a test tube containing MeCN (0.5 mL), and the resulting mixture was vortexed for 15 s and centrifuged at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g for 2 min for deproteinization. The resulting supernatant was collected and injected onto radio-HPLC analysis (Capcell Pack UG80 C18 column (4.6 mm i.d. × 250 mm), MeCN/H2O/Et3N, 45/55/0.01 (v/v/v), flow rate = 1.0 mL min−1).
000 g for 2 min for deproteinization. The resulting supernatant was collected and injected onto radio-HPLC analysis (Capcell Pack UG80 C18 column (4.6 mm i.d. × 250 mm), MeCN/H2O/Et3N, 45/55/0.01 (v/v/v), flow rate = 1.0 mL min−1).
Acknowledgements
The authors thank the staff of the National Institute of Radiological Sciences for their support with radiosynthesis and animal experiments. In addition, the first author would like to thank the Japan Society for the Promotion of Science for financial support.Notes and references
- D. Hoyer, J. P. Hannon and G. R. Martin, Pharmacol., Biochem. Behav., 2002, 71, 533 CrossRef CAS.
- J. A. Bard, J. Zgombick, N. Adham, P. Vaysse, T. A. Branchek and R. L. Weinshank, J. Biol. Chem., 1993, 268, 23422 CAS.
- T. W. Lovenberg, B. M. Baron, L. de Lecea, J. D. Miller, R. A. Prosser, M. A. Rea, P. E. Foye, M. Racke, A. L. Slone, B. W. Siegel, P. E. Danielson, J. G. Sutcliffe and M. G. Erlander, Neuron, 1993, 11, 449 CrossRef CAS.
- M. Ruat, E. Traiffort, R. Leurs, J. Tardivel-Lacombe, J. Diaz, J. M. Arrang and J. C. Schwartz, Proc. Natl. Acad. Sci. U. S. A., 1993, 90, 8547 CrossRef CAS.
- D. R. Thomasand and J. J. Hagan, Curr. Drug Targets: CNS Neurol. Disord., 2004, 3, 81–90 CrossRef.
- L. M. Paterson, R. J. Tyacke, D. J. Nutt and G. M. Knudsen, J. Cereb. Blood Flow Metab., 2010, 30, 1682–1706 CrossRef CAS PubMed.
- M. Filali, L. Lambas-Señas, H. Scarna and N. Haddjeri, Curr. Drug Targets, 2009, 10, 1109 CrossRef.
- P. B. Hedlund, L. Kelly, C. Mazur, T. Lovenberg, J. G. Sutcliffe and P. Bonaventure, Eur. J. Pharmacol., 2004, 487, 125 CrossRef CAS PubMed.
- M. R. Zhang, T. Haradahira, J. Maeda, T. Okauchi, T. Kida, S. Obayashi, K. Suzuki and T. Suhara, J. Labelled Compd. Radiopharm., 2002, 45, 857 CrossRef CAS.
- L. Lemoine, J. Andries, D. Le Bars, T. Billard and L. Zimmer, J. Nucl. Med., 2011, 52, 1811 CrossRef CAS PubMed.
- H. D. Hansen, E. Lacivita, P. Di Pilato, M. M. Herth, S. Lehel, A. Ettrup, V. L. Andersen, A. Dyssegaard, P. De Giorgio, R. Perrone, F. Berardi, N. A. Colabufo, M. Niso, G. M. Knudsen and M. Leopoldo, Eur. J. Med. Chem., 2014, 79, 152 CrossRef CAS PubMed.
- J. Colomb, G. Becker, E. Forcellini, S. Meyer, L. Buisson, L. Zimmer and T. Billard, Nucl. Med. Biol., 2014, 41, 330 CrossRef CAS PubMed.
- E. Lacivita, M. Niso, H. D. Hansen, P. Di Pilato, M. M. Herth, S. Lehel, A. Ettrup, L. Montenegro, R. Perrone, F. Berardi, N. A. Colabufo, M. Leopoldo and G. M. Knudsen, Bioorg. Med. Chem., 2014, 22, 1736 CrossRef CAS PubMed.
- H. D. Hansen, M. M. Herth, A. Ettrup, V. L. Andersen, S. Lehel, A. Dyssegaard, J. L. Kristensen and G. M. Knudsen, J. Nucl. Med., 2014, 55, 640 CrossRef CAS PubMed.
- Y. Shimoda, J. Yui, L. Xie, M. Fujinaga, T. Yamasaki, M. Ogawa, N. Nengaki, K. Kumata, A. Hatori, K. Kawamura and M. R. Zhang, Bioorg. Med. Chem., 2013, 21, 5316 CrossRef CAS PubMed.
- P. Di Pilato, M. Niso, W. Adriani, E. Romano, D. Travaglini, F. Berardi, N. A. Colabufo, R. Perrone, G. Laviola, E. Lacivita and M. Leopoldo, Rev. Neurosci., 2014, 25, 401 CAS.
- D. Hoyer, J. P. Hannon and G. R. Martin, Pharmacol., Biochem. Behav., 2002, 71, 533 CrossRef CAS.
- L. Zimmer and T. Billard, Rev. Neurosci., 2014, 25, 357 CAS.
- J. S. Kumar, R. V. Parsey, S. A. Kassir, V. J. Majo, M. S. Milak, J. Prabhakaran, N. R. Simpson, M. D. Underwood, J. J. Mann and V. Arango, Brain Res., 2013, 1507, 11 CrossRef CAS PubMed.
- D. R. Thomas, P. J. Atkinson, P. G. Hastie, J. C. Roberts, D. N. Middlemiss and G. W. Price, Neuropharmacology, 2002, 42, 74 CrossRef CAS.
- K. Varnas, D. R. Thomas, E. Tupala, J. Tiihonen and H. Hall, Neurosci. Lett., 2004, 367, 313 CrossRef CAS PubMed.
- A. K. Tiwari, J. Yui, M. Fujinaga, K. Kumata, Y. Shimoda, T. Yamasaki, L. Xie, A. Hatori, J. Maeda, N. Nengaki and M. R. Zhang, J. Neurochem., 2014, 129, 712 CrossRef CAS PubMed.
- A. K. Tiwari, M. Fujinaga, J. Yui, T. Yamasaki, L. Xie, K. Kumata, A. K. Mishra, Y. Shimoda, A. Hatori, B. Ji, M. Ogawa, K. Kawamura, F. Wang and M. R. Zhang, Org. Biomol. Chem., 2014, 12, 9621 CAS.
- M. H. Paluchowska, R. Bugno, B. Duszynska, E. Tatarczynska, A. Nikiforuk, T. Lenda, E. Chojnacka-Wojcik and E. Bioorg, Med. Chem., 2007, 15, 7116 CAS.
- J. R. Jasper, A. Kosaka, Z. P. To, D. J. Chang and R. M. Eglen, Br. J. Pharmacol., 1997, 122, 126 CrossRef CAS PubMed.
- Y. C. Cheng and W. H. Prussoff, Biochem. Pharmacol., 1973, 22, 3099 CrossRef CAS.
- P. J. Lovell, S. M. Bromidge and S. Dabbs, J. Med. Chem., 2000, 43, 342 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c4ra15833d |
| This journal is © The Royal Society of Chemistry 2015 |