Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Xanthone dimers: a compound family which is both common and privileged†
Tim
Wezeman
a,
Stefan
Bräse
*ab and
Kye-Simeon
Masters
*c
aInstitute of Organic Chemistry (IOC), Karlsruhe Institute of Technology (KIT), Fritz-Haber-Weg 6, 76131 Karlsruhe, Germany. E-mail: stefan.braese@kit.edu; Fax: (+49) 721 6084 8581
bInstitute of Toxicology and Genetics (ITG), Karlsruhe Institute of Technology (KIT), Hermann-von-Helmholtz-Platz 1, D-76344 Eggenstein-Leopoldshafen, Germany
cDiscipline of Nanotechnology and Molecular Sciences, School of Chemistry, Physics and Mechanical Engineering, Faculty of Science and Engineering, Queensland University of Technology (QUT), GPO Box 2434, Brisbane, Queensland 4001, Australia. E-mail: kye.masters@qut.edu.au
First published on 16th September 2014
Abstract
Covering: up to 2014
Xanthone dimers are a widespread, structurally-diverse family of natural products frequently found in plants, fungi and lichens. They feature an intriguing variety of linkages between the component xanthones (benzannulated chromanones). These synthetically elusive secondary metabolites are of great interest due to their broad array of bioactivities, which has led to the xanthones being designated as ‘privileged structures’. We seek herein to give an overview of all reliably-described xanthone dimers, their structures, occurrence, and the bioactivities established to date. The possible biosynthetic pathways leading to members of this family are also discussed in light of our current knowledge.
1 Introduction
Xanthones comprise a group of structurally diverse, biosynthetically intriguing, biologically active and synthetically challenging natural products. Many xanthones have been found to exhibit pronounced biological activities, for example anti-tumour effects.1–5 Xanthone chemistry is satisfyingly rich, with the conjugated donor–acceptor motif of the central B-ring ensuring that these compounds display a degree of personality greater than their apparently simple core structure might suggest.The xanthone family has been studied for over a century since the early 1900s6,7 and has since been investigated and reviewed, particularly for the monomeric species and plant-derived species.8–14 Several reviews on xanthones have been published in recent years, covering xanthones from fungi,15 lichens and bacteria,16 structure–activity relationships,2 antimalarial properties of xanthones,17 biological activity,18 chemical synthesis of xanthone cores,19–21 and their biosynthesis.22–27 Nonetheless, the field of xanthone chemistry is growing rapidly; every few years see the publication of an update on novel naturally-occurring xanthones.8,21,28–30 Analytical techniques have been optimised specifically for xanthones, including crystallography31 and chromatographic methods.32 The structure, activity and synthesis of polycyclic, yet monomeric xanthones has recently been reviewed,33 as well as the specific class of the caged xanthones.34,35 Most of these articles focus mainly on monomeric xanthones, and to date none have focused on dimeric and trimeric xanthones.
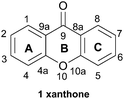
For the sake of simplicity and easy comparison between structural features, structural numbering will follow the monomeric parent xanthone core (1, Fig. 1) throughout this review.
2 Biosynthesis
The biosynthesis of dimeric xanthones is a topic which stands in need of further research progress, especially the identification and description of the key dimerisation processes (see also ref. 15 and 16). As the biosynthesis of the dimeric structures is a point of considerable interest and remains largely unknown, we will give a necessarily brief outline of the achievements so far – the full elucidation of the biosynthesis is no doubt just over the horizon.2.1 Monomer xanthones
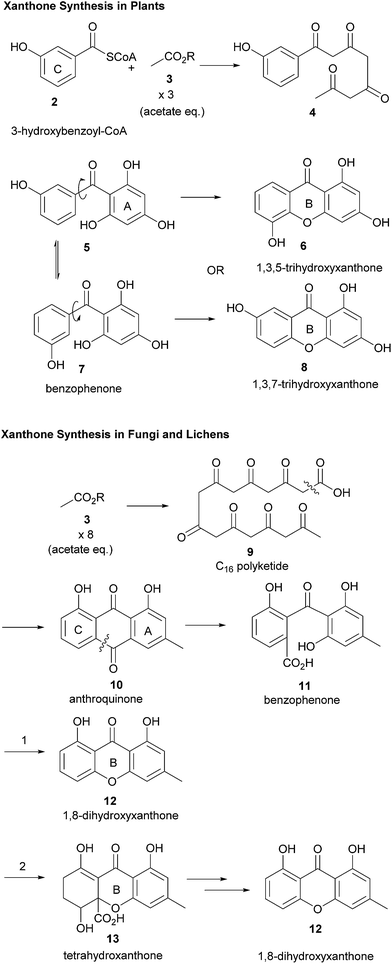
It has been known for some time that the biosynthesis of the monomeric xanthone core proceeds differently in fungi25,36 and higher plants (Scheme 1).37,38 The synthesis of the xanthones in fungi has been studied in detail since the radiolabelled acetate feeding experiments of Birch in the 1950s, showing that polyketides are the biosynthetic precursors of the xanthone core for fungi. More recently, Simpson has reported findings resulting from the sequencing of the Aspergillus nidulans genome.39 The sequencing revealed the presence of 32 clusters containing polyketide synthase (PKS) genes, which were studied by gene deletion to reveal information about the biosynthetic pathway which converts emodin via key steps of oxidative ring-scission and decarboxylative cyclisation to xanthones and eventually prenylated and cyclised products such as shamixanthone, tajixanthone and sterigmatocystin. As part of a comprehensive study on the formation of aflatoxin in Aspergillus species, Townsend and co-workers described the generation of xanthone monomers through a complex sequence of epoxidation, rearrangement, deoxygenation, Baeyer–Villiger oxidation, and further deoxygenation leading to the tricyclic xanthone core of sterigmatocystin.40 Interpretation of the sequences may differ markedly41 in terms of the sequence of events, and the structural homology of biomolecules involved in the pathway in differing species may not, in fact, be a reliable method for determining correlations between pathways.Müller and co-workers have recently discussed the importance of reductive steps in the synthesis of xanthones commencing from (polyketide-derived) anthraquinone, emodin, and passing through chrysophanol.42 Contrastingly, the biosynthesis of xanthones in plants results from a convergent synthesis of polyketide and shikimic acid pathways, whereby 3-hydroxybenzoyl-CoA (obtained from an early shikimate pathway intermediate) is condensed with a polyketide, which may be then cyclised to form the C-ring and then the B-ring.26 A consequence of these two distinct pathways, both of which pass through a freely-rotatable benzophenone intermediate, is that the pattern of oxidation between xanthones from plants and fungi frequently differs. Fungal xanthones (or their biosynthetic precursors) almost ubiquitously display C-1 and -8-hydroxylation, whilst plants frequently display C-1, -3 and -5 or -7-hydroxylation.
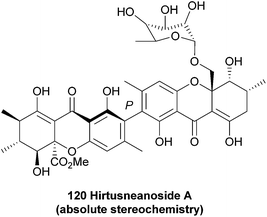
In terms of further diversification of the xanthone core, some processes are now understood better than others. The investigations of Elix and co-workers have led to a fuller understanding of some of the pathways of further structural elaboration of the xanthone nucleus in lichens. Particularly, cladistical/phylogenetic analysis of related species was studied to develop a fuller understanding of the relationship between enzymes responsible for methylation and chlorination.43,44 Prenylation of the dimers is frequently found in plant-derived xanthones.45 Pendant sugars are also found as a structural feature of dimers, see for example puniceaside C (72, plant derived) and Hirtusneanoside (120, lichenoid). It is also known that some substituents are removed from the xanthone core, such as the reduction of hydroxyl functions and reductive dearomatisation.
2.2 Xanthone dimerisation
There are a variety of ways in which dimeric and trimeric xanthones can be linked: (i) a rotatable or atropisomeric biaryl C–C bond or (ii) a biaryl ether C–O–C linkage. Also, because xanthones, particularly from plants, can be decorated with prenyl groups, the linkages are also often in the form of (iii) aryl–O–alkyl linkages, or (iv) prenyl derivative–prenyl derivative linkages. The incorporation of nitrogeneous bases to form polyaromatic antibiotic xanthones is also frequently observed, and readers are directed to the recent review by Porco and co-workers on this subject.33One of the more common ways in which dimerisation is found in the xanthones is through the biaryl linkage. Regio- and stereo-selective biaryl linkage of phenolic compounds is common in plants, bacteria, lichen, and fungi – as Bringmann and co-workers noted in their seminal work on natural biaryl linkages, “wherever in Nature phenolic aromatics can be found… the corresponding homo- or hetero-dimeric biaryls have to be expected”.46 Certainly, the discovery of the blennolides (hemisecalonic acids)47 in 2008 supported the putative dimerisation of discrete monomers as the likely method of dimer formation, rather than a more complex and difficult to imagine tandem biosynthesis pathway – the side-by-side cyclisation of a double-length polyketide to form e.g. a secalonic acid seems unlikely. Despite efforts in the detailed study of the biosynthesis of dimeric xanthones, particularly of the secalonic acids by Frank and co-workers,23,40,48–50 the dimerisation of xanthones remains a tricky subject for both the synthetic and biosynthetic chemist – the extent of the involvement of enzymes, and their nature, is at present unclear. There has been, at the time of writing, no direct observation of enzymatic dimerisation of the monomer units.
Enzymatically-mediated or not, the oxidative pathway which is most commonly invoked is shown here (Scheme 2), whereby single electron transfer deprotonation of a hydroxyxanthone, 14, leads to an intermediate which can be readily oxidised by single-electron-transfer to xanthonyl radical 16. Resonance contributors of the delocalised aryl radical, 16a–c, can then couple to electron-donors, notably the ortho and para-C positions.
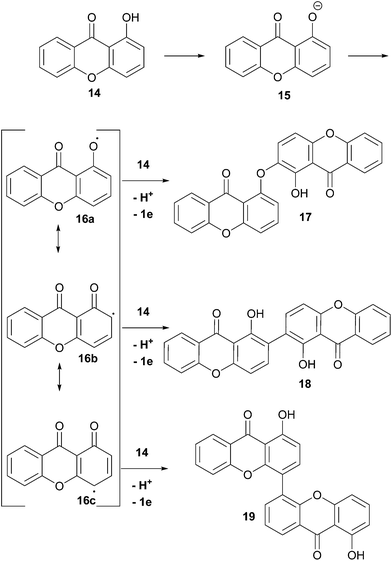 | ||
| Scheme 2 Phenolic dimerisation reactions adapted for xanthones. For clarity, coupling of 16a–c at the ortho-position of 14 is shown. | ||
2.3 Regiochemical evidence for enzyme dimerisation
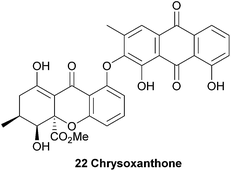
The resulting biaryl bond linkages are formed which, in the absence of an enzyme or other structurally-influencing substrate, is determined largely by the spin-density distribution of the radical generated, and presumably in some cases other inherent stereoelectronic characteristics of the substrate molecule. As Kozlowski has put it, in the absence of external factors, “the substrate typically dictates the available coupling products”.51 This leads us, as chemists, to an interesting corollary – if there are differing regio- and atropo-isomeric biaryl-linked dimers of the same substrate to be found in Nature, then it must be the case that there are external forces at play in their biosynthesis to lead to this divergence, rather than ‘spontaneous’ oxidative dimerisation with oxidants such as intracellular O2. Prominent examples supporting this line of thinking can be seen in Scheme 3, where both secalonic acid B/chrysoxanthone and secalonic acid E/talaroxanthone are presumably generated with alternative (chemo- and regio-) selectivity from the monomer units, hemisecalonic acids B and F, respectively. The oxidative dimerisation of the xanthones is thus most likely mediated by enzymes, as shown for a number of model compounds.52–54 It can be mimicked by chemical oxidases.55 In some cases, the dimerisation led to the formation of diaryl ethers (e.g. chrysoxanthone 22, Scheme 3) instead of to a C–C bond formation.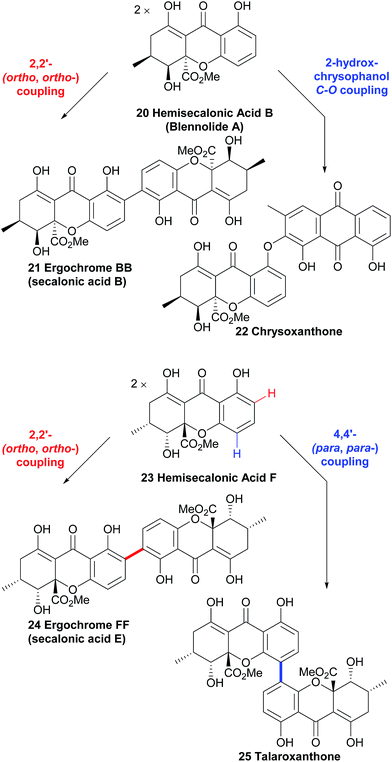 | ||
| Scheme 3 Multiple pathways from the same monomer units strongly suggest the involvement of enzymatic processes in xanthone dimerisation. | ||
Hemisecalonic acid B47 is dimerised at the ortho-positions to form secalonic acid B 21 (Scheme 3; see below for more information) in a variety of fungal species.49,148–150,153 From the same substrate in the case of the xanthone anthraquinone chrysoxanthone, the coupling of the hemisecalonic acid B with a 2-hydroxy variant of chrysophanol, a widespread polyketide natural product, dimerisation leads to the C–O bond formation of diaryl ethers i.e. chrysoxanthone.56 Another example of biosynthetic divergence is likely involved in the regioisomeric selectivity which leads to the formation of talaroxanthone 25, a 4,4′-linked xanthone dimer (from the endophytic fungi Talaromyces sp.)57 and secalonic acid E 24, a 2,2′-linked xanthone dimer (from e.g. Phoma terrestris).58,59 That a statistical mixture of the three regioisomers is produced by each of the organisms in question and the two differing regioisomers have been selectively isolated by the research groups in each case is highly unlikely (see Scheme 3). The selective discovery of these regioisomers can therefore be considered indirect evidence for the active role of yet-to-be discovered enzymes.
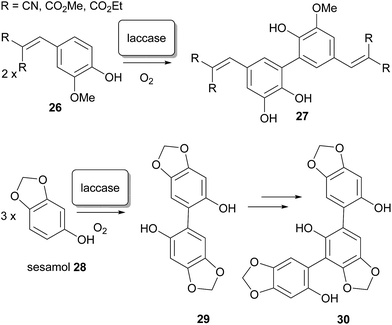
In the laboratory, regioisomeric control over the biaryl bond formation60–62 is often difficult to impose, and frequently requires innovative solutions.63–66 In natural biosynthesis, it has been demonstrated that selectivity between 2,2′- 2,4′- and 4,4′-coupling can presumably be imposed by enzymes, although little direct evidence exists for this – no enzyme has yet been identified which fulfils the role of regioselective differentiation in the dimerisation of xanthones in Nature. Nonetheless, high selectivity for such processes has been demonstrated with enzymes in the laboratory; oxidative dimerisation at a specific ortho-position to form ferulic acid derivatives has been reported by Beifuss and co-workers, and has been achieved by application of the (single electron) oxidase enzyme67,68 laccase,69 from Trametes versicolor, in the presence of O2.70 Additionally, selective coupling at the C-6 position of sesamol led to dimer intermediates and then trimers in good yield.71 The outcome was markedly different than in the application of laccase from Agaricus bisporus, and also from electrochemical non-enzymatic oxidative coupling of this substrate by the method of Waldvogel and co-workers.72,73 Aside from their potential relevance to xanthone dimerisation, the synthetic application of enzyme-catalyzed transformations has significant additional benefits: they can be performed in aqueous solvent systems and under mild reaction conditions, a remarkably broad substrate spectrum that can be expanded even more by using mediators (Scheme 4).74
It is therefore logical to support the assertion of Bringmann and co-workers that the dimerisation of phenolic aromatics (both C–C and C–O coupling) is through “ – mostly – enzymatically-assisted biogenetic pathways”.41 Based on related processes in the chemical literature on both natural products75–79 and synthesis,80–82 one possibility is that a laccase is responsible. Another possibility is a monooxygenase, such as cytochrome P450; evidence for a likely mechanism for stereoselectivity is also found in this case.
2.4 Stereochemical features of the biaryl xanthone dimer
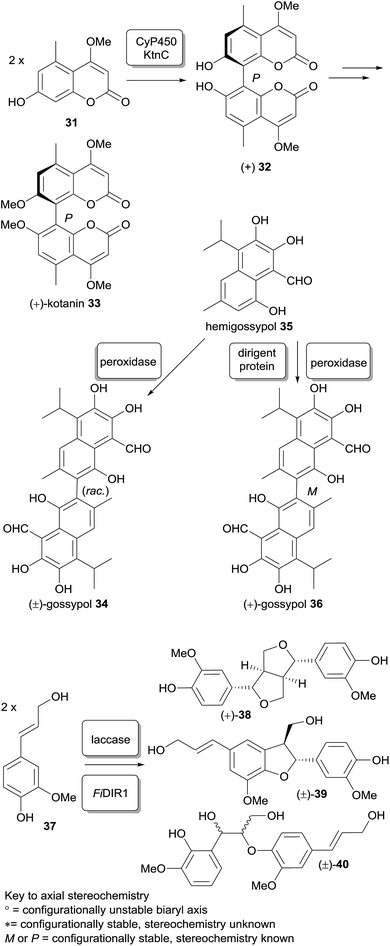
Substituted biaryls exhibit varying degrees of barrier to rotation, ranging from freely rotating (microsecond time-frame) through slow room-temperature interconversion to atropisomers which may be isolable and thereafter stable; dimeric xanthones show this feature also. Stereochemical biaryl bond formation is displayed in the secondary metabolism of bacteria, fungi, lichens and plants, and has herein been denoted alongside the structures of biaryl-linked dimers utilising the notation of Bringmann and co-workers (see Scheme 5).44 As both chiral and meso-dimers have been found in the secalonic acid series, various mechanisms for the origin of the dimers can be invoked. The reasons for this could certainly be due to the different biosynthesis routes employed between the classes of species. In fungi, all xanthone dimers display a biaryl xanthan–xanthone bond between one or more di- or tetra-hydro xanthones. It could possibly be argued that the dimerisation process could be influenced by the asymmetrically-substituted C-ring, a possible explanation for non-enzymatic axial chirality generation. However, the fully aromatic xanthone dimer ploiarixanthone 68, which is optically active ([α]25D = +23°),83 must involve axial-chirality influencing biomolecules in the stereospecific dimerisation step.The involvement of stereoselectivity-mediating proteins may prove to be the origin of stereospecific pathways in the formation of dimeric xanthones. One case where the explanation for the regio- and stereo-chemical control observed in a biphenolic coupling has been uncovered is the mechanism of selective phenol coupling in Aspergillus niger by Müller and co-workers.54 They showed that an oxidative phenol coupling is also the key stereo-divergent step in the formation of P-(+)-kotanin (33, Scheme 5) from two equivalents of coumarin 31 catalyzed by cytochrome P450 monooxygenase KtnC, which was identified by targeted gene deletion in the A. niger strain. The facially-selective interaction of the two monomer subunits with the heme-containing active site of the CyP450 was modelled in silico by homology studies and docking of both the substrates and product. Stipanovic and co-workers showed enzymatic coupling of hemigossypol (35, Scheme 5); this compound by in vitro application of a pure peroxidase only led to a racemic product (±)-gossypol, whereas the addition of a cotton-plant dirigent protein gave predominantly (+)-(S)-gossypol.84a Naturally-occurring gossypol is found as a mixture of (+) and (−) isomers, which have been found to exist in a continuum of ratios between 68 and 2% (−)-(R)-gossypol, depending on the strain of the cotton plant G. barbadense,84b possibly indicating a variation in the type or extent of dirigent protein control in the coupling. Stereoselective phenol coupling of 37 resulting in C and O–alkylation products (+)-38 or racemic 39 and 40, reminiscent of the kind of transformations of some linked and/or cyclized dimeric xanthones, has also been achieved with a combination of laccase and dirigent protein FiDIR1.85 It may be the case that for xanthones there are also mediator proteins associated with oxidase enzymes involved in the stereoselective generation of the biaryl bonds.
The absolute stereochemical assignation of regioisomerically identical xanthone dimers isolated from different biological sources is, unfortunately, in many cases incomplete. X-ray structures have been used to determine the relative and absolute stereochemistry of the stereochemical centres and axes (e.g. for phomoxanthone A 106 and phomalevone A 103).86 In some cases, however, modern DFT calculations have allowed CD spectra to be computed which were used for comparison with experimental data (blennolides).47 In particular, the quantitative consideration of vibronic effects, conformer equilibria and solvent effects were included in most of the models.87 As biological activity and their biological targets are different for different diastereomers (e.g. compare secalonic acid A and E), it is very important to address stereochemical issues of dimeric xanthones. Unfortunately, many data are not available (due to the lack of instrumentation and/or authentic material).
2.5 The true biological sources of xanthone dimers?
In some cases researchers have reported dimeric xanthones as being isolated from endophytes.57,86,88–90 Certainly culture broth production of dimeric xanthones has demonstrated that this class of molecules can be produced by fungi alone. This raises the question for ‘plant-derived’ compounds: are the plants really the true producers, or fungi? Is there a mixed origin, or is it really only the (undetected) fungi which produce the xanthone dimers?For monomeric xanthones, the plant–fungus contributions have been at least partially unveiled on at least two occasions. In its biosynthesis, the monomeric xanthone 2,7-dichlorolichexanthone (not shown) was isolated from samples of the lichen Lecanora dispersa. In cultures of the fungal species in the absence of the alga, the xanthone biosynthesis was halted.91 In an example for xanthone dimers, the eumitrins (see below) are produced by the lichen (algae-fungal symbiotic organism), and not by only the fungus in isolation.92 It is conceivable that in some situations that involve plant-microbe combinations, the biosynthesis may even involve a to-and-fro sequence of enzymatic transformations carried out by each organism. Such a scenario, if it does exist, is likely to be complex and case-specific.
3 Xanthone dimers from plants
Xanthone dimers from plants, like the monomers themselves, commonly feature prenylated and multi-prenylated cores. The presence of these many prenylated monomeric xanthones shows that prenylation occurs readily prior to dimerisation, although there is no evidence to suggest that it does not also occur post-dimerisation.3.1 Bijaponicaxanthones
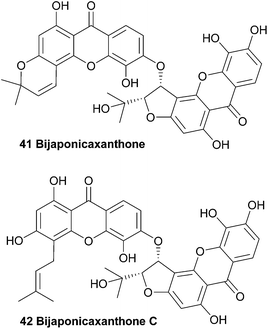
Several interesting xanthones were isolated from the ethanol extracts of the dried aerial parts of Hypericum japonicum, a Chinese medicinal plant, including one dimeric xanthone named bijaponicaxanthone (41). By comparing the NMR spectra with known compounds such as the monomeric xanthone isojacareubin (not shown), and by analysing additional 2D NMR spectra, its structure could be assigned. The relative configuration between the two chiral centres was determined to be cis for the two protons, based on their coupling constant.93–95In 2005, a similar dimeric xanthone was isolated from Hypericum japonicum. After detailed structural analysis the xanthone was found to be prenylated instead of having a fourth ring and the natural product was named bijaponicaxanthone C (42).96 Bijaponicaxanthone C was also reported to have been isolated from the roots of the Hypericum riparium plant.97
3.2 Bixanthones
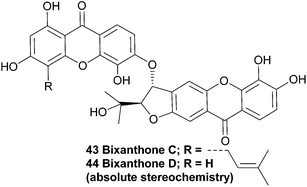
The structurally very similar bixanthones C and D (43, 44), as reported in a Chinese patent, were also isolated from Hypericum japonicum. The patent claims these compounds are active ingredients in a traditional Chinese medicine which is used to treat a variety of liver diseases, and describes the use of the flavone component of that plant species as a medicament for treating hepatic fibrosis.98 These compounds are missing OH groups on the furannulated xanthone component, presumably due to enzymatic reduction.3.3 Bigarcinenones
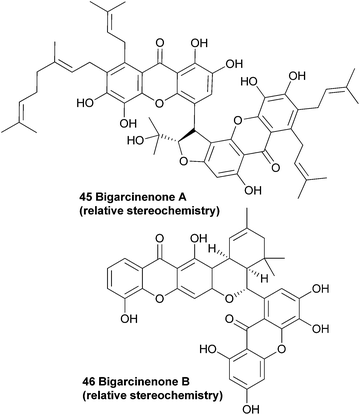
Since the plant Garcinia xanthochymus is widely used as traditional Chinese medicine, Yang et al. investigated the extracts from the bark of the plant. They observed a strong antioxidant activity (IC50: 4.6 μg mL−1, as determined by a 1,1-diphenyl-2-picrylhydrazyl (DPPH) radical scavenging bioassay) in the ethyl acetate soluble fraction from the ethanol extracts, and proceeded with a more detailed study on the contents of these extracts. Along with seven known xanthones, they found a novel bisxanthone called bigarcinenone A (45).Using mainly 2D-NMR techniques, the authors identified the dimeric xanthone to be coupled through a side-chain of one of the two subunits. Bigarcinenone A performed relatively well in a DPPH radical scavenging activity assay (IC50: 9.2 μg mL−1), outperforming all other isolated xanthones from this extract (IC50: 16.3–250 μg mL−1) as well as BHT (IC50: 20.0 μg mL−1), a well-known synthetic antioxidant.99 In 2011, bigarcinenone B was isolated from the bark of the same plant and its structure and relative stereochemistry was also elucidated using mainly 2D NMR techniques such as HMBC, HSQC and ROESY. A possible biosynthetic pathway was proposed by the authors that would lead to the unique connection through the two six membered rings. A Diels–Alder reaction between the two prenyl groups of the monomers would lead to the cyclohexene derivative, which would undergo a second cyclisation to form the final ring. The antioxidant activity of bigarcinenone B (7) was tested with a DPPH assay (IC50: 20.14 μM versus 13.16 μM for ascorbic acid) and a H2O2 assay (IC50: 2.85 μM versus 0.76 μM for ascorbic acid).100
3.4 Chiratanin
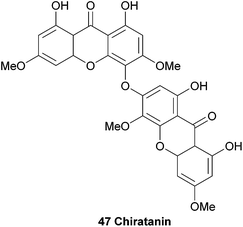
An interesting biaryl-ether linked dimeric xanthone named chiratanin (47) was isolated together with several monomeric xanthones and other secondary metabolites from the benzene extracts of the plant Swertia chirata.101 At the time, this was the first report of the occurrence of a dimeric xanthone in a higher plant. After detailed analysis of the NMR spectra, the authors identified the positions of the oxygen atoms in the molecule, and with D2O exchange experiments all the phenolic hydroxyl groups were identified. However, the actual connection of the two xanthones remained difficult to determine, and three possible structures were identified. The molecular asymmetry was instructive in determining the final structure, which they named chiratanin.3.5 Cratoxyxanthone
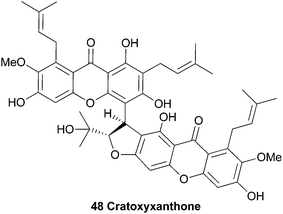
Several natural products were isolated from the bark of Cratoxylum cochinchinense in 1995 by Sim et al.102 After extensive HPLC purification of the isolated compounds, detailed high field NMR spectroscopy was used to identify a novel dimeric xanthone, named cratoxyxanthone (48).The authors speculated on a biosynthetic pathway involving the coupling of two mangostin-derived radicals, as depicted in Scheme 6. Cratoxyxanthone was also isolated from the chloroform soluble extracts of the stem bark of Garcinia mangostana in 2009.103 In this study, it was found that cratoxyxanthone had a very poor in vitro activity in cytotoxicity (HT-29 cell line) and ELISA NF-κB (p65 and p50) assays.
3.6 Garcilivins A–C
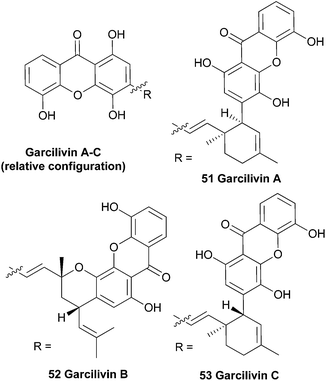
Bark from the roots of the South African plant Garcinia livingstonei was phytochemically investigated in a larger study aiming to find reversible monoamine-oxidase inhibitors for antidepressant drugs. This study resulted in the discovery of several prenylated xanthones, including three dimeric xanthones named the garcilivins (51–53).104 The structures were elucidated using extensive mass and NMR spectroscopy and by comparing with monomeric xanthone spectra.105 In 1992 it was reported that these xanthones were currently being assayed for their biological activities, but as far as the authors know, no follow-up paper was published until Pieters et al. in 2006.106 They report the isolation of several xanthones and flavonoids from a Tanzanian G. livingstonei and tested the isolated compounds for their antiparasitic activity and cytotoxicity. Garcilivin A and C showed a very interesting difference in their toxicity tests, considering they are diastereoisomers. Both compounds were tested against four parasites and garcilivin A showed a very strong non-selective activity in all the assays, whereas garcilivin C only showed significant activity against Trypanosoma brucei. Especially noteworthy is the score for cytotoxicity against MRC-5 cells of 2.0 μM for garcilivin A and 52.3 μM for garcilivin C.3.7 Garciobioxanthone
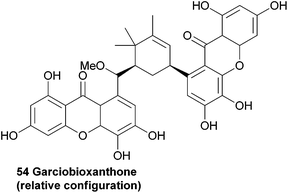
Recently, the novel garciobioxanthone (54) was isolated from the ethanol extracts from the bark of Garcina oblongifolia, along with several known compounds. Using 2D NMR techniques the relative configuration and structure was determined.1073.8 Globulixanthone E
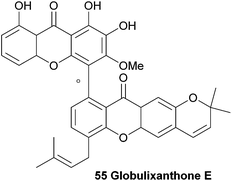
Three prenylated xanthones were isolated from the root bark of the large forest tree Symphonia globulifera and named globulixanthone C–E. The tree is widely used in Cameroon as a medicinal plant and laxative for pregnant women. Globulixanthone C and D were found to be monomeric xanthones, but globulixanthone E was found to be a bisxanthone and its structure (55) was elucidated using ESI-TOF MS, IR and NMR spectroscopic methods. Using NOESY spectra and by comparing the obtained spectra with the other isolated xanthones, the structure was determined. The three xanthones were tested for their in vitro antimicrobial activities against Staphylococcus aureus, Bacillus subtilis, Vibrio anguillarium and Escherichia coli in an agar well diffusion assay. The monomeric xanthones showed a moderate activity to S. aureus (MIC: 8.05–14.05 μg mL−1) and B. subtilis (MIC: 8.24–12.5 μg mL−1), but globulixanthone E showed a remarkably high activity against all tested organisms (MIC: 5.56–3.12 μg mL−1) except E. coli, and outperformed the positive control in the case of S. aureus (MIC: 4.51 μg mL−1versus 6.25 μg mL−1 for streptomycin sulfate).1083.9 Griffipavixanthone
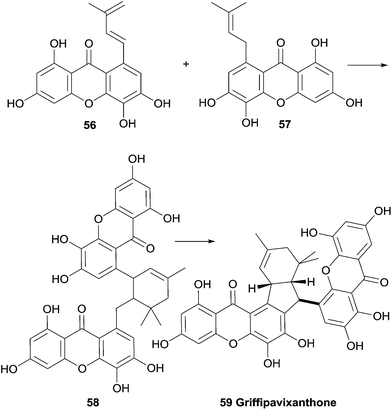
An investigation into the secondary metabolites of the Malaysian plants Garcinia pavifolia and G. griffithi lead to the discovery of a new bisxanthone named griffipavixanthone.109 Using advanced spectroscopic methods, including NOESY, HMBC and INEPT NMR studies, the relative structure was elucidated. In a later study, the bisxanthone was also found in G. maingayii.110 Griffipavixanthone was the first example of a dimeric xanthone that is connected via a 5 and a 6 membered ring.109 The authors suggest a possible biosynthetic pathway involving an initial Diels–Alder reaction of two prenyl groups on the two xanthones (Scheme 7). The resulting cyclohexene is then followed by another cyclisation, either ionic or radical, to yield the 5-membered ring. The bisxanthone was also isolated from Garcina oblongifolia107 (along with several other xanthones, such as garciobioxanthone) and G. virgate111 (along with two new monomeric xanthones named virgataxanthone A and B, not shown). The isolated compounds in the latter study were tested for antioxidant capacity, and it was noted that griffipavixanthone showed a notably high radical scavenging ability, with an EC50 lower than the references used in the study (EC50: griffipavixanthone: 11.5 μg 100 mL−1, 2,6-di-tert-butyl-4-hydroxy-anisol: 13.6 μg 100 mL−1, α-tocopherol: 13.8 μg 100 mL−1). In in vitro cell line cytotoxicity tests griffipavixanthone showed high activity against P388, LL/2 and Wehil64 cell lines with very promising ED50 between 3.40 and 6.80 μg mL−1.109 A Chinese patent reports promising anti-cancer properties for griffipavixanthone. They claim a strong inhibition effect of the bisxanthone or its salts on human lung, breast, prostatic and intestinal cancer cells, while showing no cytotoxicity to normal kidney epidermal cells. For lung cancer cells H520 in particular, they found that the cell cycle was blocked in the S stage, thereby preventing propagation of the cancers.1123.10 Hyperidixanthone
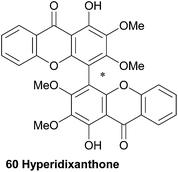
Hyperidixanthone (60) is a symmetrical dimer isolated from the plant Hypericum chinense.113 The compound was identified by a barrage of NMR techniques, including HMBC, and mass spectral data. Although hyperidixanthone is almost certainly axially chiral, the authors reported no details of optical activity.3.11 Jacarelhyperols A, B and D
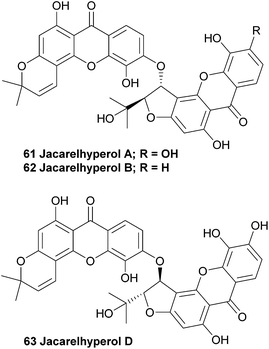
Jacarelhyperols A and B (61, 62) were isolated from Hypericum japonicum only a few years later than the bijaponicaxanthones, and the measured NMR spectra were found to be very similar to the spectra obtained from the bijaponicaxanthones. The relative structure of jacarelhyperol A was elucidated to be an epimer of bijapoincaxanthone with a trans-configuration of the two protons at the chiral centres. Jacarelhyperol B was subsequently identified as a 6′-dehydroxy variant of jacarelhyperol A. Both jacarelhyperols A and B were found to significantly inhibit platelet-activating factor (PAF)-induced hypertension in mice at 10 mg kg−1 without causing their blood pressure to rise. Therefore PAF inhibitors are considered to be potential drugs against allergic diseases.114 The isolation and structure elucidation of jacarelhyperol D from the same plant species was reported in 2007 in a short communication paper by Chen et al.1153.12 Mesuabixanthones A and B
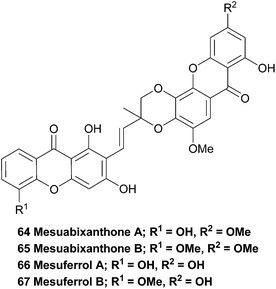
The bark of the south-east Asian tree Mesua ferrea has been widely used in traditional local medicine and is reported to exhibit antimicrobial and antiasthmatic activity. Therefore, the stem bark was investigated in an attempt to isolate the active compounds. Interestingly, the two novel bisxanthones mesuabixanthone A and B (64, 65) were isolated from an extracted fraction that showed little biological activity in their initial bioassays. Using conventional mass spectrometric and NMR spectroscopic methods they identified the two novel bisxanthones. Mesuabixanthone B differed from mesuabixanthone A only by the methylation of the hydroxyl group at C-8, as determined by a NOESY experiment. No biological testing on the isolated compounds was reported.1163.13 Mesuferrols A and B
The very similar mesuferrols A and B (66, 67) have been isolated from an acetone extract of the bark of the same tree and were purified using chromatography.117 Mesuferrol A and B were identified by means of NMR spectroscopy and HR FAB-MS.3.14 Ploiarixanthone
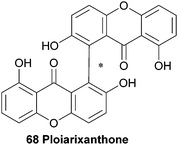
In 1990 a novel dimeric xanthone metabolite was isolated from branches of the shrub Ploiarium alternifolium. By synthesising a tetracetylated derivative and detailed comparison of the obtained NMR spectra with the data from the natural product, they identified ploiarixanthone (68) to be 8,8′-linked.83 Ploiarixanthone is optically active ([α]25D = +23°), providing strong evidence for an enzymatic dimerisation process in its formation.3.15 Puniceasides
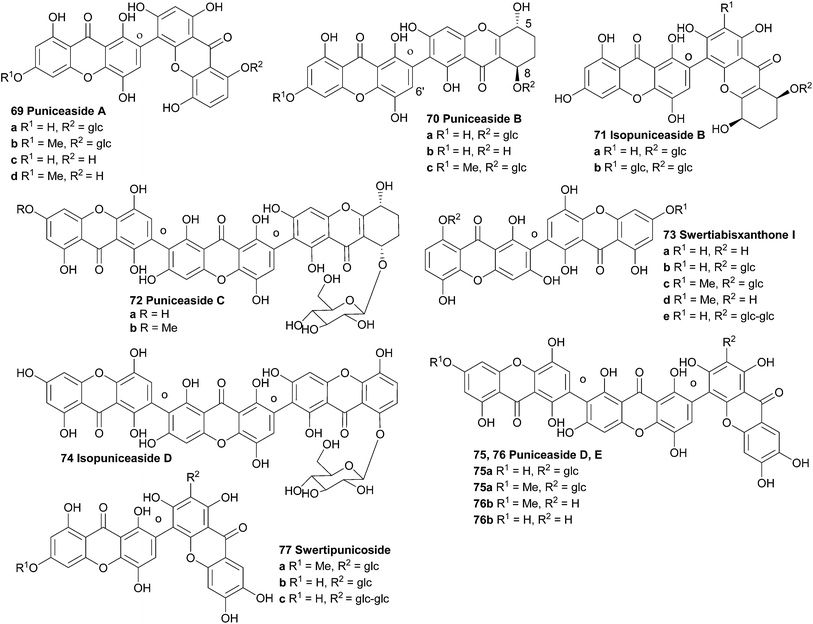
In 1991 the first isolation and structure elucidation of a bisxanthone C-glucoside named swertipunicoside (77a, Fig. 2) was reported. It was isolated from extracts from the whole plant Swertia punicea and the structure was confirmed by NMR experiments, particularly the selective INEPT technique (Insensitive Nuclei Enhanced by Polarisation Transfer).118 Only one year later the same group was also able to isolate and characterise 3-O-demethylswertipunicoside (77b) using several NMR techniques including APT, HETCOR and selective INEPT.119 Puniceasides B and C and isopuniceaside B have all been partially reduced to feature one tetrahydroxanthone core.Recently, investigations showed that 3-O-demethylswertipunicoside has potent neuroprotective capabilities. Several oxidative toxicity tests were performed on xanthone treated and untreated PC12 cells. Increased cell viability was found in a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) cell death assay where the cells were exposed to 1-methyl-4-phenylpyridiniumion (MPP+), rotenone or hydrogen peroxide. These neuroprotective effects were found to be caused by the elevation of TH and DJ-1 protein levels.120
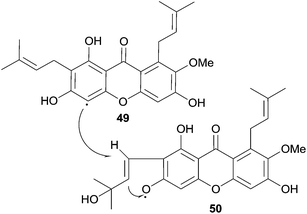
Hostettmann et al. isolated several xanthones from Gentianella amarella ssp. acuta, including one new dimeric xanthone. After structure elucidation with the use of 2D-NMR experiments and mass spectrometry they named the compound swertiabisxanthone I 8′-O-β-D-glucopyranoside (73b).121
Remarkably, its aglycone, swertiabisxanthone I (73a), was first isolated and identified more than twenty years earlier from Swertia macrosperm plants.122 Recently both the glycone and the aglycone of swertiabisxanthone I have also been isolated from Swertia punicea.123 In 2010, Guo et al. isolated two new dimeric O-glycoside xanthones, one new trimeric O-glycoside xanthone, two new trimeric C-glycoside xanthones and 12 known xanthones from the plant Swertia punicea.123 The five new xanthones were identified using extensive HRESIMS and NMR spectroscopic experiments and were named puniceasides A–E (69a, 70a, 72a, 75a, 76a). For puniceaside B the cis configuration between H-5 and H-8 was confirmed using NOESY experiments. Also, it was found that the H-6′ resonance pattern indicated that two different rotameric forms of puniceaside B exist. By heating to 120 °C in a 1HNMR experiment, the two singlets of H-6′ merged into one singlet. Puniceaside C was confirmed to be a trimeric xanthone glycoside by close examination of the results from mass spectrometric methods and HMBC NMR experiments. As far as is known, this was the first trimeric xanthone reported in the literature. By subjecting the compound to acid hydrolysis, the glycosidic moiety was found to be D-glucose. Two more trimeric xanthones were identified as puniceasides D and E. HMBC correlations showed the presence of the OMe group in puniceaside E. The absolute configurations of 75a and 76a could not be determined due to insufficient quantities of compounds. The presence of different rotameric configurations is likely, since heating the 1H NMR samples to 120 °C resulted in the merging of the two singlets of H-6′ into one singlet, similar to observations for puniceaside B. Puniceasides A–E, swertiabisxanthone I 8′-O-β-D-glucopyranoside, 3-O-demethylswertipunicoside and swertipunicoside were tested for their neuroprotective activity against hydrogen peroxide-induced PC12 cell damage. In particular puniceaside B was found to be very potent with a cell viability of 98.1 ± 6.8% at a concentration of 25 μg mL−1 compared to hydrogen peroxide treated cells. Interestingly, swertiabisxanthone I 8′-O-β-D-glucopyranoside and 3-O-demethylswertipunicoside were found to potently stimulate the damaged PC12 cells to grow, resulting in cell viability scores of 123% and 158%. In 2012, Guo et al. reported the characterisation of 16 new xanthone compounds from extracts of Swertia punicea. After studying the ESI-MS fragmentation behaviours of 17 known xanthones very closely, they could apply this knowledge to high-performance liquid chromatography diode-array detection/tandem mass spectrometric results and thus identify 11 new dimeric and 4 new trimeric xanthones (69b–d, 70b and c, 71a and b, 72b, 75b, 76b, 74, 73c–e, 77c).124
4 Xanthone dimers from fungi
Fungi have yielded some beautiful xanthone dimers, the majority of which possess a common biaryl bond as the method of their linkage. The biaryl bond is most often arranged in a 2,2′-manner with respect the common xanthone hydroxyl moiety; in a couple of cases, natural products with 2,4′-biaryl bonds have been described, as has the somewhat rarer 4,4′-biaryl linkage. Additionally, the position of methylation of the xanthone core is found at the C3, C6 or sometimes C10a positions, which can provide insight to the biosynthetic origin of the compounds (see Table S-1† for structural details).4.1 Ascherxanthones
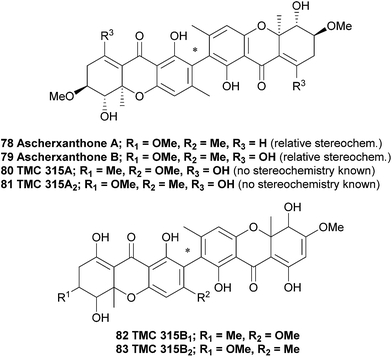
Ascherxanthone A (78) was isolated from the methanol extract of the mycelia of Aschersonia sp. BCC 8401, after significant antimalarial activity in Aschersonia cultures was observed.125 Evidence for the symmetric homodimeric structure of ascherxanthone A was provided by mass spectrometry and the presence of only 16 carbon signals in the 13C NMR spectrum. The tetrahydroxanthone structure was established on the basis of the HMBC correlations, and using COSY and HMQC NMR spectroscopic methods the entire structure was elucidated. Intense correlation from the phenolic proton to the quaternary carbon placed this carbon at C2, confirming a 2,2′-biaryl-linked dimer. Interestingly, the structure of ascherxanthone A is very close to TMC 315A1, TMC 315A2, TMC 315B1 and TMC 315B2 (80–83); four dimeric xanthones that were isolated from Ceuthospora sp. TMC1678 and reported in a Japanese patent to be RANKL (receptor activator of NF-κB ligand) antagonists that can prevent or control osteoporosis.126 It should be noted the Japanese patent does not provide any stereochemical information for TMC 315A2 (81), which may therefore be in fact identical to ascherxanthone B (79). Note that ascherxanthone A bears one less hydroxyl than the other compounds in this family, and one component of each TMC 315B1 and B2 has been reduced to the relatively rare dihydroxanthone tricycle.In a search for new agents against the rice blast fungus Magnaporthe grisea that has developed a fungicide resistance, culture broth extracts from approximately 800 fungal strains were investigated in an in vitro screening assay. A culture of Aschersonia luteola BCC 8774 was found to produce ascherxanthone B, a compound active against M. grisea with an IC90 value of 0.58 μg mL−1 The structure of ascherxanthone B (79) was elucidated using COSY, HMBC and HMQC NMR experiments and found to be almost identical to ascherxanthone A, with the only difference being the replacement of the olefinic proton at δH 6.87 (H-8) in ascherxanthone A with a chelated hydroxy resonating at δH 13.43 (C-8 hydroxyl, broad singlet). Interestingly, it was found that ascherxanthone A was nearly inactive in the same biological assay against M. grisea with an IC90 value of over 50 μg mL−1.127 However, ascherxanthone A did show significant activity against Plasmodium falciparum K1 (IC50 = 0.20 μg mL−1). Furthermore, it also showed cytotoxicity to Vero cells (IC50 0.80 μg mL−1) and three cancer cell lines (IC50 values between 1.7 and 0.16 μg mL−1).125
4.2 Chrysoxanthone
Chrysoxanthone, although not a xanthone dimer, is included here as an unusual example of the mixed-biological derivative ‘xanthraquinones’, which also include the beticolins128–131 and the xanthoquinodins.132–135 Chrysoxanthone, as reported by Anke and co-workers,56 features a form in which 2-hydroxychrysophanol is coupled to blennolide A (hemisecalonic acid B) through a biaryl ether linkage. The authors determined the novel structure with 2D NMR and mass spectral techniques, and it was found to possess antimicrobial and antifungal properties. These most likely result directly from the discrete bioactivities of each the xanthone and anthraquinone components. This compound, alongside the xanthoquinodins and beticolins, represents a biosynthetic junction of a xanthone and one of its likely biosynthetic precursors (anthraquinone). The synthesis of an ether linkage between two xanthenes has been recently reported by Sahin, Nieger and Bräse.1364.3 Dicerandrols A–C
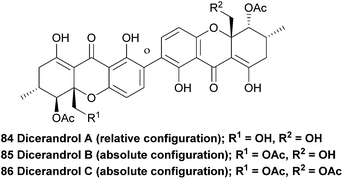
Dicerandrols A–C (84–86) were isolated from the culture broth of the Phomopsis longicolla, an endogenous fungi from the mint species Dicerandra frutescens. Extensive NMR experiments were complemented by a positive FeCl3 experiment, indicating the 2,2′-linkage mode of the dimer (the test is positive for a para-unsubstituted phenol). The 1H NMR spectrum of dicerandrol C was found to be simple, suggesting a plane of symmetry. 2D NMR experiments were used to determine the relative stereochemistry for this compound, and by inference, those of dicerandrol B and C.137 In 2013 the absolute configurations of dicerandrols B and C were determined by TDDFT electronic circular dichroism (ECD) calculations.138 It is likely that dicerandrol A has the same absolute stereochemical configuration, but to date only the relative configuration has been reported.The dicerandrols have antimicrobial activities (Bacillus subtilis and Staphylococcus aureus) with increased activities related to the decreased degree of acetylation of the molecules (i.e., A > B > C). The dicerandrols were also active against two cancer cell lines, HCT-116 and A549 (colon and lung tumour, respectively).137 In 2008, dicerandrol A was also isolated from Phomopsis sp. PSU-D15 and used in comparison studies to help identify a new member of the phomoxanthones (deacetylphomoxanthone B 101).90 Dicerandrol C was isolated from a P. longicolla obtained from red seaweed Bostrychia radicans and identified with the use of 1D and 2D NMR spectroscopic methods as well as by mass spectrometry.89 Another study performed several biological tests on dicerandrols A, B, and C that were also isolated from P. longicolla. Using disk diffusion assays they found that dicerandrols A and B possess antibacterial activity against Xanthomonas oryzae KACC 10331, a cause for bacterial blight in rice. Subsequently, dicerandrol A was also tested for antimicrobial activity against seven other X. oryzae strains, several Gram-positive and Gram-negative bacteria, a fungus and a yeast. It was found that dicerandrol A showed a relatively high activity in this broad spectrum of species, although it was often bested by commercial antibiotics. By performing growth and time-dependent production of secondary metabolites studies, it was found that the dicerandrols were produced mainly after 4 days of fermentation and that the maximal production was at day 10. This led to a maximal antibacterial activity after day 14.139 These results are useful in the production of a natural preventive medicine against bacterial blight of rice, which is very relevant since it was found that X. oryzae has developed a resistance to the available antibiotics. A more detailed continuation study on the time-dependent production of dicerandrols and other antibacterial compounds in P. longicolla confirmed that the optimum fermentation time for maximal antibacterial activity of the fungus is indeed 14 days.140 Dicerandrols A–C were also isolated in 2013 from the mycelium of a culture broth of Phomopsis longicolla S1B4 and were tested against a strain of X. oryzae. In this antimicrobial activity test the dicerandrols A–C showed a MIC of 8, 16 and 16 μg mL−1, respectively.140
The dicerandrols and the structurally similar penexanthone A were submitted to several biological activity tests versus a broad range of tumour cell lines.141 The majority of the in vitro screening tests were performed in the presence of non-malignant accessory cells, such as bone marrow stromal cells, since it was found that the activities of potential drugs can be affected by microenvironment-dependent drug resistance or sensitisation of the tumour cells.142 Among the tested compounds, dicerandrol B was found to be the most promising candidate for further investigations. It showed moderate activity against Dox40, Farage, H929, HT, OPM2 and RPMI8226 cell lines in the presence of stromal cells with IC50 values of 2.3, 1.3, 3.4, 1.3, 1.5, and 1.2 μM, respectively. More importantly, it was found that dicerandrol B showed a relatively low toxicity against human immortalised non-malignant cells, such as HS-5 bone marrow stromal cells, HOBIT osteoblast-like cells, THLE-3 hepatocytes, and SVGp12 astrocytes, compared to the values found for cancer cell lines RPMI8226 and H929. This relative selectivity is cause for further investigations into the biological activity of this compound.141 Dicerandrols A–C were also isolated from Phomopsis sp. HNY29-2B and tested for their cytotoxicity against human breast cancer, colon cancer, lung cancer, liver cancer and breast epithelial cell lines. Dicerandrol A showed a broad anti-tumour activity, but was also cytotoxic to the breast epithelial cells. Dicerandrol B and penexanthone A (101) showed also some cytotoxicity to the cancer cells, but were found to be more selective and less strongly damage the breast epithelial cells. In this assay dicerandrol C was found to be not very cytotoxic, suggesting that the free hydroxy groups are important as a key pharmacophore.143 In a study that mainly focusses on the biological activity of phomoxanthone A (106), dicerandrols B and C are also investigated. It was found that the dicerandrols were both cytotoxic to murine lymphoma cancer cell lines (IC50 values of 10 and 1.1 μM, respectively) and that dicerandrol C is slightly pro-apoptotic.138
4.4 Ergochromes
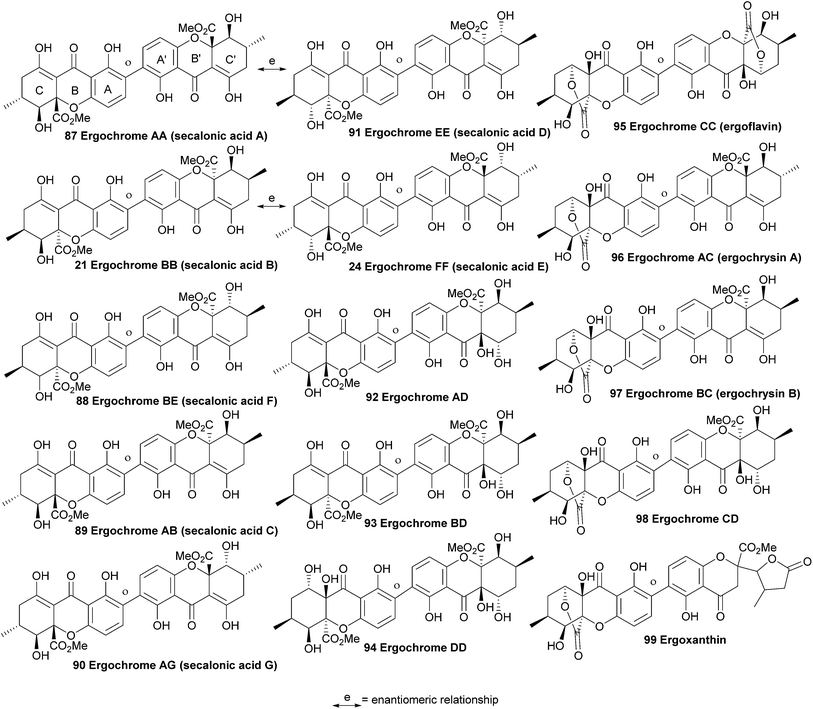
Ergochrome dimers comprise a colourful group of xanthones first reported by Kraft in 1906 as a single compound of the formula C14H14O6 after separation from Claviceps purpurea.144 The contamination of rye with ergochrome-containing C. purpurea led to epidemics known as ergotism or ‘St. Anthony's Fire’ in middle-ages Europe. The mixture of toxins produced by this fungus was responsible for the toxicity and debilitating effects on the affected population. The ergochrome dimers (21, 23, 87 to 99, Fig. 3) include the wide-spread secalonic acids, ergochrysins, ergoflavins and chrysergonic acid, many of which possess a huge variety of potent bioactivities. The ergochrome dimers are composed of several monomeric units (hemisecalonic acids A–F, aka ‘blennolides’),47 which are arranged in a variety of dimers and heterodimers. In all known secalonic acids, the methyl and methoxycarbonyl substituents are found to be trans-configured. Whilst secalonic acids are ester homodimers, other heterodimer ergochromes are known. For example, ergochrysins are lactone/ester heterodimers, ergoflavin is a homodimer,145 whilst ergoxanthin has undergone structural rearrangement to form a pendant butyrolactone ring on one half of the molecule.146Due to the early time at which they were first studied, modern spectral methods were not applicable to facilitate the structural elucidation of the ergochromes, and early structural assignations on these compounds were frequently revised in later years. For example, the initial misidentification of ergochrysin and ergoflavin as secalonic acids was later revised by Bergmann in 1932 as dimers with the formula C28H28O12.147 A number of papers followed in the 1950s to 70s, over which time more secalonic acids were isolated and identified. One major point of contention in the early days of exploration of this compound class was as to whether the natural products were 2,2′-, 4,4′- or even 2,4′-linked.148,149 The biaryl linkage was finally definitively assigned as 2,2′-linked when Mayo and Aberhart reported on the isolation, in crystalline form, of six compounds from a Portuguese ergot drug in 1965.150 In contrast to earlier reports,151 these authors showed that ergochrome dimers ubiquitously gave positive results in the Gibbs test, indicating the presence of an unsubstituted aryl methine in a position para- to the phenolic moiety, and thus a 2,2′-linkage.
A comprehensive analysis of the ergochrome constituents of C. purpurea by Franck and co-workers led to the reporting of a total of ten component compounds, including a new homodimer named ergochrome DD (94, Fig. 3), as well as the remaining six heterodimers possible with a combination of one subunit of each of those four species A, B, C and D. The majority of which they identified as the ergochromes are diastereomeric at positions 6,6′, 5,5′, 10 and 10′. The authors also identified the absolute stereochemistry at these centres.152
Büchi and co-workers reported novel secalonic acid F (88, ergochrome BE) and secalonic acid D (91, ergochrome EE) in 1977, following isolation from the fungus Aspergillus aculeatus.153 The following year, then-novel secalonic acid G (90) was isolated alongside secalonic acids A and E from Pyrenochaeta terrestris and the structure elucidated by CD and NMR spectroscopy.23
Secalonic acid D (SAD) is a major environmental toxin, being isolated from Penicillium oxalicum, a major microbial contaminant of freshly-harvested corn (in one study present on 44% of pre-harvest corn crops).154 Teratogenic effects were observed in the development of rats that were exposed to SAD injected during fetal development. SAD was lethal to mice when injected intraperitoneally in the 25–50 mg kg−1 range.155 The teratogenic and toxic nature of these compounds is alarming in light of their propensity to contaminate foodstuffs.156 The rate of birth-defects and teratogenicity-initiated spontaneous abortions in humans is also alarmingly frequent.
Despite the widespread occurrence of the ergochromes, none had been synthesised until the end of 2013, and neither had any other xanthone dimers or trimers. In early 2014, Porco and co-workers at long last described the synthesis of secalonic acids A and D, utilising a Cu(I) catalysed di-destannylative coupling.157,158 This reaction represents a significant breakthrough for the study of the xanthone dimers,62 and may open up the field to the synthesis of interesting analogues for medicinal chemistry studies. Recent advances in Ullmann coupling may make the strategy of Whalley and co-workers towards ergoflavin a more feasible route to the natural products than was originally the case in 1971.159 Bräse and co-workers have described a novel one-pot methodology for the synthesis of symmetrical biaryls160 as a part of their ongoing investigations in the synthesis of the secalonic acids. Sahin, Nieger and Bräse have also published on the oxidative coupling of various hexahydroxanthenols. The application of an iron complex as an oxidant converted 2-hydroxy substituted xanthenes to the 3,3′-bis-coupled bixanthenes.136 It appears that these two papers represent, until the end of 2013, the only successful chemistry published on the topic of xanthone or xanthene biaryl-coupling.
4.5 Neosartorin
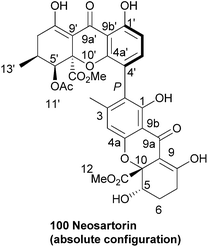
Neosartorin (100) was isolated in 1998 from the cultured mycelium of the soil mould Neosartorya fischeri. A combination of 2D NMR experiments determined that neosartorin was an isomer of eumitrin A1. However, close examination of the long range coupling constants of nuclei around the two arene units revealed a 2,4′-biaryl linkage instead of the 4,2-linkage found in eumitrin A1.161 A closer investigation of the low-intensity long-range NOEs by a 1D double-pulsed field gradient spin-echo NOESY NMR experiment resulted in confirmation of the relative configuration of the two xanthone subunits. The absolute stereochemistry of neosartorin was axially P and found to be (aR,5S,10R,5′S,6′S,10′R), this was determined on the basis of its electronic circular dichroism (ECD) spectra in conjunction162 with TDDFT-ECD calculations. They found that the nonplanar configuration of the aromatic rings connecting the two subunits was stabilised by long-range nonbonding interactions between the substituents of different xanthone units.163A taxonomical study of Aspergillus species conducted in 2005 reports the production of neosartorin in the new species A. lentulus, A. fumigatiaffinis and A. novofumigatus, but not in A. fumigatus. The presence of neosartorin was based on evaluation of UV spectral evidence. Based on these findings and other similarities the authors conclude that the taxa of Neosartorya fischeri and the Aspergillus species are chemically related.164 Neosartorin was found to show a strong activity against Gram-positive bacteria such as Staphylococcus aureus (MIC: 8 μg mL−1) and Bacillus subtilis (MIC: 4 μg mL−1), while it did not affect the tested Gram-negative bacteria. In cytotoxicity tests against several cancer cell lines it scored very badly, showing almost no notable cytotoxicity.162
4.6 Penexanthone A
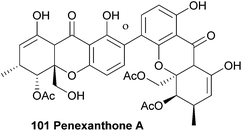
Recently, Clardy et al.141 isolated dicerandrols A–C and a novel dimeric xanthone named penexanthone A (101) from CR1642D, a Penicillium sp. obtained from the Costa Rican rainforest. The structure, including relative configuration, of penexanthone A was elucidated using mainly 2D NMR techniques (HSQC, HMBC, NOESY and ROESY) and molecular modelling with Chem3D Ultra (9.0). The two monomers were found to be connected through a 2,4′-linkage. The dicerandrols were identified by comparison of several physical and spectroscopic data sets (UV, IR, 1H NMR, [α]D and MS) with the literature values. The authors note that the structure of penexanthone A was already reported in a Korean patent, although the 1H NMR spectra was unclean, suggesting an impure sample.165 Penexanthone A is identical to monodeacetylated phomoxanthone B (vide infra). Penexanthone A was submitted to several biological activity tests versus a broad range of tumour cell lines, however the observed IC50 values indicate that penexanthone A is only weakly active in the presence of stromal cells.141 Penexanthone A was also isolated from Phomopsis sp. HNY29-2B and tested for its cytotoxicity against human breast cancer, colon cancer, lung cancer, liver cancer and breast epithelial cell lines. The dimeric xanthone showed relatively high cytotoxicity to the cancer cells and a reduced cytotoxicity towards the immortalised breast epithelial cells, which is a very useful feature when developing new anti-cancer drugs.1434.7 Penicillixanthone
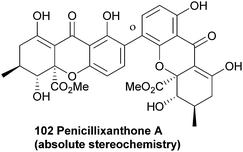
The 2,4′-biaryl-linked dimer penicillixanthone was reported around ten years ago88,166 by Chinese groups, who described the absolute stereochemistry. This compound has been re-isolated in 2013.1674.8 Phomalevones
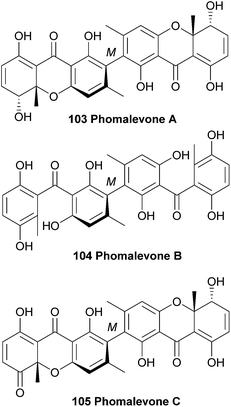
In 2011, phomalevones A–C (103–105) were isolated from a Hawaiian isolate of Phoma sp. (MYC-1734 = NRRL 39060; Cucurbitariaceae). The structures of phomalevones A–C were determined by detailed analysis of NMR and MS data.168 The absolute configuration of the sp3 stereogenic centres of phomalevone A were revealed by analysis of NMR spectra of Mosher type derivatives.Phomalevone A was found to resemble ascherxanthone B. The 13C NMR spectrum of phomalevone B showed only 15 carbon signals, thus indicating a symmetrical, homodimeric structure. Via HR-ESI-MS data it was determined that phomalevone B is an isomer of phomalevone A. However, the NMR spectra showed a significant downfield shift of the only carbonyl signal relative to its location in the spectrum of phomalevone A and the presence of two sp2 carbon signals instead of sp3 signals as in the spectrum of phomalevone A. By comparing the NMR spectroscopic data with that of acremonidin D,169 a symmetrical dimer with two benzophenone monomer units connected via a methylene linkage, a structural assignment could be made.168 The occurrence of phomalevone A together with the benzophenone analogue, phomalevone B, is consistent with several reports that hydroxylated benzophenones are very likely to be key intermediates in xanthone biosynthesis.25,170,171 HMBC spectra of phomalevone C revealed its heterodimeric nature, with one subunit the same as the monomer unit in phomalevone A, while the other has one secondary alcohol group oxidised to a carbonyl group. Phomalevones A–C showed antibacterial activity in agar disk diffusion assays at 100 μg per disk against Bacillus subtilis and Staphylococcus aureus. Additionally, phomalevone B was also active against Candida albicans and Escherichia coli and phomalevone C showed activity against Fusarium verticillioides and Aspergillus flavus (IC50 value of 4 μg mL−1, compared to an IC50 value of 5 μg mL−1 for the antifungal standard Nystatin).168
4.9 Phomoxanthones
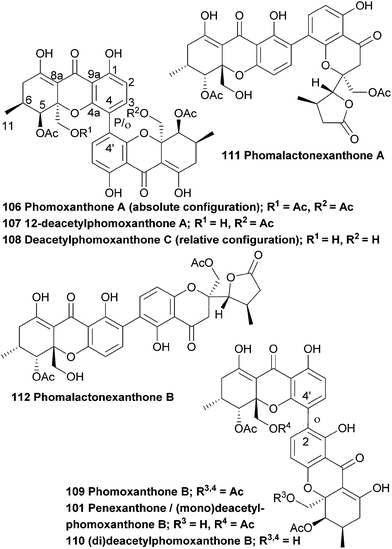
The antimalarial activity of methanolic extracts of mycelia of the endophytic fungus Phomopsis sp. BCC 1323 led to researchers isolating and identifying phomoxanthones A and B (106, 109). The relative stereochemistry could be determined by synthesising a deacetyl derivative of phomoxanthone A and comparing the change in coupling signals. Phomoxanthone A and B were the first reported examples of naturally occurring xanthone dimers with 4,4′ or 2,4′ linkages.172 In 2008, the deacetyl derivative of phomoxanthone B (110) was isolated from the same fungus species by Kirtikara et al.90 Phomolactonexanthone A and B are both derived either from coupling of a lactone-analogue of a xanthone (as the blennolides)47 and a xanthone itself, or from further derivatisation from a dimeric xanthone.Biological tests with deacetylphomoxanthone B showed a higher antibacterial effect against the pathogen Xanthomonas oryzae KACC 10331, a cause for bacterial blight in rice, than a positive control with 2,4-diacetyphloroglucinol.139 Deacetylphomoxanthone B and C (108) and phomolactonexanthone A and B (111, 112), that were isolated from Phomopsis sp. HNY29-2B, were tested for their cytotoxicity against a series of human cancer lines and it was found that while deacetylphomoxanthone B has a reasonable cytotoxicity against all tested cancer cell lines, deactylphomoxanthone C and the phomolactonexanthones were virtually inactive against all cell lines.143 Phomoxanthone A and B showed antimalarial (Plasmodium falciparum K1, multi drug resistant strain, IC50 of 0.11 and 0.33 μg mL−1, compared to IC50 of 0.16 and 0.0011 μg mL−1 for chloroquine diphosphate and artemisinin, respectively) and antitubercular activity (Mycobacterium tuberculosis, H37Ra strain, IC50 of 0.5 and 6.25 μg mL−1, compared to IC50 of 0.05 and 2.5 μg mL−1 for isoniazid and kanamycin sulfate, respectively). Biological tests on cancer cell lines (KB cells, BC-1 cells and Vero cells) showed that the natural products are cytotoxic (IC50 μg mL−1; 0.51–1.4 for phomoxanthone A and 0.7–4.1 for phomoxanthone B), although it was noted that standard drugs are more potent in both the bacterial and cancer cells tests. The deacetyl derivative was also investigated in these assays and was found to be mostly inactive in all the tests – it is speculated that this might be due to its lower lipophilicity.172
The absolute configuration (aS,5R,6R,10 aR,5′R,6′R,10a′R) of phomoxanthone A was determined by a combination of single crystal X-ray analysis with measured and calculated CD spectra, in 2005.86 However, in 2013 the absolute configuration was revised to be (aR,5S,6-S,10 aS,5′S,6′S,10a′S), when a team of researchers performed a detailed X-ray analysis study and TDDFT-ECD calculations after they suspected that the configuration might have been wrongfully assigned.138 The suspicion arose when they isolated phomoxanthone A together with dicerandrols B and C from Phomopsis longicolla and found that all their stereocenters had the S absolute configuration. In this same study they also found a new natural product from the same endophyte, which was identified by NMR studies as 12-deactylphomoxanthone A (107).
Biological tests showed that phomoxanthone A has promising activity against Gram-positive bacteria and fungi.86 In 2013 Lee et al. isolated several secondary metabolites from the mycelium of a culture broth of Phomopsis longicolla S1B4, including dicerandrols A–C (84–86) and a compound they named deacetylphomoxanthone B, 101 (which is in fact a monodeacetylphomoxanthone and is identical to penexanthone A). This compound exhibits a strong antimicrobial activity against gammaproteobacteria Xanthomonas oryzae with a MIC of 32 μg mL−1, compared to dicerandrols A–C which showed 8, 16 and 16 μg mL−1 respectively.140 Phomoxanthone A has also been found to strongly inhibit the proliferation of the murine lymphoma cell line L5178Y, as well as other cell lines, including some cisplatin resistant cancer cell lines.138 Remarkably, the cytotoxicity of phomoxanthone A towards the lymphoma cells was found to be increased over 100-fold with respect to healthy blood cells. A semisynthetic fully deacetylated phomoxanthone A showed no activity in this assay. Proksch et al. also analysed the apoptosis inducing potential of phomoxanthone A by flow cytometric analyses of hypodiploid nuclei in Jurkat T cells.138 The presence of these hypodiploid nuclei is indicative of aspartate-directed cysteine protease (caspase) induced DNA fragmentation, which is an indicator for apoptosis. They found that among all tested substances the 4,4′-linked phomoxanthone A and deacetylphomoxanthone A scored highest on these tests and actively induce apoptosis. On top of that, they also found that they activate immune effector functions in murine immune cells, such as primary T lymphocytes, NK cells and macrophages, as was observed by the upregulation of cell-type specific activation markers. During a preliminary structure–activity assessment the authors conclude that the presence of the acetyl groups and the 4,4′-linked system in xanthone dimers 106 and 107 are important to maintain the interesting biological properties, possibly because of an underlying mechanism where there is a necessity to pass the cellular membrane. The deacetylated xanthones are less lipophilic and therefore often score very low on biological tests.
When comparing the overall activity of the phomoxanthones with other similar xanthones, it must indeed be noted that the acetylated species, such as the dicerandrols and neosartorin, often show quite interesting biological properties, whereas more often the non-acetylated compounds either were not tested, or did not show promising activities. One could speculate that this is possibly due to enhanced cell penetration, however, in the case of the dicerandrols, the more acetylated dicerandrol C was found to be much less cytotoxic to cancer cells than the dicerandrols A and B that have free hydroxyl groups.
4.10 Rugulotrosins
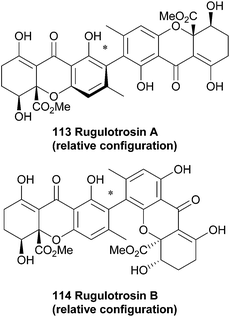
Rugulotrosins A and B (113, 114) were isolated from cultures of Penicillium sp. derived from Australian soil samples. Rugulotrosin A is symmetrically 2,2′-linked while rugulotrosin B is a 2,4′ linked dimer. The structures of these compounds were determined by NMR and MS techniques, and supported by single crystal X-ray analysis of rugulotrosin A. The compounds were of considerable activity against several microbial species (Bacillus subtilis, Enterococcus faecalis and Bacillus cereus).1734.11 Talaroxanthone
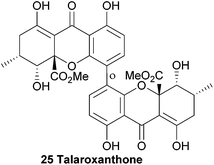
The identity of talaroxanthone (25) was reported by Koolen and co-workers in 2013. This compound, featuring a 4,4′-linked biaryl motif, was isolated from the culture broth of DgCr22.1b, an endophytic Talaromyces sp. fungi isolated from the healthy tissue of the medicinal plant Duguetia stelechantha in the Amazonian rainforest. The yellow powder that was subsequently isolated by chromatographic techniques was determined to be the structure shown by HRMS and several 2D NMR techniques. Whether the medicinal qualities of the plant material are due to the presence of this fungal-derived compound is as yet unclear.574.12 Xanthonol
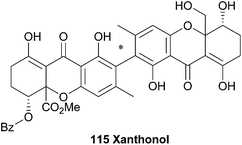
The unsymmetrical 2,2′-biaryl-linked dimeric xanthone, xanthonol (115), was isolated from the fermentation broth of a non-sporulating fungi found in the leaf litter of the plant Manikara bidentata. Xanthonol bears the methyl-substituents in the aryl positions, rather than the 6,6′-methylation of the ergochromes etc. Also interesting is the benzoylated alcohol at the C5′ position. Xanthonol is an anthelmintic, as revealed by tests upon the larvae of Lucilia sericata, Aedes aegypti, and Haemonchus contortus.1745 Xanthone dimers from lichens
5.1 Eumitrins
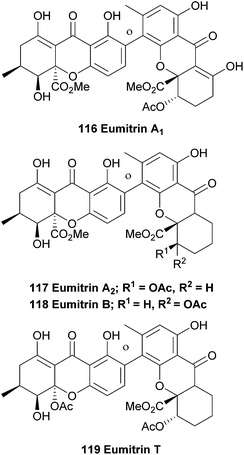
In 1967 Asahina first reported the occurrence of yellow pigments named eumitrins A1, A2, B and T (116–119) isolated from the lichen Usnea baileyi.175 Eumitrins A and B were later isolated from benzene extracts of the dried lichen thalli.176 Shibata et al. found that the fraction of eumitrin A could be separated into A1 and A2 by recrystallisation and elucidated the chemical structures of eumitrins A1, A2 and B by spectroscopic methods.177 By measuring differences in chemical shifts in NMR experiments after acetylation of the compounds, they were able to assign the biphenyl junction in these dimers as a 4,2′-linkage. Using single-crystal X-ray analysis of a tribromo derivative of eumitrin B they confirmed the structures of eumitrins A1, A2 and B. Interestingly, these xanthones have been reduced down from their normal oxidation states to bear a non-oxygenated/non-aromatic cyclohexenyl C-ring.In 2002 a Japanese patent was filed, claiming that eumitrins A1 and A2 could be used as inhibitors for nitric oxide formation by macrophages and thus be useful for a wide range of illnesses.178 Elix et al. report the extraction of secalonic acid A and eumitrin T from the lichen-forming fungus Physconia distort. Interestingly, they found that these dimeric xanthones were only found when intact lichen thalli were investigated and not in axenic cultures of the fungus, indicating a clear difference in metabolic pathways between the lichen and the axenic fungus.92 Recently, the same group investigated the contents of 18 lichen obtained from the mountainous areas of Malaysia using TLC and HPLC analyses. Amongst the secondary metabolites found in the lichen Usnea baileyi they found eumitrins A1, A3, B1 and B2. However, the structures of eumitrins A3 and B2 are not yet elucidated.179
5.2 Hirtusneanoside
Hirtusneanoside A (120) was first isolated from the lichen Usnea hirta in 2007 by Řezanaka and Sigler, and found to be a L-rhamnose-O-deoxyglycoside of an unsymmetrical dimeric tetrahydroxanthone.180 Fascinatingly, hirtusneanoside features an additional methyl in relation to the secalonic acids; the biosynthetic origin of this methyl group is as yet unknown. Hirtusneanoside A is effective against Staphylococcus aureus and Bacillus subtilis.5.3 Unnamed dimeric xanthones
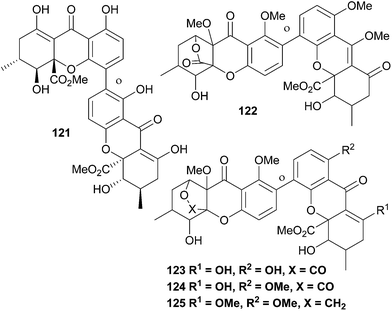
A number of unnamed xanthones have been reported, some due to the lack of full proof of their structure, some as parts of patent applications (Fig. 4). In compound 125, the lactone bridge appears to have been reductively transformed to a furan.6 Conclusions
Dimeric xanthones are a structurally highly diverse group of compounds from various kingdoms of organisms, including plants, fungi and lichens. We have reviewed their origins, structural features, and, where applicable, bioactivities. Within the last five years, a number of monomeric xanthones have been prepared16 and the first synthesis of ergochromes has been achieved. We assume that in the next decade or so – with the further advent of arylation methods suitable for the unique requirements of these species – the synthesis of dimeric xanthones will be tackled with success.7 Acknowledgements
K.-S. M. acknowledges the generous support of a Vice-Chancellor’s Research Fellowship from QUT. The research of the Bräse group has been supported by the DFG (BR 1730), the Helmholtz association and the European Union (Marie-Curie ITN ECHONET) (grant no. 316379). Co-operation between the Bräse and Masters groups is supported by an ATN-DAAD International Collaboration Award (2014–2015).8 Notes and references
- Y. Na, J. Pharm. Pharmacol., 2009, 61, 707–712 CrossRef CAS.
- M. M. M. Pinto, M. E. Sousa and M. S. Nascimento, Curr. Med. Chem., 2005, 12, 2517–2538 CrossRef CAS.
- N. Pouli and P. Marakos, Anti-Cancer Agents Med. Chem., 2009, 9, 77–98 CrossRef CAS.
- S. Malhotra, G. Shakya, A. Kumar, B. W. Vanhoecke, A. L. Cholli, H. G. Raj, L. Saso, B. Ghosh, M. E. Bracke, A. K. Prasad, S. Biswal and V. S. Parmar, ARKIVOC, 2008, 119–139 CrossRef CAS.
- T. Shan, Q. Ma, K. Guo, J. Liu, W. Li, F. Wang and E. Wu, Curr. Med. Chem., 2011, 11, 666–677 CAS.
- F. Kraft, Arch. Pharm., 1906, 244, 79 CrossRef PubMed.
- I. Carpenter, H. D. Locksley and F. Scheinmann, Phytochemistry, 1969, 8, 2013–2025 CrossRef CAS.
- F. Imperato, Curr. Top. Phytochem., 2004, 6, 63–93 CAS.
- K. Hostettmann and H. Wagner, Phytochemistry, 1977, 16, 821–829 CrossRef CAS.
- M. Afzal and J. M. Al-Hassan, Heterocycles, 1980, 14, 1173–1205 CrossRef CAS.
- G. J. Bennett and H. H. Lee, Phytochemistry, 1989, 28, 967–998 CrossRef CAS.
- K. Hostettmann and M. Hostettmann, Methods Plant Biochem., 1989, 1, 493–508 CAS.
- H. M. G. Alhazimi and G. A. Miana, J. Chem. Soc. Pak., 1990, 12, 174–188 CAS.
- S. Mandal, P. C. Das and P. C. Joshi, J. Indian Chem. Soc., 1992, 69, 611–636 CAS.
- S. Bräse, A. Encinas, J. Keck and C. F. Nising, Chem. Rev., 2009, 109, 3903–3990 CrossRef PubMed.
- K.-S. Masters and S. Bräse, Chem. Rev., 2012, 112, 3717–3776 CrossRef CAS PubMed.
- M. Riscoe, J. X. Kelly and R. Winter, Curr. Med. Chem., 2005, 12, 2539–2549 CrossRef CAS.
- M. M. M. Pinto and R. A. P. Castanheiro, Nat. Prod., 2009, 520–675 CAS.
- M. E. Sousa and M. M. M. Pinto, Curr. Med. Chem., 2005, 12, 2447–2479 CrossRef CAS.
- C. M. G. Azevedo, C. M. M. Afonso and M. M. M. Pinto, Curr. Org. Chem., 2012, 16, 2818–2867 CrossRef CAS.
- C.-H. Yang, L. Ma, Z.-P. Wei, F. Han and J. Gao, Chin. Herb. Med., 2012, 4, 87–102 CAS.
- H. R. El-Seedi, M. A. El-Barbary, D. M. H. El-Ghorab, L. Bohlin, A. K. Borg-Karlson, U. Goransson and R. Verpoorte, Curr. Med. Chem., 2010, 17, 854–901 CrossRef CAS.
- I. Kurobane, L. C. Vining, A. G. McInnes, J. A. Walter and J. L. C. Wright, Tetrahedron Lett., 1978, 19, 1379–1382 CrossRef.
- A. Ichihara and H. Oikawa, Biosci., Biotechnol., Biochem., 1997, 61, 12–18 CrossRef CAS.
- J. G. Hill, T. T. Nakashima and J. C. Vederas, J. Am. Chem. Soc., 1982, 104, 1745–1748 CrossRef CAS.
- C.-Z. Wang, U. H. Maier, M. Keil, M. H. Zenk, A. Bacher, F. Rohdich and W. Eisenreich, Eur. J. Biochem., 2003, 270, 2950–2958 CrossRef CAS.
- B. Franck, Angew. Chem., Int. Ed., 1969, 8, 251–260 CrossRef CAS PubMed.
- H. R. El-Seedi, D. M. H. El-Ghorab, M. A. El-Barbary, M. F. Zayed, U. Goransson, S. Larsson and R. Verpoorte, Curr. Med. Chem., 2009, 16, 2581–2626 CrossRef CAS.
- V. Peres, T. J. Nagem and F. F. de Oliveira, Phytochemistry, 2000, 55, 683–710 CrossRef CAS.
- L. M. M. Vieira and A. Kijjoa, Curr. Med. Chem., 2005, 12, 2413–2446 CrossRef CAS.
- L. Gales and A. M. Damas, Curr. Med. Chem., 2005, 12, 2499–2515 CrossRef CAS.
- T. Bo and H. W. Liu, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2004, 812, 165–174 CrossRef CAS PubMed.
- D. K. Winter, D. L. Sloman and J. A. Porco, Nat. Prod. Rep., 2013, 30, 382–391 RSC.
- O. Chantarasriwong, A. Batova, W. Chavasiri and E. A. Theodorakis, Chem.–Eur. J., 2010, 16, 9944–9962 CrossRef CAS PubMed.
- N. Anantachoke, P. Tuchinda, C. Kuhakarn, M. Pohmakotr and V. Reutrakul, Pharm. Biol., 2012, 50, 78–91 CrossRef CAS PubMed.
- A. J. Birch, J. Baldas, J. R. Hlubucek, T. J. Simpson and P. W. Westerman, J. Chem. Soc., Perkin Trans. 1, 1976, 898–904 RSC.
- H. G. Floss, Lloydia, 1968, 31, 427 Search PubMed.
- G. J. Bennett and H. H. Lee, J. Chem. Soc., Chem. Commun., 1988, 619–620 RSC.
- T. J. Simpson, ChemBioChem, 2012, 13, 1680–1688 CrossRef CAS PubMed.
- K. M. Henry and C. A. Townsend, J. Am. Chem. Soc., 2005, 127, 3724–3733 CrossRef CAS PubMed.
- J. F. Sanchez, R. Entwistle, J.-H. Hung, J. Yaegashi, S. Jain, Y.-M. Chiang, C. C. C. Wang and B. R. Oakley, J. Am. Chem. Soc., 2011, 133, 4010–4017 CrossRef CAS PubMed.
- M. A. Schätzle, S. M. Husain, S. Ferlaino and M. Müller, J. Am. Chem. Soc., 2012, 134, 14742–14745 CrossRef PubMed.
- J. A. Elix, H. M. Chappell and J. Hui, Bryologist, 1991, 94, 304–307 CrossRef CAS.
- J. A. Elix and C. E. Crook, Bryologist, 1992, 95, 52–64 CrossRef CAS.
- S. Tarcz, X. Xie and S.-M. Li, RSC Adv., 2014, 4, 17986–17992 RSC.
- G. Bringmann, C. Guennther, M. Ochse, O. Schupp and S. Tasler, Prog. Chem. Org. Nat. Prod., 2001, 82, 1–249 CAS.
- W. Zhang, K. Krohn, Z. Ullah, U. Florke, G. Pescitelli, L. Di Bari, S. Antus, T. Kurtan, J. Rheinheimer, S. Draeger and B. Schulz, Chem.–Eur. J., 2008, 14, 4913–4923 CrossRef CAS PubMed.
- B. Franck, F. Hüper, D. Gröger and D. Erge, Chem. Ber., 1968, 101, 1954–1969 CrossRef CAS PubMed.
- I. Kurobane, L. C. Vining and A. G. McInnes, J. Antibiot., 1979, 32, 1256–1266 CrossRef CAS.
- B. Franck, G. Bringmann and G. Flohr, Angew. Chem., 1980, 92, 483–484 CrossRef CAS PubMed.
- M. C. Kozlowski, B. J. Morgan and E. C. Linton, Chem. Soc. Rev., 2009, 38, 3193–3207 RSC.
- S. E. Bode, D. Drochner and M. Müller, Angew. Chem., Int. Ed., 2007, 46, 5916–5920 CrossRef CAS PubMed.
- R. F. Helm, L. Zhentian, T. Ranatunga, J. Jervis and T. Elder, Basic Life Sci., 1999, 66, 83–99 CAS.
- C. G. Girol, K. M. Fisch, T. Heinekamp, S. Gunther, W. Hüttel, J. Piel, A. A. Brakhage and M. Müller, Angew. Chem., Int. Ed., 2012, 51, 9788–9791 CrossRef PubMed.
- J. B. Hewgley, S. S. Stahl and M. C. Kozlowski, J. Am. Chem. Soc., 2008, 130, 12232–12233 CrossRef CAS PubMed.
- A. Schüffler, J. C. Liermann, H. Kolshorn, T. Opatz and H. Anke, Tetrahedron Lett., 2009, 50, 4813–4815 CrossRef PubMed.
- H. H. F. Koolen, L. S. Menezes, M. P. Souza, F. M. A. Silva, F. G. O. Almeida, A. Q. L. de Souza, A. Nepel, A. Barison, F. H. da Silva, D. E. Evangelista and A. D. L. de Souza, J. Braz. Chem. Soc., 2013, 24, 880–883 CAS.
- C. C. Howard and R. A. W. Johnstone, J. Chem. Soc., Perkin Trans. 1, 1973, 2440–2444 RSC.
- C. C. Howard and R. A. W. Johnstone, J. Chem. Soc., Chem. Commun., 1973, 464 RSC.
- R. Pummerer, H. Puttfarcken and P. Schopflocher, Ber. Dtsch. Chem. Ges., 1925, 58, 1808–1820 CrossRef PubMed.
- T. Pal and A. Pal, Curr. Sci., 1996, 71, 106–108 CAS.
- K. S. Feldman, K. J. Eastman and G. Lessene, Org. Lett., 2002, 4, 3525–3528 CrossRef CAS PubMed.
- B. Elsler, D. Schollmeyer, K. M. Dyballa, R. Franke and S. R. Waldvogel, Angew. Chem., Int. Ed., 2014, 53, 5210–5213 CAS.
- T. Wezeman, K.-S. Masters and S. Bräse, Angew. Chem., Int. Ed., 2014, 53, 4524–4526 CrossRef CAS PubMed.
- K.-S. Masters and S. Bräse, Angew. Chem., Int. Ed., 2013, 52, 866–869 CrossRef CAS PubMed.
- K.-S. Masters, A. Bihlmeier, W. Klopper and S. Bräse, Chem.–Eur. J., 2013, 19, 17827–17835 CrossRef CAS PubMed.
- E. I. Solomon, A. J. Augustine and J. Yoon, Dalton Trans., 2008, 3921–3932 RSC.
- H. Claus, Arch. Microbiol., 2003, 179, 145–150 CAS.
- S. Riva, Trends Biotechnol., 2006, 24, 219–226 CrossRef CAS PubMed.
- M.-A. Constantin, J. Conrad and U. Beifuss, Green Chem., 2012, 14, 2375–2379 RSC.
- M.-A. Constantin, J. Conrad and U. Beifuss, Tetrahedron Lett., 2012, 53, 3254–3258 CrossRef CAS PubMed.
- A. Kirste, M. Nieger, I. M. Malkowsky, F. Stecker, A. Fischer and S. R. Waldvogel, Chem.–Eur. J., 2009, 15, 2273–2277 CrossRef CAS PubMed.
- I. M. Malkowsky, C. E. Rommel, R. Frohlich, U. Griesbach, H. Putter and S. R. Waldvogel, Chem.–Eur. J., 2006, 12, 7482–7488 CrossRef CAS PubMed.
- H. Leutbecher, M. A. Constantin, S. Mika, J. Conrad and U. Beifuss, Tetrahedron Lett., 2011, 52, 605–608 CrossRef CAS PubMed.
- A. Wells, M. Teria and T. Eve, Biochem. Soc. Trans., 2006, 34, 304–308 CrossRef CAS PubMed.
- I. W. C. E. Arends, Y. X. Li, R. Ausan and R. A. Sheldon, Tetrahedron, 2006, 62, 6659–6665 CrossRef CAS PubMed.
- F. d'Acunzo, P. Baiocco and C. Galli, New J. Chem., 2003, 27, 329–332 RSC.
- E. Fritz-Langhals and B. Kunath, Tetrahedron Lett., 1998, 39, 5955–5956 CrossRef CAS.
- A. Potthast, T. Rosenau, C. L. Chen and J. S. Gratzl, J. Mol. Catal. A: Chem., 1996, 108, 5–9 CrossRef CAS.
- S. Witayakran and A. J. Ragauskas, Adv. Synth. Catal., 2009, 351, 1187–1209 CrossRef CAS PubMed.
- A. Mikolasch and F. Schauer, Appl. Microbiol. Biotechnol., 2009, 82, 605–624 CrossRef CAS PubMed.
- A. Kunamneni, S. Camarero, C. Garcia-Burgos, F. J. Plou, A. Ballesteros and M. Alcalde, Microb. Cell Fact., 2008, 7, 32 CrossRef PubMed.
- G. J. Bennett, H. H. Lee and T. K. Lowrey, Tetrahedron Lett., 1990, 31, 751–754 CrossRef CAS.
- (a) J. G. Liu, R. D. Stipanovic, A. A. Bell, L. S. Puckhaber and C. W. Magill, Phytochemistry, 2008, 69, 3038–3042 CrossRef CAS PubMed; (b) J. Liu, R. D. Stipanovic, A. A. Bell, L. S. Puckhaber and C. W. Magill, J. Agric. Food Chem., 2009, 57, 566–571 CrossRef PubMed.
- B. Pickel, M.-A. Constantin, J. Pfannstiel, J. Conrad, U. Beifuss and A. Schaller, Angew. Chem., Int. Ed., 2010, 49, 202–204 CrossRef CAS PubMed.
- B. Elsässer, K. Krohn, U. Flörke, N. Root, H.-J. Aust, S. Draeger, B. Schulz, S. Antus and T. Kurtán, Eur. J. Org. Chem., 2005, 4563–4570 CrossRef PubMed.
- I. Warnke and F. Furche, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2012, 2, 150–166 CrossRef CAS PubMed.
- L. Wen, Z. Guo, F. Liu, Q. Wan, Z. Yu, Y. Lin and L. Fu, Huaxue Yanjiu Yu Yingyong, 2009, 21, 198–202 CAS.
- C. Erbert, A. A. Lopes, N. S. Yokoya, N. A. J. C. Furtado, R. Conti, M. T. Pupo, J. L. C. Lopes and H. M. Debonsi, Bot. Mar., 2012, 55, 435–440 CAS.
- V. Rukachaisirikul, U. Sommart, S. Phongpaichit, J. Sakayaroj and K. Kirtikara, Phytochemistry, 2008, 69, 783–787 CrossRef CAS PubMed.
- C. Leuckert, V. Ahmadjian, C. F. Culberson and A. Johnson, Mycologia, 1990, 82, 370–378 CrossRef CAS.
- M. C. Molina, A. Crespo, C. Vicente and J. A. Elix, Plant Physiol. Biochem., 2003, 41, 175–180 CrossRef CAS.
- Q.-L. Wu, S.-P. Wang, L.-J. Du, J.-S. Yang and P.-G. Xiao, Phytochemistry, 1998, 49, 1395–1402 CrossRef CAS.
- P. Fu, T. Li, R. Liu, W. Zhang, W. Zhang and H. Chen, Tianran Chanwu Yanjiu Yu Kaifa, 2004, 16, 511–513 CAS.
- L. Zhang, Y. Jin and J. Tian, Zhongguo Yaoxue Zazhi, 2007, 42, 341–344 CAS.
- P. Fu, W. D. Zhang, T. Z. Li, R. H. Liu, H. L. Li, W. Zhang and H. S. Chen, Chin. Chem. Lett., 2005, 16, 771–773 CAS.
- M. F. Tala, P. D. Tchakam, H. K. Wabo, F. M. Talontsi, P. Tane, J. R. Kuiate, L. A. Tapondjou and H. Laatsch, Cardiovasc. Drug Rev., 2013, 7, 65–68 CAS.
- W. Zhang, P. Fu, H. Li, C. Zhang, J. Su and R. Liu, Chinese Patent no. CN1660307A, 2005.
- F.-F. Zhong, Y. Chen and G.-Z. Yang, Helv. Chim. Acta, 2008, 91, 1695–1703 CrossRef CAS PubMed.
- Y. Chen, H. Fan, G.-Z. Yang, Y. Jiang, F.-F. Zhong and H.-W. He, Helv. Chim. Acta, 2011, 94, 662–668 CrossRef CAS PubMed.
- S. Mandal and A. Chatterjee, Tetrahedron Lett., 1987, 28, 1309–1310 CrossRef CAS.
- G.-L. Sia, G. J. Bennett, L. J. Harrison and K.-Y. Sim, Phytochemistry, 1995, 38, 1521–1528 CrossRef CAS.
- A.-R. Han, J.-A. Kim, D. D. Lantvit, L. B. S. Kardono, S. Riswan, H. Chai, d. B. E. J. Carcache, N. R. Farnsworth, S. M. Swanson and A. D. Kinghorn, J. Nat. Prod., 2009, 72, 2028–2031 CrossRef CAS PubMed.
- I. Sordat-Diserens, M. Toyota, M. Hamburger, B. Sordat and K. Hostettmann, Bull. Liaison–Groupe Polyphenols, 1990, 15, 175–178 CAS.
- I. Sordat-Diserens, M. Hamburger, C. Rogers and K. Hostettmann, Phytochemistry, 1992, 31, 3589–3593 CrossRef CAS.
- Z. H. Mbwambo, M. C. Kapingu, M. J. Moshi, F. Machumi, S. Apers, P. Cos, D. Ferreira, J. P. J. Marais, B. D. Vanden, L. Maes, A. Vlietinck and L. Pieters, J. Nat. Prod., 2006, 69, 369–372 CrossRef CAS PubMed.
- S. Feng, Y. Jiang, J. Li, S. Qiu and T. Chen, Nat. Prod. Res., 2014, 28, 81–85 CrossRef CAS PubMed.
- A. E. Nkengfack, P. Mkounga, M. Meyer, Z. T. Fomum and B. Bodo, Phytochemistry, 2002, 61, 181–187 CrossRef CAS.
- Y.-J. Xu, S.-G. Cao, X.-H. Wu, Y.-H. Lai, B. H. K. Tan, J. T. Pereira, S. H. Goh, G. Venkatraman, L. J. Harrison and K.-Y. Sim, Tetrahedron Lett., 1998, 39, 9103–9106 CrossRef CAS.
- S. Hartati, H.-B. Wang, L. B. S. Kardono, S. Kosela and G.-W. Qin, Zhongguo Tianran Yaowu, 2007, 5, 272–276 CAS.
- J. Merza, M.-C. Aumond, D. Rondeau, V. Dumontet, R. A.-M. Le, D. Seraphin and P. Richomme, Phytochemistry, 2004, 65, 2915–2920 CrossRef CAS PubMed.
- S. Feng and T. Chen, Shenzhen Fairy Lake Botanical Garden Management Office, Peop. Rep. China, CN102440985A, 2012.
- W. Wang, Y. H. Zeng, K. Osman, K. Shinde, M. Rahman, S. Gibbons and Q. Mu, J. Nat. Prod., 2010, 73, 1815–1820 CrossRef CAS PubMed.
- K. Ishiguro, S. Nagata, H. Oku and M. Yamaki, Planta Med., 2002, 68, 258–261 CrossRef CAS PubMed.
- W.-D. Zhang, P. Fu, R.-H. Liu, T.-Z. Li, H.-L. Li, W. Zhang and H.-S. Chen, Fitoterapia, 2007, 78, 74–75 CrossRef CAS PubMed.
- S. Singh, A. I. Gray and P. G. Waterman, Nat. Prod. Lett., 1993, 3, 53–58 CrossRef CAS.
- M. Iinuma, H. Tosa, T. Tanaka and S. Riswan, Heterocycles, 1996, 43, 1999–2004 CrossRef CAS.
- P. Tan, C.-Y. Hou, Y.-L. Liu, L.-J. Lin and G. A. Cordell, J. Org. Chem., 1991, 56, 7130–7133 CrossRef CAS.
- P. Tan, C.-Y. Hou, Y.-L. Liu, L.-J. Lin and G. A. Cordell, Phytochemistry, 1992, 31, 4313–4315 CrossRef CAS.
- S. P. Zhang, X. G. Du and X. P. Pu, Biol. Pharm. Bull., 2010, 33, 1529–1533 CAS.
- A. Urbain, A. Marston, L. S. Grilo, J. Bravo, O. Purev, B. Purevsuren, D. Batsuren, M. Reist, P. A. Carrupt and K. Hostettmann, J. Nat. Prod., 2008, 71, 895–897 CrossRef CAS PubMed.
- H.-M. Zhou, Y.-L. Liu, G. Blaskó and G. A. Cordell, Phytochemistry, 1989, 28, 3569–3571 CrossRef CAS.
- X.-G. Du, W. Wang, S.-P. Zhang, X.-P. Pu, Q.-Y. Zhang, M. Ye, Y.-Y. Zhao, B.-R. Wang, I. A. Khan and D.-A. Guo, J. Nat. Prod., 2010, 73, 1422–1426 CrossRef CAS PubMed.
- X.-G. Du, W. Wang, Q.-Y. Zhang, J. Cheng, B. Avula, I. A. Khan and D.-A. Guo, Rapid Commun. Mass Spectrom., 2012, 26, 2913–2923 CrossRef CAS PubMed , S2913/2911–S2913/2920.
- M. Isaka, S. Palasarn, K. Kocharin and J. Saenboonrueng, J. Nat. Prod., 2005, 68, 945–946 CrossRef CAS PubMed.
- N. Sakurai, H. Akatsuka, J. Minakami, M. Nishio and O. Kono, Tanabe Seiyaku Co., Ltd., Japan, JP2004075625A, 2004.
- C. Chutrakul, T. Boonruangprapa, R. Suvannakad, M. Isaka, P. Sirithunya, T. Toojinda and K. Kirtikara, J. Appl. Microbiol., 2009, 107, 1624–1631 CrossRef CAS PubMed.
- M.-L. Milat and J.-P. Blein, J. Chromatogr. A, 1995, 699, 277–283 CrossRef CAS.
- C. Goudet, M.-L. Milat, H. Sentenac and J.-B. Thibaud, Mol. Plant-Microbe Interact., 2000, 13, 203–209 CrossRef CAS PubMed.
- P.-H. Ducrot, J. Einhorn, L. Kerhoas, J.-Y. Lallemand, M.-L. Milat, J.-P. Blein, A. Neuman and T. Prangé, Tetrahedron Lett., 1996, 37, 3121–3124 CrossRef CAS.
- P.-H. Ducrot, J.-Y. Lallemand, M.-L. Milat and J.-P. Blein, Tetrahedron Lett., 1994, 47, 8797–8800 CrossRef.
- J. Y. Ueda, M. Takagi and K. Shin-ya, J. Antibiot., 2010, 63, 615–618 CrossRef CAS PubMed.
- K. Matsuzaki, N. Tabata, H. Tomoda, Y. Iwai, H. Tanaka and S. Omura, Tetrahedron Lett., 1993, 34, 8251–8254 CrossRef CAS.
- N. Tabata, H. Tomoda, Y. Iwai and S. Omura, J. Antibiot., 1995, 49, 267–271 CrossRef.
- N. Tabata, H. Tomoda, K. Matsuzaki and S. Omura, J. Am. Chem. Soc., 1993, 115, 8558–8564 CrossRef CAS.
- H. Sahin, M. Nieger and S. Bräse, Eur. J. Org. Chem., 2009, 2009, 5576–5586 CrossRef PubMed.
- M. M. Wagenaar and J. Clardy, J. Nat. Prod., 2001, 64, 1006–1009 CrossRef CAS PubMed.
- D. Rönsberg, A. Debbab, A. Mándi, V. Vasylyeva, P. Böhler, B. Stork, L. Engelke, A. Hamacher, R. Sawadogo, M. Diederich, V. Wray, W. Lin, M. U. Kassack, C. Janiak, S. Scheu, S. Wesselborg, T. Kurtán, A. H. Aly and P. Proksch, J. Org. Chem., 2013, 78, 12409–12425 CrossRef PubMed.
- C. Lim, J. Kim, J. N. Choi, K. Ponnusamy, Y. Jeon, S.-U. Kim, J. G. Kim and C. H. Lee, J. Microbiol. Biotechnol., 2010, 20, 494–500 CAS.
- J. N. Choi, J. Kim, K. Ponnusamy, C. Lim, J. G. Kim, M. J. Muthaiya and C. H. Lee, J. Antibiot., 2013, 66, 231–233 CrossRef CAS PubMed.
- S. Cao, D. W. McMillin, G. Tamayo, J. Delmore, C. S. Mitsiades and J. Clardy, J. Nat. Prod., 2012, 75, 793–797 CrossRef CAS PubMed.
- D. W. McMillin, J. Delmore, J. Negri, M. Ooi, S. Klippel, C. V. Miduturu, N. S. Gray, P. G. Richardson, K. C. Anderson, A. L. Kung and C. S. Mitsiades, PLoS One, 2011, 6, e20226 CAS.
- B. Ding, J. Yuan, X. Huang, W. Wen, X. Zhu, Y. Liu, H. Li, Y. Lu, L. He, H. Tan and Z. She, Mar. Drugs, 2013, 11, 4961–4972 CrossRef CAS PubMed.
- F. Kraft, Arch. Pharm., 1906, 244, 336–359 CrossRef PubMed.
- J. W. ApSimon, J. A. Corran, N. G. Creasey, W. Marlow, W. B. Whalley and K. Y. Sim, J. Chem. Soc., 1965, 4144–4156 RSC.
- W. Bergmann, Ber. Dtsch. Chem. Ges. A, 1932, 65, 1489–1494 CrossRef PubMed.
- W. Bergmann, Ber. Dtsch. Chem. Ges. A, 1932, 65, 1486–1488 CrossRef PubMed.
- B. Franck and G. Baumann, Chem. Ber., 1963, 96, 3209–3216 CrossRef CAS PubMed.
- B. Franck and E. M. Gottschalk, Angew. Chem., 1964, 76, 438 Search PubMed.
- D. J. Aberhart, Y. S. Chen, P. de Mayo and J. B. Stothers, Tetrahedron, 1965, 21, 1417–1432 CrossRef CAS.
- B. Franck and G. Baumann, Chem. Ber., 1966, 99, 3875–3883 CrossRef CAS PubMed.
- B. Franck, G. Baumann and U. Ohnsorge, Tetrahedron Lett., 1965, 6, 2031–2037 CrossRef.
- R. Andersen, G. Buchi, B. Kobbe and A. L. Demain, J. Org. Chem., 1977, 42, 352–353 CrossRef CAS.
- W. W. Carlton, J. Tuite and P. Mislivec, Toxicol. Appl. Pharmacol., 1968, 13, 372–387 CrossRef CAS.
- P. B. Mislivec and J. Tuite, Mycologia, 1970, 62, 67–74 CrossRef CAS.
- A. Ciegler, A. W. Hayes and R. F. Vesonder, Appl. Environ. Microbiol., 1980, 39, 285–287 CAS.
- T. Qin and J. A. Porco, Angew. Chem., 2014, 126, 3171–3174 CrossRef PubMed.
- T. Qin and J. A. Porco, Angew. Chem., Int. Ed., 2014, 53, 3107–3110 CrossRef CAS PubMed.
- J. W. Hooper, W. Marlow, W. B. Whalley, A. D. Borthwick and R. Bowden, J. Chem. Soc. D, 1971, 111–112 RSC.
- C. F. Nising, U. K. Schmid, M. Nieger and S. Bräse, J. Org. Chem., 2004, 69, 6830–6833 CrossRef CAS PubMed.
- B. Proksa, D. Uhrin, T. Liptaj and M. Sturdikova, Phytochemistry, 1998, 48, 1161–1164 CrossRef CAS.
- A. R. B. Ola, A. Debbab, A. H. Aly, A. Mandi, I. Zerfass, A. Hamacher, M. U. Kassack, H. Brotz-Oesterhelt, T. Kurtan and P. Proksch, Tetrahedron Lett., 2014, 55, 1020–1023 CrossRef CAS PubMed.
- T. Liptaj, T. N. Pham, B. Proksa and D. Uhrin, Chirality, 2001, 13, 545–547 CrossRef CAS PubMed.
- S.-B. Hong, S.-J. Go, H.-D. Shin, J. C. Frisvad and R. A. Samson, Mycologia, 2005, 97, 1316–1329 CrossRef CAS.
- C. H. Lee, J. G. Kim, C. S. Lim and J. Y. Kim, Korean Patent no. KR2011021258 A2020110304, 2011.
- T. Jiang, L. Tian, A. Guo, H. Fu, Y. Pei and W. Lin, Yaoxue Xuebao, 2002, 37, 271–274 CAS.
- J. Bao, Y.-L. Sun, X.-Y. Zhang, Z. Han, H.-C. Gao, F. He, P.-Y. Qian and S.-H. Qi, J. Antibiot., 2013, 66, 219–223 CrossRef CAS PubMed.
- S. H. Shim, J. Baltrusaitis, J. B. Gloer and D. T. Wicklow, J. Nat. Prod., 2011, 74, 395–401 CrossRef CAS PubMed.
- H. He, R. Bigelis, E. H. Solum, M. Greenstein and G. T. Carter, J. Antibiot., 2003, 56, 923–930 CrossRef CAS.
- W. Schmidt and L. Beerhues, FEBS Lett., 1997, 420, 143–146 CrossRef CAS.
- H. Kachi and T. Sassa, Agric. Biol. Chem., 1986, 50, 1669–1671 CrossRef CAS.
- M. Isaka, A. Jaturapat, K. Rukseree, K. Danwisetkanjana, M. Tanticharoen and Y. Thebtaranonth, J. Nat. Prod., 2001, 64, 1015–1018 CrossRef CAS PubMed.
- M. Stewart, R. J. Capon, J. M. White, E. Lacey, S. Tennant, J. H. Gill and M. P. Shaddock, J. Nat. Prod., 2004, 67, 728–730 CrossRef CAS PubMed.
- J. G. Ondeyka, A. W. Dombrowski, J. P. Polishook, T. Felcetto, W. L. Shoop, Z. Guan and S. B. Singh, J. Antibiot., 2006, 59, 288–292 CrossRef CAS PubMed.
- Y. Asahina, J. Jpn. Bot., 1967, 42, 1 Search PubMed.
- M. Nuno, J. Jpn. Bot., 1971, 46, 294 Search PubMed.
- D.-M. Yang, N. Takeda, Y. Iitaka, V. Sankawa and S. Shibata, Tetrahedron, 1973, 29, 519–528 CrossRef CAS.
- K. Oshima, Y. Fujimiya, M. Soda, F. Takano and S. Fushitani, Netairin Saisei Gijutsu Kenkyu Kumiai, Japan, JP2002047181A, 2002.
- L. B. Din, Z. Zakaria, M. W. Samsudin and J. A. Elix, Sains Malays., 2010, 39, 901–908 CAS.
- T. Rezanka and K. Sigler, J. Nat. Prod., 2007, 70, 1487–1491 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: In order to provide a clear overview of the structural features, natural sources and biological activities of the discussed xanthones, three tables detailing these topics have been added as ESI. See DOI: 10.1039/c4np00050a. |
| This journal is © The Royal Society of Chemistry 2015 |