Fluorescence probing of metal-ion-mediated hybridization of oligonucleotides†
Sharmin Taherpour* and
Tuomas Lönnberg
Department of Chemistry, University of Turku, Vatselankatu 2, FIN-20014, Turku, Finland. E-mail: shtahe@utu.fi
First published on 5th January 2015
Abstract
The sensitivity of fluorescence of pyrrolocytosine-containing oligonucleotides to changes in their secondary structure has been harnessed to monitor the hybridization of modified metal-ion-chelating oligonucleotides with their unmodified counterparts. With short double-helical oligonucleotides, quenching of fluorescence correlated well with the length and, hence, Tm of the duplex provided that the pyrrolo-dC residue was incorporated within the duplex. Furthermore, hybridization of the metal-ion-chelating oligonucleotides was greatly enhanced on addition of 1 eq. of Cu2+, as evidenced by both Tm and fluorometric measurements. As an example of a case where interpretation of a conventional UV-melting profile is challenging, application of the fluorometric method was extended to probing hybridization of short metal-ion-carrying oligonucleotides with the trinucleotide bulge motif of TAR RNA models. In this case the results were more ambiguous, presumably due to weak hybridization with the short target sequence.
Introduction
The past decade has seen the emergence of non-coding small RNA molecules, such as siRNA and miRNA, as targets for chemotherapy and diagnostics.1 These RNAs regulate gene expression post-transcriptionally and can suppress viral2,3 and oncogenic4,5 RNAs. On the other hand, a number of miRNAs have been identified as oncogenic themselves.5,6 Most types of cancer have, indeed, highly characteristic miRNA profiles that could be exploited in cancer diagnostics.7 The TAR element of HIV shares the general secondary structure of pre-miRNAs, i.e. a hairpin loop and a double-helical stem incorporating a trinucleotide bulge.8 In fact, the TAR element has been identified as a source of viral miRNAs.2,9 The bulge and loop regions of TAR RNA serve as binding sites of HIV's trans-activator of transcription (Tat) protein10 and positive transcription elongation factor complex (P-EFTb),11 respectively, making them attractive targets for anti-viral therapeutics.12The most obvious approach for targeting nucleic acid sequences is through hybridization with a complementary oligonucleotide.13 In the case of pre-miRNA or the TAR element of HIV, however, the single-stranded regions are too short for efficient hybridization at physiological temperature, rendering this strategy largely dependent on the RISC machinery to unwind the pre-miRNA to a longer single-stranded target. One way to overcome this problem would be to replace some or all of the canonical Watson–Crick base pairs with metal-ion-mediated base pairs between modified and natural nucleobases. A number of stabilizing metal-ion-mediated base pairs have, indeed, been described in the literature.14 In most cases, however, the pairing was achieved by sandwiching the metal ion between two modified or two natural nucleobases, limiting the utility of these systems in targeting short biologically relevant RNAs. In fact, besides our own previous works, we are aware of only one report of a metal-ion-mediated base pair between a modified and a natural nucleobase in an oligonucleotide environment.15
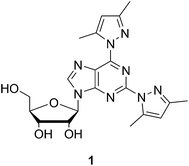
In a previous study, we have shown that the tridentate 2,6-bis(3,5-dimethylpyrazol-1-yl)purine ribonucleoside (1), when incorporated at the 3′-termini of short oligonucleotides, greatly stabilizes the duplexes formed by these oligonucleotides in the presence of Cu2+ and, to a lesser extent, Zn2+.16 While discrimination between the natural nucleobases to complete the metal-ion-mediated base pair was rather modest, fidelity of base pairing in the Watson–Crick region of the oligonucleotides was retained. In the present study we explore the potential of short oligonucleotides incorporating the chelating nucleoside 1 for selective hybridization with the bulge motif of an HIV TAR RNA model. The similarity in secondary structure of this model and pre-miRNA makes the results relevant also from the point of view of pre-miRNA recognition.
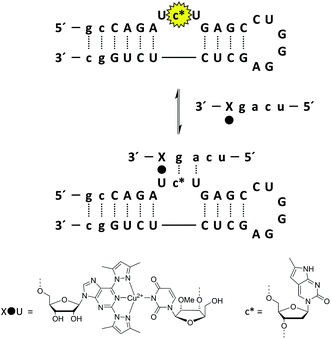
In addition to the desired hybridization between the bulge and the metal-ion-carrying oligonucleotide, the TAR RNA model contains two additional short double-helical regions, giving rise to a highly convoluted UV melting profile. Furthermore, even if a melting temperature could be obtained for the duplex between the metal-ion-carrying oligonucleotide and the TAR RNA model, the site of hybridization could not be unambiguously established. To avoid these problems, the interaction of the modified oligonucleotides with the TAR RNA models has been studied by using a fluorescence-based method. Accordingly, a pyrrolo-2′-deoxycytidine17 residue has been incorporated at various sites of the target oligonucleotide and its fluorescence emission measured in the presence and absence of the metal-ion-chelating oligonucleotide and Cu2+ (Scheme 1). The fluorescence emission of pyrrolo-dC is quenched when this nucleoside is embedded within a double helix,18 providing information on the site of hybridization.
Results and discussion
Oligonucleotide synthesis
The 15- and 9-mer target oligonucleotides ON1a, ON1b and ON2, as well as the 29-mer TAR RNA models ON3a, ON3b and ON3c, with different placement of the fluorescent pyrrolo-dC residue (Table 1), were assembled on an automated synthesizer from commercial phosphoramidite monomers. In the case of the TAR RNA models, two 2′-O-methyl ribonucleotides were inserted to both ends of the chain to increase stability of the stem region and to protect the models against decomposition by exonucleases. To ensure folding to the desired natural tertiary structure, the rest of the sequence was native RNA. In the synthesis of the 15- and 9-mer targets, in turn, 2′-O-methylated building blocks were used throughout the sequences for stability and ease of synthesis. As Cu2+ is known to interact with nucleic acids predominantly through the base moieties,19 methylation of the 2′-oxygens should not affect the coordination chemistry of the oligonucleotides. The 2′-O-methyl-RNA oligonucleotides were released from the support and deprotected by conventional ammonolysis (33% aq. NH3, 5 h at 55 °C), whereas for the RNA oligonucleotides, a mixture of ethanol and 33% aq. NH3 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, v/v, 16 h at 55 °C) was used instead.
3, v/v, 16 h at 55 °C) was used instead.
| Sequencea | |
|---|---|
| a Capital letters refer to ribonucleotide and lowercase letters to 2′-O-methylribonucleotide residues. For the structure of the pyrrolo-2′-deoxycytidine residue (c*), see Scheme 1. | |
| ON1a | 5′-aaugcagugc*cguaa-3′ |
| ON1b | 5′-aaugc*agugccguaa-3′ |
| ON1c | 5′-aaugcagugccguaa-3′ |
| ON2 | 5′-ccggc*gcgc-3′ |
| ON3a | 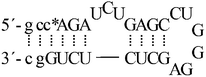 |
| ON3b | 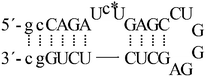 |
| ON3c | 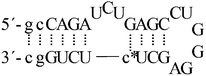 |
Assembly of the metal-ion-chelating 4-, 5-, and 6-mer oligonucleotides ON4a, ON4b, ON4c, ON5a and ON5b (Table 2), bearing the modified nucleoside 1 (denoted by the letter X in the sequences) at the 3′-terminus, was carried out on a solid support obtained by immobilizing the 3′-O-TBDMS-protected derivative of 1 on LCAA-CPG, as reported previously.16 In all cases, the couplings proceeded on this support with normal (approximately 99%) efficiency. Due to the lability of the modified nucleoside 1 to nucleophiles, a shorter ammonia treatment (33% aq. NH3, 2 h at 55 °C) was used with these oligonucleotides.
| Sequencea | |
|---|---|
| a Lowercase letters refer to 2′-O-methylribonucleotide residues. The capital letter X refers to the residue of the metal-ion-chelating nucleoside 1. | |
| ON4a | 5′-ggcX-3′ |
| ON4b | 5′-cggcX-3′ |
| ON4c | 5′-acggcX-3′ |
| ON5a | 5′-ucagX-3′ |
| ON5b | 5′-ugagX-3′ |
| ON6 | 5′-uuacggcacugcauu-3′ |
| ON7 | 5′-aaaaaaaaaaaaaaa-3′ |
| ON8 | 5′-gcgcXccgg-3′ |
In all cases (where applicable), the TBDMS protections were removed with 1.53 M triethylamine trihydrofluoride in dry DMSO (2 h at 55 °C). The crude oligonucleotides were purified by RP-HPLC and characterized by ESI-MS. The concentrations were determined UV-spectrophotometrically using molar absorptivities calculated by an implementation of the nearest-neighbors method.20
Metal-ion-promoted hybridization of single-stranded oligonucleotides
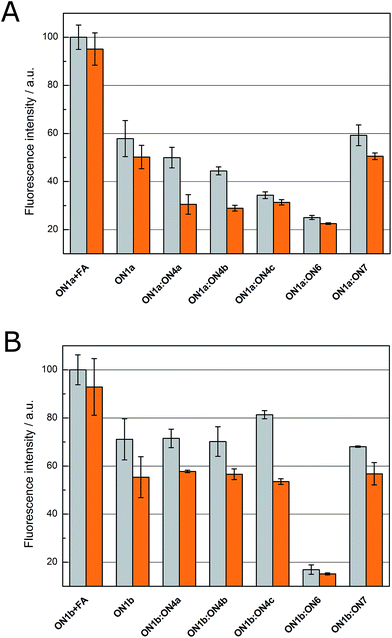
For validation on a system where conventional Tm measurements yield unambiguous results, the fluorometric studies were first carried out with the 15-mer 2′-O-Me-RNA oligonucleotides ON1a and ON1b, differing only in the placement of the fluorescent pyrrolo-dC residue. These oligonucleotides were designed to have essentially identical hybridization properties but significant quenching of the fluorescence emission was only expected when hybridization takes place at a region containing the pyrrolo-dC residue (Fig. 1). The intensity of fluorescence emission of ON1a and ON1b, as well as the duplexes formed with their short modified counterparts ON4a, ON4b and ON4c, was recorded at 460 nm (λex = 337 nm) in the presence and absence of 1 eq. of Cu2+ (Fig. 2). For a reference representing the maximal quenching of pyrrolo-dC fluorescence, ON1a and ON1b were also hybridized with the fully complementary unmodified 15-mer ON6 (Table 2). Finally, for a negative control with a non-complementary sequence, the fluorescence emission of ON1a and ON1b was measured in the presence of the A15 oligonucleotide ON7 (Table 2). The measurements were performed at room temperature in a 20 mmol L−1 cacodylate buffer at pH 7.4 and ionic strength of 0.10 mol L−1 (adjusted with NaClO4). The concentration of the oligonucleotides and Cu2+ (where applicable) was 2.0 μmol L−1. To examine to what extent the pyrrolo-dC fluorescence is affected by base-stacking or transient intramolecular base-pairing of the labeled single-stranded oligonucleotide, the emission of ON1a and ON1b was also measured in otherwise the same conditions except that formamide (40%, v/v) was added as a denaturant.Single-stranded ON1a and ON1b both exhibited strong emission at 460 nm and in both cases the emission was markedly quenched upon hybridization with ON6. This behavior is expected as in both ON1a:ON6 and ON1b:ON6, the fluorescent pyrrolo-dC residue is embedded within a relatively long double helix. Introduction of the non-complementary oligonucleotide ON7, in turn, had hardly any impact on the fluorescence of ON1a and ON1b. Under denaturing conditions (40% aq. formaldehyde), a significant increase in emission was observed with both ON1a and ON1b, indicating a degree of quenching of the pyrrolo-dC fluorescence even in single-stranded oligonucleotides, in all likelihood due to base-stacking.
In the absence of divalent metal ions, the fluorescence of the duplexes formed by ON1a with the short modified oligonucleotides ON4a, ON4b and ON4c monotonously decreased with increasing length of the short oligonucleotide, consistent with the fact that in these duplexes the pyrrolo-dC residue is part of the double-helical region. However, even with the longest duplex (ON1a:ON4c), quenching was less pronounced than with the full-length ON1a:ON6. In the case of ON1b, where the pyrrolo-dC residue lies outside of the region forming the double helix, the fluorescence was insensitive to the presence of the short modified oligonucleotides.
In the case of single-stranded ON1a and ON1b, as well as the duplexes formed by ON1b with the short modified oligonucleotides, addition of 1 eq. of Cu2+ quenched the fluorescence by 10–15%, presumably due to coordination of Cu2+ by the pyrrolocytosine base. With ON1a:ON4a, ON1a:ON4b and ON1a:ON4c, in turn, fluorescence emission dropped almost to the baseline level observed with the 15-mer double helix ON1a:ON6. Unlike in the absence of divalent metal ions, in the presence of Cu2+ the fluorescence of these short duplexes was no longer dependent on the length of the double-helix. As could be expected, the greatest effect upon introduction of Cu2+ was observed with the shortest modified oligonucleotide ON4a, in which case the emission was quenched by nearly 40%.
In addition to the well-documented effects of base-stacking, fluorescence of the pyrrolo-dC residue could also be quenched by absorption of the Cu2+-mediated base pair at the excitation wavelength used in the fluorometric measurements (337 nm). To explore this possibility, UV spectrum of the chelating nucleoside 1 was recorded at varying concentrations of CuSO4 (Fig. 3). On increasing [Cu2+], hyperchromic shifts at 250 and 330 nm and a hypochromic shift at 270 nm were observed. These shifts leveled off at [Cu2+] > 50 μM (5 eq.), consistent with coordination of Cu2+ by 1 (data presented as ESI†). The same experiment was also carried out with the ON1c:ON4c duplex, with similar results (data presented as ESI†). From these results, the absorption of the Cu2+-mediated base pair at 337 nm under conditions of the fluorometric measurements could be estimated as 0.01. Given this low value, interference of the pyrrolocytosine excitation by absorbance of the Cu2+-mediated base pair does not play a major role.
For comparison with the fluorometric data, melting temperatures of the oligonucleotide duplexes were measured under the same conditions as the fluorescence intensities (Table 3). Nearly identical results were obtained with ON1a and ON1b, as would be expected given the fact that these two oligonucleotides differ only in the location of the pyrrolo-dC residue and that this modified nucleoside closely mimics native cytidine in Watson–Crick base pairing. The latter point was further verified by measuring the melting temperature of the ON1c:ON6 duplex, an unmodified counterpart of ON1a:ON6 and ON1b:ON6. The recorded Tm, 75.5 °C, was very close to respective value of ON1a:ON6.
| Cu2+ | ON1a | ON1b | ||
|---|---|---|---|---|
| − | + | − | + | |
| ON4a | 10.8 ± 0.5 | 28.6 ± 0.4 (+17.8) | 11.1 ± 0.7 | 29 ± 3 (+17.9) |
| ON4b | 27.6 ± 1.3 | 51.5 ± 0.3 (+23.9) | 25.3 ± 0.8 | 51.9 ± 0.5 (+26.6) |
| ON4c | 42.9 ± 0.2 | 54.6 ± 1.6 (+11.7) | 41.9 ± 0.3 | 55.3 ± 0.4 (+13.4) |
| ON6 | 74.0 ± 0.3 | 74.6 ± 0.2 (+0.6) | 70.9 ± 0.3 | 71.9 ± 0.1 (+1.0) |
In line with previous results on short self-complementary oligonucleotides bearing the metal-ion-chelating nucleoside 1 at the 3′-terminus, the duplexes formed with ON4a, ON4c and, especially, ON4b, were greatly stabilized in the presence of 1 eq. of Cu2+, suggesting Cu2+-mediated base pairing between 1 and U. As previously reported,21 the 2,6-bis(3,5-dimethylpyrazol-1-yl)purine base of 1 may undergo partial hydrolysis at high temperatures in the presence of Cu2+ but the resulting 2-(3,5-dimethylpyrazol-1-yl)hypoxanthine base exhibits similar affinity in metal-ion-mediated base pairing.
In all of the cases discussed above, the fluorescent pyrrolo-dC residue has been either unpaired or engaged in canonical Watson–Crick base pairing. To study the impact of Cu2+-mediated base pairing of pyrrolocytosine on its fluorescence, the 9-mer oligonucleotide ON2 was hybridized with its metal-ion-chelating complement ON8 and the emission intensity measured under the conditions described above (Fig. 4). The duplex between ON8 and its unmodified complement placing cytosine opposite to the metal-ion-chelating 2,6-bis(3,5-dimethylpyrazol-1-yl)purine has been reported to be greatly stabilized by 1 eq. of Cu2+ and this stabilization was attributed to Cu2+-mediated base pairing between these two nucleobases.16 Given the similarity of the Watson–Crick face of cytosine and pyrrolocytosine, formation of a Cu2+-mediated base pair was expected also in the middle of the ON2:ON8 duplex.
The fluorescence of single-stranded ON2 was similar to single-stranded ON1a or ON1b and insensitive to the presence of Cu2+. The behavior of the ON2:ON8 duplex, on the other hand, was dramatically different: in the absence of Cu2+, ON2:ON8 emitted twice as strongly as single-stranded ON2. The most likely explanation for this increase in fluorescence intensity is that in the ON2:ON8 duplex, the bulky 2,6-bis(3,5-dimethylpyrazol-1-yl)purine base forces the pyrrolocytosine out of the base stack, exposing it even more than in single-stranded ON2. A similar effect has been reported for complexes of the methyl-specific endonuclease McrBC with pyrrolo-dC-modified oligonucleotides, in which case flipping-out of the cytosine base was also verified by X-ray crystallography.22 Addition of 1 eq. of Cu2+ led to quenching of the fluorescence, comparable to the level observed with ON1a:ON4a, ON1a:ON4b and ON1a:ON4c. Evidently upon formation of the Cu2+-mediated base-pair between pyrrolocytosine and 2,6-bis(3,5-dimethylpyrazol-1-yl)purine, the former becomes part of the base stack again. This interpretation receives further support from our previous studies with related duplexes: in the absence of divalent metal ions, the bulky modified residues favored an abasic site over any of the natural nucleobases in the opposite strand but in the presence of Cu2+ the situation was reversed, owing to formation of Cu2+-mediated base pairs.21
Recognition of the bulge motif of TAR RNA by short metallo-oligonucleotides
Encouraged by the capacity of even the shortest metal-ion-chelating oligonucleotide tested (the 4-mer ON4a) to hybridize with its unmodified complements with high affinity and sequence-specificity, we next set out to explore the potential of such oligonucleotides in the recognition of the bulge motif of RNA. Fluorescence of the three target models (ON3a, ON3b and ON3c), incorporating the pyrrolo-dC residue at different locations of the strand, was quantified in the presence and absence of oligonucleotides ON5a and ON5b under the same conditions as described above for the single-stranded targets (Fig. 5). In ON3b, the pyrrolo-dC residue formed a part of the trinucleotide bulge (Scheme 1), whereas in ON3a and ON3c it was embedded within the double-helical stem. In other words, only ON3b is expected to experience a marked loss of fluorescence upon hybridization with ON5a or ON5b. The short oligonucleotides ON5a and ON5b, in turn, were both complementary to the bulge region of ON3a, ON3b and ON3c but differed in their 5′-terminal sequences. Accordingly, ON5a, but not ON5b, could potentially invade the double helix next to the loop motif of the TAR RNA models.Despite the different environment surrounding the pyrrolo-dC residue in ON3a, ON3b and ON3c, similar fluorescence emissions were recorded for these three oligonucleotides in the absence of Cu2+. Contrary to what could have been expected, addition of 1 eq. of the short modified oligonucleotide ON5a decreased the fluorescence of ON3a by 29% and increased the fluorescence of ON3b by 7%. With ON3c, a 6% increase was observed. Addition of ON5b, in turn, led to quenching with all the three models (30%, 8% and 5% for ON3a, ON3b and ON3c, respectively). Clearly, the relationship between secondary structure and fluorescence is much more complicated with the TAR RNA models than with linear oligonucleotides.
Introduction of 1 eq. of Cu2+ resulted in approximately 15% quenching of fluorescence with all of the TAR models. The magnitude of quenching was comparable to that observed with single-stranded ON1a and ON1b and probably attributable to coordination of Cu2+ by the pyrrolocytosine base. Of the putative duplexes formed with the short metal-ion-chelating oligonucleotides ON5a and ON5b, only ON3b:ON5a, ON3b:ON5b and ON3c:ON5a exhibited appreciable quenching of fluorescence by Cu2+. As expected, the greatest effect was observed with ON3b:ON5a but one should be careful interpreting these results as fluorescence of ON3b:ON5a in the absence of divalent metal ions appeared anomalously high. For comparison, with ON3b:ON5b the quenching was lower than what was observed with ON3b alone.
Despite the fact that significant quenching of the fluorescence of the single-stranded oligonucleotide ON1a took place upon Cu2+-promoted hybridization with the modified 5-mer ON4b and even the 4-mer ON4a, hybridization of the 5-mer metal-ion-carrying oligonucleotides ON5a and ON5b with the bulge region of the TAR RNA models could not be unambiguously confirmed. Evidently neither ON5a nor ON5b is able to invade the double-helical stem between the bulge and the loop motifs and the three unpaired bases constituting the bulge are not enough for significant hybridization, even when one of the base pairs would be a stronger metal-ion-mediated one. Furthermore, these three bases may not be as readily available to base pairing as those of a longer single-stranded region.
Conclusions
Pyrrolocytosine was proven to be a good probe for metal-ion-mediated base-pairing in oligonucleotides. Within a double helix, the decrease of the fluorescence emission of the pyrrolo-dC correlated well with the thermal stability of the double helix. When placed opposite to a sterically demanding base, such as the 2,6-bis(3,5-dimethylpyrazol-1-yl)purine employed in the present study, pyrrolocytosine flips out of the base stack, resulting in a great increase of the fluorescence emission. Attempts to study the binding of short metal-ion-carrying oligonucleotides to the trinucleotide bulge region of pyrrolo-dC-labeled TAR RNA models was less successful, probably owing to insufficient affinity of the former to the short target sequence.Experimental section
General methods
Mass spectra were recorded on a Bruker Daltonics micrOTOF-Q ESI-MS spectrometer. For the preparation of HPLC elution buffers, freshly distilled triethylamine was used. Other chemicals were commercial products that were used as received.Oligonucleotide synthesis
The modified oligonucleotides were prepared using an ABI 3400 DNA/RNA synthesizer following previously published protocols.16,23 Accordingly, the metal-ion-chelating 4-, 5-, and 6-mer oligonucleotides ON4a, ON4b, ON4c, ON5a and ON5b were assembled on a solid support obtained by immobilizing the 3′-O-TBDMS-protected derivative of the modified nucleoside 1 on LCAA-CPG. Synthesis of the other oligonucleotides was carried out on appropriate commercial supports according to conventional phosphoramidite strategy. The oligonucleotides were released from the supports and the base and phosphate protections removed by treatment with either 33% aq. NH3 at 55 °C (2 h for ON4a, ON4b, ON4c, ON5a and ON5b, 5 h for ON1a, ON1b, ON2 and ON6) or a mixture of ethanol and 33% aq. NH3 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, v/v) at 55 °C (16 h for ON3a, ON3b and ON3c). Where applicable, the TBDMS protections were removed with 1.53 M triethylamine trihydrofluoride in dry DMSO (2 h at 55 °C). The crude oligonucleotides were purified by RP-HPLC on a Hypersil ODS C18 column (250 × 4.6 mm, 5 μm) eluting with a linear gradient (10–40% during 25 min) of acetonitrile in 0.1 mol L−1 aq. triethylammonium acetate. The identity of the purified oligonucleotides was established by ESI-MS and their concentration determined UV-spectrophotometrically using molar absorptivities calculated by an implementation of the nearest-neighbors method.20
3, v/v) at 55 °C (16 h for ON3a, ON3b and ON3c). Where applicable, the TBDMS protections were removed with 1.53 M triethylamine trihydrofluoride in dry DMSO (2 h at 55 °C). The crude oligonucleotides were purified by RP-HPLC on a Hypersil ODS C18 column (250 × 4.6 mm, 5 μm) eluting with a linear gradient (10–40% during 25 min) of acetonitrile in 0.1 mol L−1 aq. triethylammonium acetate. The identity of the purified oligonucleotides was established by ESI-MS and their concentration determined UV-spectrophotometrically using molar absorptivities calculated by an implementation of the nearest-neighbors method.20
Melting temperature studies
The UV melting curves were acquired by monitoring the temperature dependence of the absorbance at 260 nm (data collection 0.5 °C min−1) on a Perkin-Elmer Lambda 35 UV-vis spectrophotometer equipped with a Peltier temperature controller. The measurements were carried out at pH 7.4 (20 mmol L−1 cacodylate buffer) and ionic strength of 0.10 mol L−1 (adjusted with NaClO4). The concentration of the oligonucleotides and CuSO4 (where applicable) was 2.0 μmol L−1. The Tm values were determined as maximum values of the first derivatives of the melting curves.Fluorometric studies
Fluorescence emission spectra were recorded between 400 and 600 nm using a Cary Eclipse fluorescence spectrophotometer with an excitation wavelength of 337 nm. The scan rate was 120 nm min−1 with a slit of 5 nm and a PMT voltage of 600 V. The composition of the samples was the same as described above for the Tm measurements. Before fluorescence measurements all samples were quickly heated to approximately 90 °C and then allowed to slowly cool down. The measurements were carried out at room temperature.Acknowledgements
We gratefully acknowledge the financial support of the Academy of Finland.References
- Therapeutic Applications of RNAi: Methods and Protocols, ed. J. F. Reidhaar-Olson and C. M. Rondinone, 2009, vol. 555, p. 1 CrossRef CAS PubMed; K. M. Nelson and G. J. Weiss, Mol. Cancer Ther., 2008, 7, 3655 CrossRef CAS PubMed; H. Ling, M. Fabbri and G. A. Calin, Nat. Rev. Drug Discovery, 2013, 12, 847 CrossRef PubMed; C. Schöniger and C. Arenz, Bioorg. Med. Chem., 2013, 21, 6115 CrossRef PubMed; D. S. Karolina and K. Jeyaseelan, in Advanced Delivery and Therapeutic Applications of RNAi, John Wiley and Sons, Ltd, 2013, pp. 439–482 CrossRef PubMed; R. Garzon, G. Marcucci and C. M. Croce, Nat. Rev. Drug Discovery, 2010, 9, 775 CrossRef PubMed; J. A. Broderick and P. D. Zamore, Gene Ther., 2011, 18, 1104 CrossRef PubMed; G. Condorelli, M. V. G. Latronico and G. W. Dorn, Eur. Heart J., 2010, 31, 649 CrossRef PubMed.
- D. L. Ouellet, I. Plante, C. Barat, M. J. Tremblay and P. Provost, in siRNA and miRNA Gene Silencing, ed. M. Sioud, Humana Press, 2009, pp. 1–19 Search PubMed.
- S. Swaminathan, D. D. Murray and A. D. Kelleher, AIDS, 2012, 26, 1325–1334 CrossRef CAS PubMed; D. Boden, O. Pusch, R. Silbermann, F. Lee, L. Tucker and B. Ramratnam, Nucleic Acids Res., 2004, 32, 1154 CrossRef PubMed; R. Nathans, C.-Y. Chu, A. K. Serquina, C.-C. Lu, H. Cao and T. M. Rana, Mol. Cell, 34, 696 CrossRef PubMed.
- G. K. Scott, A. Goga, D. Bhaumik, C. E. Berger, C. S. Sullivan and C. C. Benz, J. Biol. Chem., 2007, 282, 1479 CrossRef CAS PubMed; S. M. Johnson, H. Grosshans, J. Shingara, M. Byrom, R. Jarvis, A. Cheng, E. Labourier, K. L. Reinert, D. Brown and F. J. Slack, Cell, 2005, 120, 635 CrossRef PubMed; Y. S. Lee and A. Dutta, Genes Dev., 2007, 21, 1025 CrossRef PubMed; C. D. Johnson, A. Esquela-Kerscher, G. Stefani, M. Byrom, K. Kelnar, D. Ovcharenko, M. Wilson, X. Wang, J. Shelton, J. Shingara, L. Chin, D. Brown and F. J. Slack, Cancer Res., 2007, 67, 7713 CrossRef PubMed; M. S. Kumar, S. J. Erkeland, R. E. Pester, C. Y. Chen, M. S. Ebert, P. A. Sharp and T. Jacks, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 3903 CrossRef PubMed; L. He, X. He, L. P. Lim, E. de Stanchina, Z. Xuan, Y. Liang, W. Xue, L. Zender, J. Magnus, D. Ridzon, A. L. Jackson, P. S. Linsley, C. Chen, S. W. Lowe, M. A. Cleary and G. J. Hannon, Nature, 2007, 447, 1130 CrossRef PubMed; G. J. Weiss, L. T. Bemis, E. Nakajima, M. Sugita, D. K. Birks, W. A. Robinson, M. Varella-Garcia, P. A. Bunn, J. Haney, B. A. Helfrich, H. Kato, F. R. Hirsch and W. A. Franklin, Ann. Oncol., 2008, 19, 1053 CrossRef PubMed; B. Kefas, J. Godlewski, L. Comeau, Y. Li, R. Abounader, M. Hawkinson, J. Lee, H. Fine, E. A. Chiocca, S. Lawler and B. Purow, Cancer Res., 2008, 68, 3566 CrossRef PubMed.
- S. Sassen, E. Miska and C. Caldas, Virchows Arch., 2008, 452, 1 CrossRef CAS PubMed; L.-A. MacFarlane and P. R. Murphy, Curr. Genomics, 2010, 11, 537 CrossRef PubMed.
- J. T. Mendell, Cell, 2008, 133, 217 CrossRef CAS PubMed; L. He, J. M. Thomson, M. T. Hemann, E. Hernando-Monge, D. Mu, S. Goodson, S. Powers, C. Cordon-Cardo, S. W. Lowe, G. J. Hannon and S. M. Hammond, Nature, 2005, 435, 828 CrossRef PubMed; F. Petrocca, R. Visone, M. R. Onelli, M. H. Shah, M. S. Nicoloso, I. de Martino, D. Iliopoulos, E. Pilozzi, C.-G. Liu, M. Negrini, L. Cavazzini, S. Volinia, H. Alder, L. P. Ruco, G. Baldassarre, C. M. Croce and A. Vecchione, Cancer Cell, 2008, 13, 272 CrossRef PubMed.
- J. Lu, G. Getz, E. A. Miska, E. Alvarez-Saavedra, J. Lamb, D. Peck, A. Sweet-Cordero, B. L. Ebert, R. H. Mak, A. A. Ferrando, J. R. Downing, T. Jacks, H. R. Horvitz and T. R. Golub, Nature, 2005, 435, 834 CrossRef CAS PubMed; S. Volinia, G. A. Calin, C.-G. Liu, S. Ambs, A. Cimmino, F. Petrocca, R. Visone, M. Iorio, C. Roldo, M. Ferracin, R. L. Prueitt, N. Yanaihara, G. Lanza, A. Scarpa, A. Vecchione, M. Negrini, C. C. Harris and C. M. Croce, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 2257 CrossRef PubMed; L. Zhang, J. Huang, N. Yang, J. Greshock, M. S. Megraw, A. Giannakakis, S. Liang, T. L. Naylor, A. Barchetti, M. R. Ward, G. Yao, A. Medina, A. O'Brien-Jenkins, D. Katsaros, A. Hatzigeorgiou, P. A. Gimotty, B. L. Weber and G. Coukos, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 9136 CrossRef PubMed.
- C. Dingwall, I. Ernberg, M. J. Gait, S. M. Green, S. Heaphy, J. Karn, A. D. Lowe, M. Singh, M. A. Skinner and R. Valerio, Proc. Natl. Acad. Sci. U. S. A., 1989, 86, 6925 CrossRef CAS.
- D. L. Ouellet, I. Plante, P. Landry, C. Barat, M.-È. Janelle, L. Flamand, M. J. Tremblay and P. Provost, Nucleic Acids Res., 2008, 36, 2353 CrossRef CAS PubMed; Z. Klase, R. Winograd, J. Davis, L. Carpio, R. Hildreth, M. Heydarian, S. Fu, T. McCaffrey, E. Meiri, M. Ayash-Rashkovsky, S. Gilad, Z. Bentwich and F. Kashanchi, Retrovirology, 2009, 6, 18 CrossRef PubMed.
- A. D. Frankel, Curr. Opin. Genet. Dev., 1992, 2, 293 CrossRef CAS; S. Roy, U. Delling, C. H. Chen, C. A. Rosen and N. Sonenberg, Genes Dev., 1990, 4, 1365 CrossRef; C. Dingwall, I. Ernberg, M. J. Gait, S. M. Green, S. Heaphy, J. Karn, A. D. Lowe, M. Singh and M. A. Skinner, EMBO J., 1990, 9, 4145 Search PubMed.
- S. Richter, H. Cao and T. M. Rana, Biochemistry, 2002, 41, 6391 CrossRef CAS PubMed.
- A. Davidson, T. C. Leeper, Z. Athanassiou, K. Patora-Komisarska, J. Karn, J. A. Robinson and G. Varani, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 11931 CrossRef CAS PubMed; D. Raghunathan, V. M. Sánchez-Pedregal, J. Junker, C. Schwiegk, M. Kalesse, A. Kirschning and T. Carlomagno, Nucleic Acids Res., 2006, 34, 3599 CrossRef PubMed.
- M. Takahashi, N. Yamada, H. Hatakeyama, M. Murata, Y. Sato, N. Minakawa, H. Harashima and A. Matsuda, Nucleic Acids Res., 2013, 41, 10659 CrossRef CAS PubMed; G. F. Deleavey and M. J. Damha, Chem. Biol., 2012, 19, 937 CrossRef PubMed.
- G. H. Clever and M. Shionoya, Coord. Chem. Rev., 2010, 254, 2391 CrossRef CAS PubMed; Y. Takezawa and M. Shionoya, Acc. Chem. Res., 2012, 45, 2066 CrossRef PubMed; G. H. Clever, C. Kaul and T. Carell, Angew. Chem., Int. Ed., 2007, 46, 6226 CrossRef PubMed; J. Müller, Eur. J. Inorg. Chem., 2008, 2008, 3749 CrossRef; P. Scharf and J. Müller, ChemPlusChem, 2013, 78, 20 CrossRef.
- F.-A. Polonius and J. Müller, Angew. Chem., Int. Ed., 2007, 46, 5602 CrossRef CAS PubMed.
- S. Taherpour, H. Lönnberg and T. Lönnberg, Org. Biomol. Chem., 2013, 11, 991 CAS.
- D. A. Berry, K.-Y. Jung, D. S. Wise, A. D. Sercel, W. H. Pearson, H. Mackie, J. B. Randolph and R. L. Somers, Tetrahedron Lett., 2004, 45, 2457 CrossRef CAS PubMed.
- R. A. Tinsley and N. G. Walter, RNA, 2006, 12, 522 CrossRef CAS PubMed.
- K. Burger, Biocoordination chemistry: coordination equilibria in biologically active systems, Ellis Horwood, 1990 Search PubMed.
- C. R. Cantor, M. M. Warshaw and H. Shapiro, Biopolymers, 1970, 9, 1059 CrossRef CAS PubMed; M. J. Cavaluzzi and P. N. Borer, Nucleic Acids Res., 2004, 32, e13 CrossRef PubMed; A. V. Tataurov, Y. You and R. Owczarzy, Biophys. Chem., 2008, 133, 66 CrossRef PubMed.
- S. Taherpour, O. Golubev and T. Lönnberg, J. Org. Chem., 2014, 79, 8990 CrossRef CAS PubMed.
- R. Sukackaite, S. Grazulis, G. Tamulaitis and V. Siksnys, Nucleic Acids Res., 2012, 40, 7552 CrossRef CAS PubMed.
- S. Taherpour and T. Lönnberg, J. Nucleic Acids, 2012, 2012, 196485 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: HPLC traces and mass spectra for oligonucleotides ON4a, ON4b, ON4c and ON5b; fluorescence emission spectra for oligonucleotides ON1a, ON1b, ON2, ON3a, ON3b and ON3c and their various duplexes; UV absorption of 1 at 330 nm as a function of [CuSO4]; UV spectra of duplex ON1c:ON4c at various concentrations of CuSO4. See DOI: 10.1039/c4ra15136d |
| This journal is © The Royal Society of Chemistry 2015 |