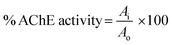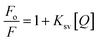Novel pyridinium oximes: synthesis, molecular docking and in vitro reactivation studies†
Poojaab,
S. Aggarwalab,
A. K. Tiwari*a,
V. Kumara,
R. Pratapb,
G. Singhb and
A. K. Mishra*a
aDivision of Cyclotron and Radiopharmaceutical Sciences, Institute of Nuclear Medicine and Allied Sciences, Brig. S. K. Mazumdar Road, Delhi-110054, India. E-mail: anjani7797@rediffmail.com; akmishra63@gmail.com
bDepartment of Chemistry, University of Delhi, Delhi-110054, India
First published on 24th February 2015
Abstract
A computational approach has been attempted for the screening of 4-pyridoxinium (4P) ring based reactivators for paraoxon inhibited AChE. The oxime molecules were designed with the common 4P skeleton and varying the carbon linkers. Initially, the AChE binding capability was assessed by molecular docking with PDB:2WHP and 3ZLV, which showed important interactions with Ser298, Try124 and Trp286. These computational results were validated by an in vitro AChE binding assay, which showed binding affinities in the range of 10–90%. Finally, reactivation potency was calculated as % reactivation on paraoxon inhibited eelAChE in the concentration range of 10−5 to 10−7 M. It was observed that introduction of an aliphatic linker attached to 4-pyridoxime has a high binding affinity and hence, may act as a good reactivator as compared to the aromatic pyridoximes.
1. Introduction
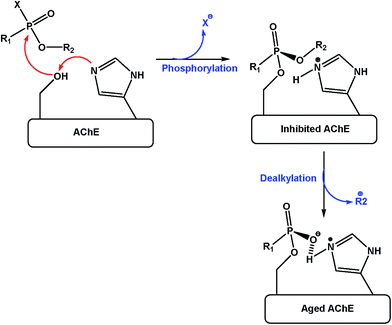
Acetylcholinesterase (AChE) is a very important enzyme for the hydrolysis of neurotransmitter acetylcholine at the cholinergic synaptic site. It has two binding sites, first is the active site known as the A-site and second is a peripheral anionic site known as the PAS or P-site. Hydrolysis of acetylcholine takes place via hydrolyzing the serine residue by the active site of the AChE enzyme (Scheme 1).1–6 The major inhibitors of AChE are nerve gas agents, which include different classes of organophosphorus (OP) compounds which bind with the –OH of the serine residue and block the action of AChE towards the hydrolysis of ACh into choline and acetic acid.7–9 This blockage by these OP agents is responsible for the high level of ACh at the cholinergic sites leading to failure of the neurotransmission process and can cause various neurological diseases of the CNS. The most well known OP agents are sarin, soman, tabun, VX and paraxon (Fig. 1). To inhibit the action of OP agents and to reactivate OP-inhibited AChE, various pyridoxime derivatives (Fig. 2) have been synthesized and reported in the literature e.g., 2-PAM, HI-6 and obidoxime.10–13 Pyridoxime derivatives have substitution (hydroxoiminomethyl) at their 2, 3 or 4 positions. They may interact at the active site and break the covalent bond formed between the OP agents and –OH of the serine residue of OP-inhibited AChE. 4-Pyridoxime derivatives were observed to have high potency to act as reactivators of OP-inhibited AChE as compared to 2 and 3 pyridoximes. It was also observed that the positive charge on the nitrogen of the pyridoxime ring makes it more potent for reactivation but it increases the hydrophilicity of the oxime moiety making it less lipophilic to cross the blood–brain barrier (BBB), which is the most important lipoidal membrane. Therefore, various modifications have been done and different non-ionic oxime derivatives were reported in the literature, but it was found that they have less reactivation potency, as compared to ionic oximes. This implies that the positive charge on pyridoxime ring is important. Furthermore, carbon chain linker was introduced at the –N position of pyridoxime derivatives to give lipophilicity to the molecule for enhancing its permeability. The studied chain with the 4-pyridoxime are alkyl, alkoxyl, and aromatic carbon chains.14,15 It was found that as the carbon chain length increases, lipophilicity and permeability to cross BBB increases and % reactivation towards the OP-inhibited AChE also increases. The 4-pyridoxime derivatives have higher lipophilicity when compared with 2-PAM, e.g., 30% for 4-octyl-PAM.1These oximes have been studied on the species like rodents, which are similar to human.16 We used eelAChE due to the easy and readily availability, which is one of the most explored species in preliminary studies. Though other studies have been also performed previously, to show variation in the binding and reactivation potency of oximes with different species.17–19
Here, we report the synthesis of six 4-pyridoxime derivatives having aliphatic and aromatic carbon chain linker attached at the nitrogen (–N) of the pyridoxime moiety. These derivatives were selected on the basis of preliminary computational studies and comparison with the previously synthesized oxime derivatives. Finally, in vitro AChE binding and reactivation study was performed to see the efficacy of these molecules for further applications.
2. Experimental
2.1 Chemicals
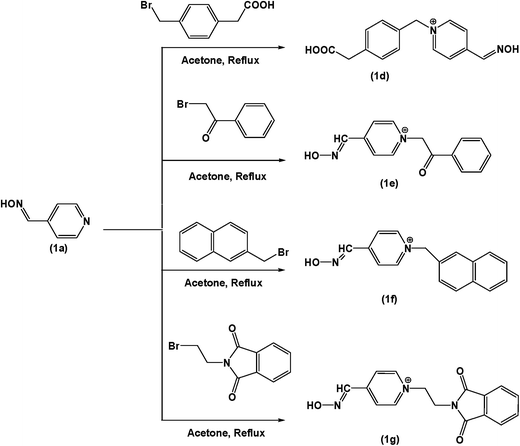
4-Pyridine carboxaldehyde, hydroxylamine hydrochloride, 1,3-dibromo propane, 1-bromo butyrl chloride, 4-bromomethyl phenyl acetic acid, 2-bromomethylnaphthalene, N-(2-bromoethyl) phthalimide, 2-bromoacetophenone, paraoxon, eel acetylcholinesterase (eelAChE) enzyme, acetylthiocholine iodide, (DTNB), purchased from Sigma-Aldrich Co, USA. Thin layer chromatography on aluminum plates coated with silica gel 1160, F254, CHCl3, acetone, isopropanol, sodium monophosphate, and sodium diphosphate purchased from Merck, Germany.Six different oximes were synthesized on the basis of preliminary computational prediction which are presented in Fig. 3. The synthetic schemes are shown as Scheme 2 and 3.
2.2 Synthesis of pyridine-4-carbaldehyde oxime (4P)
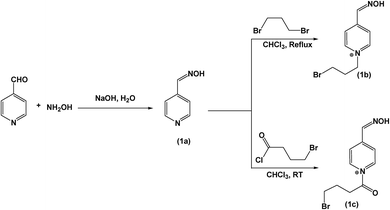
A solution of hydroxylamine hydrochloride (9.34 mmol, 650 mg) in water was stirred on a magnetic stirrer. The pH of the reaction mixture was maintained at 6–7 throughout the reaction by using 1 M NaOH solution.20 Aqueous solution of 4-pyridine carbaldehyde (4.67 mmol, 0.44 mL) was added dropwise. Reaction mixture was kept on heating at 50 °C for 12 h. After that solvent was evaporated on rota evaporator and final product was dried under vacuum.2.3 Synthesis of 1-(3-bromo-propyl)-4-(hydroxyimino-methyl)-pyridinium (4PBP)
Pyridoxinium ion was synthesized by using 4-pyridoxime (2.03 mmol, 250 mg) in dried CHCl3 and 1,3-dibromo propane (10.15 mmol, 1.18 mL) in CHCl3.21 This reaction mixture was stirred first at room temperature for 10 min after that allowed at reflux condition of temperature 60 °C for 72 h. White precipitate was appeared in the reaction mixture, which was filtered and dried under vacuum.2.4 Synthesis of 1-(4-bromo-butyryl)-4-(hydroxyimino-methyl)-pyridinium (4PBB)
4PBB was synthesised similar to the procedure for 4PBP by adding 4-bromo butyryl chloride (2.23 mmol, 0.28 mL) in CHCl3 in place of 1,3-dibromo propane. Reaction mixture was stirred at room temperature for 1 h after that white precipitates appeared in the reaction mixture. It was filtered and solvent was evaporated under vacuum on rota evaporator.2.5 Synthesis of 1-(4-carboxymethyl-benzyl)-4-(hydroxyimino-methyl)-pyridinium (4PCB)
4PCB was synthesised by the reaction between 4-pyridoxime (2.03 mmol, 250 mg) in dried acetone and 4-(bromo methyl) phenyl acetic acid (3.65 mmol, 1.680 g).22 Reaction mixture was stirred at refluxing for 28 h after that precipitates appeared in the reaction mixture. It was filtered and solvent was evaporated under vacuum on rota evaporator.2.6 Synthesis of 4-(hydroxyimino-methyl)-1-(2-oxo-2-phenyl-ethyl)-pyridinium (4OA)
In a two neck round bottom flask, reaction of 4-pyridoxime (1.00 mmol, 124 mg) with 2-bromoacetophenone (2.50 mmol, 497.6 mg) was added in dry acetone. Reflux conditions were maintained for 15 h. The progress of the reaction was checked using TLC.2.7 Synthesis of 4-(hydroxyimino-methyl)-1-naphthalen-2-ylmethyl-pyridinium (4ON)
Synthesis of ‘4-(hydroxyimino-methyl)-1-naphthalen-2-ylmethyl-pyridinium’ (4ON) was carried out by a substitution reaction of 4-pyridoxime with bromo-functionalized naphthalene moiety. 4-Pyridoxime (1.00 mmol, 123.7 mg) was added in a two neck round bottom flask and dissolved in dry acetone. Then 2-bromomethylnaphthalene (2.50 mmol, 552.7 mg) was added to the reaction mixture and stirring at reflux was continued for 15 h. The TLC was monitored at regular intervals.2.8 Synthesis of 1-[2-(1,3-dioxo-1,3-dihydro-isoindol-2-yl)-ethyl]-4-(hydroxyimino-methyl)-pyridinium (4OPh)
To a solution of 4-pyridoxime (1.00 mmol, 123.7 mg) in dry acetone, N-(2-bromoethyl) phthalimide (2.50 mmol, 635.2 mg) was added. The reaction mixture was kept at stirring under reflux conditions for 15 h and the completion of the reaction was checked by TLC.2.9 Computational approach for AChE interaction
The X-ray crystal structure of sarin phosphonylated (PDB entry 2WHP, having resolution of 2.20 Å)23 and tabun complexed acetylcholinesterase (PDB entry 3ZLV, having resolution of 2.50 Å)24 were used for molecular docking studies. The following criteria were kept in mind before choosing the protein–ligand complexes: non-covalent binding between protein and ligand, crystallographic resolution less than or equal to 2.5 Å and known experimental binding data. Molecular modeling investigations were carried out using an advanced molecular docking program GLIDE, 5.8.25 During the docking process, initially GLIDE performs a complete systematic search of the conformational, orientation and positional space of the docked ligand eliminating unwanted conformations using scoring followed by energy optimization. Preparation of the protein for docking included removal of unnecessary heteroatom and solvent coupled with addition of hydrogen atoms, bond order for crystal ligand and protein were adjusted and minimized up to 0.30 Å RMSD.2.10 Binding study with AChE
Mixture of 750 μL enzyme (eelAChE, 0.1 mg in 20 mL sodium phosphate buffer of pH 7.4), 750 μL acetylthiocholine iodide as a substrate (550 μM) and 750 μL oxime solution in the range of concentration 1 mM to 1 nM was incubated for 12 h at 25 °C. A parallel experiment was running in which 750 μL buffer was used in place of test compound in the enzyme solution as a positive control. In each concentration sample 750 μL DTNB solution (80 μM, prepared in phosphate buffer of pH 7.4) was added and absorbance was recorded at 412 nm at every 5 min for 30 min.26,27Enzyme activity was calculated by using the formula:
2.11 Reactivation study towards paraoxon inhibited AChE
To study the reactivation on inhibited AChE, the foremost necessity was to know about the inhibited enzyme activity. So, 30 μL enzyme was incubated with 10 μL paraoxon (10−2 to 10−5 M) for 15 min to achieve >95% inhibition. Thereafter, a mixture of 40 μL inhibited enzyme, 40 μL acetylthiocholine iodide and 40 μL PBS buffer was incubated for 15 min. Then, absorbance (Ai) was measured at 412 nm by adding 40 μL DTNB at different time intervals, 0 min, 15 min, 30 min, 45 min and 60 min.For reactivation study, a mixture of 40 μL inhibited enzyme, 40 μL acetylthiocholine iodide and 40 μL test oxime (10−3 to 10−8 M) was incubated for 15 min and absorbance (Ar) was recorded at 412 nm by adding 40 μL DTNB at different time interval. A parallel positive control experiment was done by taking 40 μL enzyme and 40 μL buffer in place of inhibited enzyme and test oxime respectively.1,15,26
% AChE reactivation was calculated by using the formula:
| % AChE reactivation = (Ar − Ai/Ao − Ai) × 100 |
2.12 Fluorometric assay with human serum albumin (HSA)
Binding constant of the oximes with HSA was calculated by the fluorometric experiments. The fluorescence intensity was recorded at λem = 350 nm at the fixed λex = 280 nm for the different concentration of oximes from 4 × 10−4 to 10−5 M in PBS of pH = 7.4 at 25 °C while the concentration of HSA was fixed at 2 × 10−5 M. The change in intensity at 350 nm (tryptophan) was used to calculate the binding constant (K).28,29 The binding constant was calculated by the Stern–Volmer equation:30where Fo and F denotes the fluorescence intensity in the absence and presence of quencher pyridoxime, respectively. Ksv is the Stern–Volmer quenching constant (or binding constant) and [Q] is the concentration of quencher.
3. Result and discussion
3.1 Synthesis of 4P
4-Pyridoxime was synthesized by the hydrolysis reaction between 4-pyridine carbaldehyde and hydroxylamine as shown in Scheme 2. The product was obtained in high yield of 97.17% by evaporating solvent and its structure was confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S1†).1H NMR (400 MHz, DMSO): δ = 7.5 (d, 2H, ArH), 8.1 (d, 1H, –CH proton), 8.5 (d, 2H, ArH), 13C NMR (100 MHz, DMSO): δ = 120.94 (ArC), 141.05 (ArC), 146.73 (ArC), 150.68 (–CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NOH), m/z (ESI-LCMS): calculated mass [M+] = 122.05, [M + H+] = 123.4.
NOH), m/z (ESI-LCMS): calculated mass [M+] = 122.05, [M + H+] = 123.4.
3.2 Synthesis of 4PBP
N-(3-Bromo-propyl)-4-pyridoxime was synthesized by the N-alkylation reaction between 4-pyridoxime and 1,3-dibromo propane as shown in Scheme 2. The product was obtained as brown precipitate with moderate yield of 60.34% and its structure was confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S2†).1H NMR (400 MHz, D2O): δ = 8.75 (d, 2H, ArH), 8.66 (s, 1H, ArH), 8.25 (d, 2H, ArH), 4.65 (s, D2O), 3.43 (t, 2H, –CH2), 2.66 (t, 2H, –CH2), 2.14 (m, 2H, –CH2), 13C NMR (100 MHz, D2O): δ = 149.19 (–CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NOH), 146.23 (ArC), 144.62 (ArC), 125.13 (ArC), 57.70, 32.61, 31.58, m/z (ESI-LCMS): calculated mass [M+] = 243.01, [M + H+] = 244.9, with M: (M + 2)
NOH), 146.23 (ArC), 144.62 (ArC), 125.13 (ArC), 57.70, 32.61, 31.58, m/z (ESI-LCMS): calculated mass [M+] = 243.01, [M + H+] = 244.9, with M: (M + 2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 bromo isotopic pattern.
1 bromo isotopic pattern.
3.3 Synthesis of 4PBB
N-Alkylation reaction between 4-pyridoxime and 4-bromo butyryl chloride yields N-(4-bromo-butyryl)-4-pyridoxime with 59.24% as shown in Scheme 2. Its structure was confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S3†).1H NMR (400 MHz, D2O): δ = 8.74 (d, 2H, ArH), 8.66 (s, 1H, ArH), 8.59 (d, 2H, ArH), 4.65 (s, D2O), 3.42 (t, 2H, –CH2), 2.66 (t, 2H, –CH2), 2.41 (m, 2H, –CH2), 13C NMR (100 MHz, D2O): δ = 183.22 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 146.78 (–CH
O), 146.78 (–CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NOH), 141.49 (ArC), 130.47 (ArC), 124.17 (ArC), 30.33, 27.94, 26.48, m/z (ESI-LCMS): calculated mass [M+] = 271, [M + H+] = 271.3, with M: (M + 2)
NOH), 141.49 (ArC), 130.47 (ArC), 124.17 (ArC), 30.33, 27.94, 26.48, m/z (ESI-LCMS): calculated mass [M+] = 271, [M + H+] = 271.3, with M: (M + 2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 bromo isotopic pattern.
1 bromo isotopic pattern.
3.4 Synthesis of 4PCB
N-Alkylation between 4-pyridoxime and 4-bromomethyl phenyl acetic acid gives N-(4-carboxymethyl-benzyl)-4-pyridoxime in the form of yellow colored compound as shown in Scheme 2. 78.45% yield was obtained and structure confirmation was done by 1H NMR, 13C NMR and mass spectroscopy (shown as S4†).1H NMR (400 MHz, D2O): δ = 8.70 (d, 2H, ArH), 8.19 (s, 1H, –CH), 8.01 (d, 2H, ArH), 7.22–7.31 (dd, 4H, ArH), 5.62 (s, 2H, –CH2), 4.65 (s, D2O peak), 3.59 (s, 2H, –CH2); 13C NMR (100 MHz, D2O): δ = 176.63 (–COOH), 148.93 (–CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NOH), 146.22, 144.48, 136.23, 131.58, 130.65, 129.48, 124.91 (ArC's), 63.86, 40.39, m/z (ESI-LCMS): calculated mass [M+] = 271.11, [M+] = 271.3, [M + Na+] = 294.
NOH), 146.22, 144.48, 136.23, 131.58, 130.65, 129.48, 124.91 (ArC's), 63.86, 40.39, m/z (ESI-LCMS): calculated mass [M+] = 271.11, [M+] = 271.3, [M + Na+] = 294.
3.5 Synthesis of 4OA
4-(Hydroxyimino-methyl)-1-(2-oxo-2-phenyl-ethyl)-pyridinium was synthesized by the N-alkylation reaction between 4-pyridoxime and 2-bromoacetophenone as shown in Scheme 3. The product was obtained by evaporating solvent as a white powder in moderate yield of 72.01% and its structure has been confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S5†).1H NMR (CDCl3, 400 MHz): δ values (ppm) = 4.19 (s, 2H, CH2), 7.51–7.69 (m, 3H, ArH), 7.94–7.96 (d, 2H, ArH), 8.04–8.07 (m, 1H, ArH) 8.25 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 8.69–8.71 (d, 2H, ArH), 8.88 (s, 1H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 60.4 (CH2), 114.6 (ArCH), 128.1 (ArCH), 128.5 (ArCH), 129.5 (ArCH), 132.7 (ArCH), 133.6 (ArCH), 135.8 (ArCH), 143.3 (ArCH), 143.9 (ArC), 144.6 (ArC), 145.8 (CH), 192.1 (C
N), 8.69–8.71 (d, 2H, ArH), 8.88 (s, 1H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 60.4 (CH2), 114.6 (ArCH), 128.1 (ArCH), 128.5 (ArCH), 129.5 (ArCH), 132.7 (ArCH), 133.6 (ArCH), 135.8 (ArCH), 143.3 (ArCH), 143.9 (ArC), 144.6 (ArC), 145.8 (CH), 192.1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). ESI-MS: m/z = 241.2 [M]+.
O). ESI-MS: m/z = 241.2 [M]+.
3.6 Synthesis of 4ON
4-(Hydroxyimino-methyl)-1-naphthalen-2-ylmethyl-pyridinium was synthesized by the N-alkylation reaction between 4-pyridoxime and 2-bromomethylnaphthalene as shown in Scheme 3. The product was obtained by evaporating solvent as a pale yellow powder in moderate yield of 65.12% and its structure has been confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S6†).1H NMR (CDCl3, 400 MHz): δ values (ppm) = 2.91 (s, 2H, CH2), 7.27–7.7.81 (m, 7H, ArH), 8.17–8.19 (d, 2H, ArH), 8.45 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 8.91–8.93 (d, 2H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 64.3 (CH2), 124.7 (ArCH), 125.5 (ArCH), 127.2 (ArCH), 127.5 (ArCH), 127.7 (ArCH), 128.1 (ArCH), 128.9 (ArCH), 129.5 (ArCH), 130.2 (ArCH), 132.8 (ArC), 133.1 (ArC), 144.3 (ArC), 146.0 (ArC), 148.7 (CH). ESI-MS: m/z = 263.3 [M]+.
N), 8.91–8.93 (d, 2H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 64.3 (CH2), 124.7 (ArCH), 125.5 (ArCH), 127.2 (ArCH), 127.5 (ArCH), 127.7 (ArCH), 128.1 (ArCH), 128.9 (ArCH), 129.5 (ArCH), 130.2 (ArCH), 132.8 (ArC), 133.1 (ArC), 144.3 (ArC), 146.0 (ArC), 148.7 (CH). ESI-MS: m/z = 263.3 [M]+.
3.7 Synthesis of 4OPh
1-[2-(1,3-Dioxo-1,3-dihydro-isoindol-2-yl)-ethyl]-4-(hydroxyimino-methyl)-pyridinium (1 g) was synthesized by the N-alkylation reaction between 4-pyridoxime and N-(2-bromoethyl)phthalimide as shown in Scheme 3. The product was obtained by evaporating solvent as a white powder in moderate yield of 61.34% and its structure has been confirmed by 1H NMR, 13C NMR and mass spectroscopy (shown as S7†).1H NMR (CDCl3, 400 MHz): δ values (ppm) = 4.18–4.21 (t, 2H, CH2), 4.76–4.79 (t, 2H, CH2), 7.71–7.75 (m, 4H, ArH), 8.04–8.06 (d, 2H, ArH), 8.24 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 8.75–8.77 (d, 2H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 38.3 (CH2), 59.6 (CH2), 123.7 (ArCH), 124.9 (ArCH), 130.7 (ArCH), 135.1 (ArCH), 144.9 (ArC), 146.0 (ArC), 149.4 (CH), 169.7 (C
N), 8.75–8.77 (d, 2H, ArH). 13C NMR (CDCl3, 100 MHz): δ values (ppm) = 38.3 (CH2), 59.6 (CH2), 123.7 (ArCH), 124.9 (ArCH), 130.7 (ArCH), 135.1 (ArCH), 144.9 (ArC), 146.0 (ArC), 149.4 (CH), 169.7 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). ESI-MS: m/z = 296.1 [M]+.
O). ESI-MS: m/z = 296.1 [M]+.
3.8 Computational approach for AChE interaction
Molecular modeling studies were performed on synthesized oxime molecules 4PBP, 4PBB, 4PCB, 4OA, 4ON and 4OPh for the justification of their interaction with OP-inhibited hAChE. It is important for a reactivator of AChE to bind with the enzyme which was revealed in the form of Glide Gscore. Binding affinity of synthesized 4-pyridoximes varies by changing the alkyl chain substituted to the pyridinium nitrogen of the ring. It was observed that the Glide Gscore for aliphatic chain alkyl group attached to pyridoximes are high (−9.920, −11.021 for 4PBP and −10.793, −10.902 for 4PBB) as compared to the aromatic (−9.951, −9.760 for 4PCB, −10.250, −8.927 for 4OA, −8.639, −8.261 for 4ON and −9.202, −8.244 for 4OPh) with both the PDB's 2WHP and 3ZLV (detail shown in S8†). This variation in Glide Gscores is due to difference in the type of linkers in each oxime which affect the binding with various amino acid residues. These synthesized oximes showed binding with Trp286 and Tyr124 via π–π interaction with the aromatic moiety of amino acid and pyridinium ring, which is an important interaction for oximes to reactivate nerve gas inhibited AChE.26 These amino acid residues are present at peripheral anionic binding site of enzyme AChE. It is known that attack of OP on the –OH of serine residue present at the esteric site is important for any reactivator to bind with the serine residue and its bond breakage was previously formed by OP agents. Oximes showed binding with Ser298 of AChE by hydroxyimino moiety via hydrogen bonding interaction, except for aromatic linker attached oximes (4OA, 4ON and 4OPh). In 4PBP and 4PBB extra interactions were observed including π–π interaction with the amino acid residue Tyr72. While, aromatic linker attached pyridoximes showed less interaction as compared to aliphatic linker due to rigidity in the structure. The 2-D diagrams of oximes with PDB 2WHP and 3ZLV were shown in Fig. 4 and 5, respectively. These results were compared with the well known AChE reactivator 2-PAM (2D interaction shown in Fig. 6). The reliability of the docking program Glide used in this study was verified evaluating its performances in reproducing the geometry of bound inhibitors taken from cocrystallized complexes (2WHP and 3ZLV). The docking calculation reproduces the actual orientation of these molecules as shown in S9.† This was further validated by MD simulation and MMGBSA analysis. Furthermore, to validate this computational binding there in vitro binding and reactivation potency were studied towards OP-inhibited AChE.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 394 atoms, to MD simulation which was subsequently carried out by using following procedure: first, default 1.2 ns run of MD simulation at constant temperature of 300 K and with all atoms having positions restrained except for the water molecules inside the box under the NPT ensemble. This initial MD was necessary in order to allow the equilibration of the solvent molecules around the protein. Another production run was carried out which included: full MD simulations of 5 ns at 300 K with no restrictions with 2 fs of integration time. After completion, simulations quality and event analysis was done.
394 atoms, to MD simulation which was subsequently carried out by using following procedure: first, default 1.2 ns run of MD simulation at constant temperature of 300 K and with all atoms having positions restrained except for the water molecules inside the box under the NPT ensemble. This initial MD was necessary in order to allow the equilibration of the solvent molecules around the protein. Another production run was carried out which included: full MD simulations of 5 ns at 300 K with no restrictions with 2 fs of integration time. After completion, simulations quality and event analysis was done.| ΔGbind = ΔEMM + ΔGsolv + ΔGSA |
MMGBSA energy profile with 3ZLV and 2WHP are shown in S10 and S11† along with 3D interactive poses in S12† and simulation interactions for the NPT ensemble 4PBB-2WHP presented in S13† for 5 ns.
3.9 Binding study with AChE
An oxime can only reactivate AChE if it goes to the site of OP-inhibited AChE and bind to inhibit it. To get more insight of the reactivation of inhibited AChE, binding affinity of the synthesized oximes was evaluated with eelAChE by measuring their inhibitory activity using Ellman method. Binding experiment was performed in which eelAChE was incubated with different concentrations (10−3 to 10−9 M) of oximes for 12 h to achieve the maximum inhibition. Binding affinity was also measured for the precursor 4-pyridoxime to compare. The in vitro data is shown in Fig. 7. It was observed that all the synthesized oximes showed change in binding affinity by changing the concentration. Out of six synthesized oximes, 4PBB and 4PCB showed highest binding affinity. At the concentrations 10−9 to 10−7 M, 4PBP have less inhibitory activity as compared to 4PBB and 4PCB. But at the concentrations 10−6, 10−4 and 10−3 M it showed inhibitory activity of 1%, 1% and 50% for 4PBP, 15%, 80% and 95% for 4PBB, 40%, 92% and 60% for 4PCB, 36%, 29% and 74% for 4OA, 44%, 45% and 52% for 4ON, 30%, 87% and 89% for 4OPh and 10%, 40% and 60% for precursor 4P respectively. It was also observed that at the biological concentration of 10−9, 10−8 and 10−7 M these oximes do not show much efficient inhibitory activity as compared to the concentration of 10−6, 10−4 and 10−3 M. They showed optimum inhibitory activity at the concentration range of 10−4 M.3.10 Reactivation study towards paraoxon inhibited AChE
To validate the binding affinity of synthesized pyridoximes with AChE, in vitro reactivation experiment was demonstrated with the paraoxon inhibited eel-AChE.10 Enzyme was incubated with the nerve gas paraoxon about 30 min to achieve maximum inhibition. Then enzyme activity was determined by performing well known Ellman assay27 using DTNB as a coloring agent. This assay was performed on the synthesized pyridoximes (4PBP, 4PBB, 4PCB, 4OA, 4ON and 4OPh) as well as with the synthesized precursor 4P. These reactivation potencies were also compared with the currently used AChE reactivator 2-PAM. The results of reactivation experiment were shown in Fig. 8, which showed reactivation % by varying the concentration of pyridoximes.This experiment showed that the reactivation potency of the oximes varied by changing the linker at the pyridinium nitrogen of the ring. It was observed that the reactivation potency for the synthesized oximes 4PBP and 4PBB were high as compared to the aromatic ring bearing pyridoximes (4PCB, 4OA, 4ON and 4OPh). Results showed 61.82%, 89.09%, 76.36% reactivation potency for 4PBP, 76.36%, 65.45%, 81.82% for 4PBB, −32.73%, 32.73%, 21.82% for 4PCB, −0.04%, −0.19%, −0.03% for 4OA, 3.00%, 5.00%, −1.00% for 4ON and −0.04%, 0.02%, −0.02% for 4OPh at the concentration of 10−5 M, 10−6 M and 10−7 M respectively. These results were also compared with the precursor 4P which showed reactivation potency 56.36%, 49.09%, 56.36% for the same concentration range as for the products. Reactivation potency was evaluated for the known reactivator 2-PAM which showed 16.39%, 9.18%, 2.30% reactivation at the concentration 10−5 M, 10−6 M and 10−7 M respectively. For the aromatic ring bearing pyridoximes, reactivation was also studied at high concentration of 10−3 M and 10−4 M. It was found that these oximes able to reactivate paraoxon inhibited AChE at high concentrations with the % reactivation of, 22.73%, 36.36% for 4PCB at 10−3 M and 10−4 M, 20.00%, for 4OA at 10−3 M, 49.51%, 42.95% for 4ON at 10−3 M and 10−4 M and 42.95%, 15.08% for 4OPh at 10−3 M and 10−4 M respectively. This data stated as we move from aliphatic to aromatic linker, reactivation potency decreases and also the synthesized oximes showed high reactivation potency at low concentration range between 10−6 to 10−7 M. Secondly, these data showed better reactivation than known reactivator 2-PAM. In 2-PAM, oxime group was at 2nd position of the pyridinium ring while it was at 4th position in our synthesized oximes. Synthesized oxime precursor 4P is also more effective than the 2-PAM at low concentrations.
Reactivation potency for 2-PAM decreased as the concentration decreases which showed that they are good reactivators at the high concentration but not at the low concentration. Both the aliphatic chain linked with pyridoximes, 4PBP and 4PBB showed similar reactivation potencies, but if observed at each concentration then it was found that 4PBB have good reactivation potency (81.82%) at the concentration 10−7 M whereas 4PBP showed excellent reactivation potency (89.09%) at 10−6 M.
3.11 HSA binding assay
Interaction of ligand with HSA was analyzed by in vitro fluorometric assay. HSA shows emission because of aromatic amino acids tryptophan, tyrosine and phenylalanine residues present in it. Quenching of fluorescence intensity of tyrosine was observed if it is present near an amino, a carboxyl group or a tryptophan, and phenylalanine with very low quantum yield. Small molecules bound to HSA tend to change in intrinsic fluorescence intensity due to tryptophan residue in the oxime system. Hence, fluorescence quenching was observed by addition of oxime in the concentration range of 0.4–0.05 mM to the fixed concentration of HSA (0.02 mM) at 350 nm (Fig. 9(a–d)). This quenching indicates the formation of ligand–HSA complex which leads to microenvironment change in the protein and hence showed decrease in fluorescence intensity. | ||
| Fig. 9 Emission plot for ligand–HSA fluorescence quenching with different oximes. # (a) 4P (b) 4PBP (c) 4PBB (d) 4PCB. | ||
The binding constants of the synthesized oximes as well as precursor 4P were calculated by Stern–Volmer plot and these data were shown in Fig. 10. Stern–Volmer binding constants were calculated as 1.17 × 103 M−1, 2.29 × 103 M−1, 0.90 × 103 M−1 and 0.75 × 103 M−1 for 4P, 4PBB, 4PBP and 4PCB respectively. These binding constant values suggested that the newly synthesized oximes have ability to bind with HSA for their transportation at the specific site in the body to reactivate inhibited AChE.
4. Conclusion
In this study mono-quaternary oximes were synthesized as reactivator for paraoxon inhibited eelAChE by variation in the carbon chain attached with the –N position of 4-pyridoxime moiety. These oximes were studied for their binding affinity with eelAChE. Further, this binding was validated with the molecular modeling studies on the basis of Glide Gscore, π–π interaction and hydrophobic interaction on the active site of AChE. These synthesized oximes were found to be effective reactivators towards the paraoxon inhibited eelAChE as compared to the known reactivator 2-PAM. 4PBB and 4PBP have high reactivation potency as compared to 4PCB. Furthermore, they also showed drug likeliness behavior by showing effective pharmacokinetics properties with human serum albumin.Acknowledgements
We thank Dr R. P. Tripathi, Director, INMAS, DRDO for providing the necessary facilities and CSIR for financial assistance to the first and second authors during the course of the research work.Notes and references
- M. C. De Koning, M. J. A. Joosen, D. Noort, A. Van Zuylen and M. C. Tromp, Bioorg. Med. Chem., 2011, 19, 588–594 CrossRef CAS PubMed.
- Z. Kovarik, N. Macek, R. K. Sit, Z. Radic, V. V. Fokin, K. B. Sharpless and P. Taylor, Chem.–Biol. Interact., 2013, 203, 77–80 CrossRef CAS PubMed.
- T. H. Kim, K. Kuca, D. Jun and Y. S. Jung, Bioorg. Med. Chem. Lett., 2005, 15, 2914–2917 CrossRef CAS PubMed.
- K. Musilek, K. Kuca, D. Jun, V. Dohnal and M. Dolezal, Bioorg. Med. Chem. Lett., 2006, 16, 622–627 CrossRef CAS PubMed.
- K. Musilek, O. Holas, K. Kuca, D. Jun, V. Dohnal and M. Dolezal, Bioorg. Med. Chem. Lett., 2006, 16, 5673–5676 CrossRef CAS PubMed.
- K. Musilek, J. Roder, M. KomLoova, O. Holas, M. Hbinova, M. Pohanka, V. Dohnal, V. Opletalova, K. Kuca and Y. S. Jung, Bioorg. Med. Chem. Lett., 2011, 21, 150–154 CrossRef CAS PubMed.
- S. B. Bharate, L. Guo, T. E. Reeves, D. M. Cerasoli and C. M. Thompson, Bioorg. Med. Chem., 2010, 18, 787–794 CrossRef CAS PubMed.
- J. Acharya, A. K. Gupta, A. Mazumder and D. K. Dubey, Toxicol. In Vitro, 2008, 22, 525–530 CrossRef CAS PubMed.
- G. Sinko, J. Brglez and Z. Kovarik, Chem.–Biol. Interact., 2010, 187, 172–176 CrossRef CAS PubMed.
- S. B. Bharate, L. Guo, T. E. Reeves, D. M. Cerasoli and C. M. Thompson, Bioorg. Med. Chem. Lett., 2009, 19, 5101–5104 CrossRef CAS PubMed.
- K. Musilek, O. Holas, D. Jun, V. Dohnal, F. G. Moore, V. Opletalova, M. Dolezala and K. Kuca, Bioorg. Med. Chem., 2007, 15, 6733–6741 CrossRef CAS PubMed.
- K. Kuca, J. Bielavsky, J. Cabal and M. Bielavska, Tetrahedron Lett., 2003, 44, 3123–3125 CrossRef CAS.
- F. Ekstrom, J. C. Astot and Y. P. Pang, Clin. Pharmacol. Ther., 2007, 82, 282–293 CrossRef CAS PubMed.
- A. Luettringhaus and I. Hagedorn, Drug Res., 1964, 14, 1–5 CAS.
- J. Acharya, D. K. Dubey, A. K. Srivastava and S. K. Raza, Toxicol. In Vitro, 2011, 25, 251–256 CrossRef CAS PubMed.
- J. J. Kassa, J. Toxicol., Clin. Toxicol., 2002, 40, 803–816 CrossRef CAS PubMed.
- F. Worek, G. Reiter, P. Eyer and L. Szinicz, Arch. Toxicol., 2002, 76, 523–529 CrossRef CAS PubMed.
- J. G. Clement and N. Erhardt, Arch. Toxicol., 1994, 68, 648–659 CrossRef CAS.
- W. K. Berry, Biochem. Pharmacol., 1971, 20, 1333–1334 CrossRef CAS.
- X. Q. Feng, Z. S. Zeng, Y. H. Liang, F. E. Chen, C. Pannecouque, J. Balzarini and E. D. Clercq, Bioorg. Med. Chem., 2010, 18, 2370–2374 CrossRef CAS PubMed.
- S. R. Chennamaneni, V. Vobalaboina and A. Garlapati, Bioorg. Med. Chem. Lett., 2005, 15, 3076–3080 CrossRef CAS PubMed.
- A. K. Sikder, A. K. Ghosh and D. K. Jaiswal, J. Pharm. Sci., 1993, 82, 258–261 CrossRef CAS.
- F. Ekstrom, A. Hornberg, E. Artursson, L. G. Hammarstrom, G. Schneider and Y. P. Pang, PLoS One, 2009, 4(6), e5957 Search PubMed.
- E. Artursson, P. O. Andersson, C. Akfur, A. Linusson, S. Borjegren and F. Ekstrom, Biochem. Pharmacol., 2013, 85, 1389–1397 CrossRef CAS PubMed.
- R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, J. Med. Chem., 2006, 49, 6177–6196 CrossRef CAS PubMed.
- K. Musilek, M. KomLoova, O. Holas, A. Horova, M. Pohanka, F. G. Moore, V. Dohnal, M. Dolezal and K. Kuca, Bioorg. Med. Chem., 2011, 19, 754–762 CrossRef CAS PubMed.
- G. L. Ellman, D. K. Courtney, V. Andreas and R. M. Featherstone, Biochem. Pharmacol., 1961, 7, 88–95 CrossRef CAS.
- H. H. Sun, J. Zhang, Y. Z. Zhang, L. Y. Yang, L. L. Yuan and Y. Liu, Mol. Biol. Rep., 2012, 39, 5115–5123 CrossRef CAS PubMed.
- P. Bourassa, S. Dubeau, G. M. Maharvi, A. H. Fauq, T. J. Thomas and H. A. T. Riahi, Biochimie, 2011, 93, 1089–1101 CrossRef CAS PubMed.
- H. X. Zhang and P. Mei, J. Solution Chem., 2009, 38, 351–355 CrossRef CAS.
- N. Chadha, A. K. Tiwari, V. Kumar, S. Lal, M. D. Milton and A. K. Mishra, J. Biomol. Struct. Dyn., 2014, 29, 1–13 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: ESI contains associated figures and tables. See DOI: 10.1039/c4ra14696d |
| This journal is © The Royal Society of Chemistry 2015 |