Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Food derived microRNAs
Anika E.
Wagner
*,
Stefanie
Piegholdt
,
Martin
Ferraro
,
Kathrin
Pallauf
and
Gerald
Rimbach
Institute of Human Nutrition and Food Science, Christian-Albrechts-University, Hermann-Rodewald-Str. 6, 24118 Kiel, Germany. E-mail: wagner@molecularnutrition.uni-kiel.de; Fax: +49 431 880 2628; Tel: +49 431 880 5313
First published on 22nd January 2015
Abstract
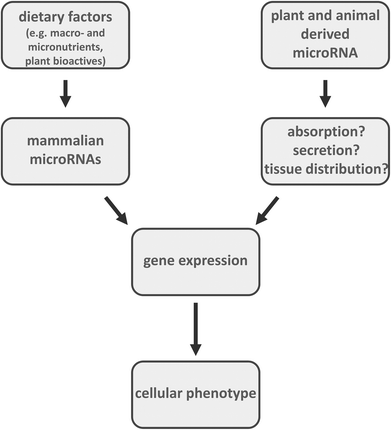
Foods provide fats, carbohydrates, and proteins as well as vitamins, minerals and trace elements. These dietary factors may influence cellular processes by regulating endogenous microRNA expression. MicroRNAs are non-coding regulatory molecules which affect gene expression at the post transcriptional level. It has been shown that plant and animal derived foods also contain microRNA. Yet, it is unclear if and to what extent plant and animal food derived microRNAs are absorbed by mammals. Thus, future studies need to better address absorption, tissue distribution and function of dietary plant and animal derived microRNAs in the context of human health and disease.
Over the last decade it has become apparent that nutrition does not only provide fats, carbohydrates, proteins, vitamins, minerals and trace elements. Beside macro- and micronutrients, plant and animal derived foods also contain considerable amounts of so-called microRNAs.
MicroRNAs are evolutionarily conserved small non-coding RNAs consisting of on average 22 nucleotides.1–5 They mediate post-transcriptional changes in gene expression usually through binding to the 3′ untranslated region (3′UTR) of target mRNAs with their seed region (nucleotides 2–7). Depending on the base pair complementarity between the microRNA and the target mRNA, the target mRNA is either degraded (perfect match) or the ribosomal translation is blocked (imperfect binding) which in turn influences the cellular phenotype (for microRNA processing see Fig. 1).
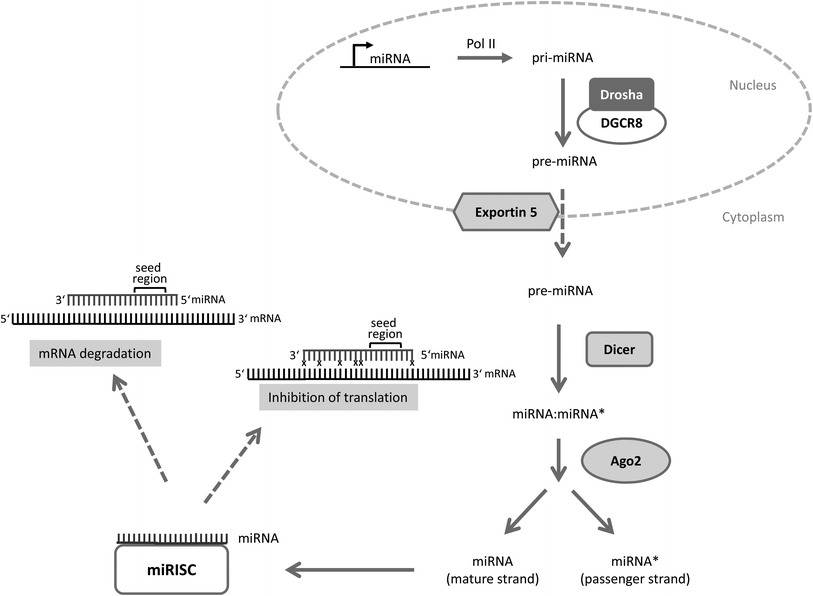 | ||
Fig. 1 microRNA processing. Primary microRNAs (pri-miRNAs) are specifically transcribed from microRNA genes by polymerase II (Pol II). Pri-microRNA is cleaved by an enzyme complex consisting of Drosha and DGCR8 (DiGeorge critical region 8) into a precursor microRNA (pre-miRNA) of ∼70 nucleotides which is exported from the nucleus into the cytoplasm by the nuclear export protein exportin 5. Once in the cytoplasm the pre-miRNA is processed by the enzyme Dicer into a miRNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) miRNA* duplex which is then unwound by Ago2 into two single strands. Ago2 is part of the miR-RISC (microRNA – RNA induced silencing complex; miR-RISC). The miRNA strand is the mature strand whereas miRNA* represents the passenger strand which is usually degraded. However, some passenger strands enter the RISC and function as a “normal” microRNA. The mature miRNA-strand connects to the RISC and finally targets the mRNAs. Depending on the complementarity between mRNA and microRNA either the translation is inhibited or the mRNA is degraded. x indicates mismatch between microRNA and mRNA (modified according to ref. 4). miRNA* duplex which is then unwound by Ago2 into two single strands. Ago2 is part of the miR-RISC (microRNA – RNA induced silencing complex; miR-RISC). The miRNA strand is the mature strand whereas miRNA* represents the passenger strand which is usually degraded. However, some passenger strands enter the RISC and function as a “normal” microRNA. The mature miRNA-strand connects to the RISC and finally targets the mRNAs. Depending on the complementarity between mRNA and microRNA either the translation is inhibited or the mRNA is degraded. x indicates mismatch between microRNA and mRNA (modified according to ref. 4). | ||
It has become increasingly evident that microRNAs are involved in the regulation of a large number of cellular processes.6 MicroRNAs are found in all human cells and currently almost 2000 sequences are listed in the microRNA database http://www.mirbase.org. Other important microRNA databases include http://www.microRNA.org, http://www.ncrna.org and http://www.mirdb.org. It has been hypothesized that as many as 60% of the human protein-coding genes may contain microRNA binding sites.7 In line with this estimation, microRNAs play a role in the genesis of obesity, diabetes, neurodegenerative diseases and cancer.8,9 Consistently, microRNAs were shown to influence insulin production in pancreatic β-cells by regulating transcriptional repressors,10 miR-33 and other microRNAs were shown to affect lipid metabolism,11 miR-15 and miR-16 promote apoptosis through inhibiting the expression of pro-apoptotic Bcl-212 and miR-376b and miR-30a regulate autophagy by targeting pro-autophagic beclin 1.13,14 Given these implications for health and disease, interest in developing microRNAs as therapeutics for illnesses is rising and a first liposome-based microRNA mimic (MRX34) has reached phase I clinical trials.15
Diet may also influence cellular processes by regulating microRNA expression16 (Fig. 2). Food components such as vitamins (e.g. vitamin D, vitamin E, folate) and secondary plant metabolites (e.g. epigallocatechingallate, curcumin, resveratrol, quercetin, isothiocyanates) were reported as modulators of microRNA levels.6,17–19 Moreover, there has been a report that microRNAs from food may regulate gene expression in mammals. It was shown that miR-168a from rice is transported from the gastrointestinal tract to target organs in mice. In murine liver, this plant-derived microRNA bound to low-density lipoprotein receptor adapter protein 1 (LDLRAP1) mRNA and inhibited LDLRAP1 expression, thereby interfering with cholesterol transport.20 Another recent study published by the same group suggests that in influenza-infected mice honeysuckle exhibits its anti-viral potency via microRNA-2911.21 Yet, a so-called cross-kingdom regulation by microRNAs is still under debate22–24 since a different group of researchers could not detect plant microRNAs after feeding in mice25 and studies in humans did not lead to detectable amounts of microRNAs in the plasma after consumption of fruits26 and broccoli sprouts.27 In contrast, Lukasik and Zielenkiewicz28 identified plant-derived microRNAs in exosomes of human and porcine breast milk and, by using a bioinformatics approach, predicted their potential human target mRNAs. Interestingly, Melnik and co-workers29 relate the atopy-preventing effect of raw cow milk to exosomal miR-155 which is present in high amounts in raw cow milk (especially colostrum) and also in human breast milk. miR-155 is known to play a pivotal role in the development of the immune system and especially in the activity of regulatory T cells (Treg).29–31 While naked microRNAs are highly unstable and are quickly degraded by omnipresent RNases several papers note that circulating microRNAs in blood and various body fluids are highly stable which predisposes them as a potential biomarker for malignant and non-malignant diseases. This exceptionally high stability is owed to the fact that microRNAs can either form complexes with proteins or be packed into small vesicles originating from endosomal membrane compartments or the plasma membrane itself.32 A high number of microRNAs in human breast milk are present in microvesicles or exosomes which protect them from rapid degradation by RNases.33,34 It has been proposed that microRNAs are involved in cell–cell communication and play an important role in the immune system maturation of the infant.33,34 There is evidence that exosomes are transported by human cells through phagocytosis and carrier-mediated processes. It was shown that cow-milk-derived exosomes are transported by human intestinal cells following a saturated kinetic process which can be inhibited at low temperatures as well as through an elimination of exosome surface proteins.35
Recently Zempleni and co-workers have demonstrated that humans seem to absorb cow milk derived microRNAs in significant concentrations and that these microRNAs seem to affect gene expression in human white blood cells, cultured kidney cells, and mouse liver. In fact, post prandial pharmacokinetic analyses suggested that miR-29b and miR-200c were absorbed in humans and, most importantly, the expression of RUNX2 (runt-related transcription factor 2), a target of miR-29b, was modulated after milk consumption indicating that microRNAs in milk may be considered as bioactive food compounds regulating gene expression and signal transduction in humans. Interestingly, the authors could also show that endogenous microRNA-synthesis cannot compensate for the milk-derived microRNAs in mice fed a microRNA depleted diet.27
Based on dietary surveys of European consumers, we conducted a literature search into the microRNA composition of commonly consumed food such as milk, meat, cereals and oil plants like rape. We searched Pubmed for studies up to 2014 and included 15 foods in our final analysis and if data were available we filtered the results for the 15 most abundant miRNAs present in the corresponding food items (Table 1).
| Food | Ref. | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||
| Pork (sus scrofa) | miRNA | 1 | 378 | 143-3p | 133a-3p | 30a-5p | 206 | let-7f | 148a | 10b | 127 | 140 | 30d | 542-3p | 21 | let-7a | 36 |
| Poultry (gallus gallus) | miRNA | 206 | let-7c | let-7j | 103 | let-7f | 130a | 107 | 130b | let-7b | 128 | 16c | 15b | 221 | 21 | 125b | 37 |
| Wheat (triticum aestivum) | miRNA | 156a | 168a | 167a | 166a | 156c | 1869 | 1074 | 894 | 2199 | 172a | 1132 | 2111-3p | 2911 | 166k | 1135 | 38 |
| Barley (hordeum vulgare) | miRNA | 168-5p | 156 | 166b/c | 5048 | 159a/b | 168-3p | 1120 | 5049 | 171 | 444b | 5052 | 397 | 1126 | 5050 | 1436 | 39 |
| Corn (zea mays) | miRNA | 319b | 319a | 319d | 319c | 171e | 171j | 171i | 166l | 166a | 171d | 166g | 166m | 166e | 166d | 166f | 40 |
| Rape (brassica napus) | miRNA | 156 | 159 | 166 | 172 | 158 | 171 | 319 | 824 | 160 | 167 | 827 | 165 | 403 | 395 | 390 | 41 |

|
|||||||||||||||||
| Beef1 (bos taurus) | miRNA | 133a | 26a | 99a | 103 | 150 | 574-3p | 411 | 107 | 652 | 423-5p | 122 | 42 | ||||
| Milk (raw) (bos taurus) | miRNA | 21 | 99a | 30d | 148a | 200c | 26a | 26b | 43 | ||||||||
| Broccoli (brassica oleracea) | miRNA | 398b | 319b | 395b | 394a | 414b | 162a | 158e | 44 | ||||||||
| Rice (oryza sativa) | miRNA | 1861e/g/k/m | 1861b/f/i/l | 164e | 167d/f–h/j | 397b | 519a–j | 5337 | 5338 | 45 | |||||||
| Apple (golden delicious) (malus domesticus) | miRNA | 396 | 159 | 858 | 482 | 167 | 399 | 164 | 165/166 | 172 | 162 | 171 | 156 | 535 | 168 | 395 | 46 |
| Orange (citrus sinensis) | miRNA | 172a | 164a | 166j | 167d | 479 | 156a | 167a | 162a | 159a | 473a | 164d | 156b | 399a | 482c | 171b | 47 |
| Potato (root) (solanum tuberosum) | miRNA | 164 | 165 | 162 | 168 | 171 | 167 | 159 | 48 | ||||||||
| Tomato (solanum lycopersicum) | 172 | 166 | 396 | 164 | 162 | 157 | 159 | 168 | 403 | 156 | 827 | 167 | 165 | 171 | 390 | 49 | |
| Grape (vitis vinifera) | 166 | 159 | 164 | 167 | 165 | 168 | 827 | 319 | 396 | 172 | 162 | 156 | 403 | 390 | 479 | 50 | |
Sophisticated tools (e.g. microRNA.org) allow the identification of corresponding mRNA targets of these food-derived microRNAs. It may be possible to predict the regulatory function of foods on the basis of their microRNA composition. However, it has to be taken into account that one microRNA normally targets several mRNAs and one mRNA can be targeted by several microRNAs. In addition it has to be considered that the biological activity of microRNAs does not necessarily correlate with their abundance but rather with their presence in exosomes. However before comprehensive food microRNA data bases are released, it needs to be assured if and to what extent microRNAs are absorbed in the gastrointestinal tract. Moreover, the mechanisms of intestinal absorption, bioavailability, tissue distribution and function of exogenous microRNAs should be established.
Conflict of interest
Anika E. Wagner, Stefanie Piegholdt, Martin Ferraro, Kathrin Pallauf and Gerald Rimbach declare that they have no conflict of interest.References
- G. Egger, G. Liang, A. Aparicio and P. A. Jones, Nature, 2004, 429, 457–463 CrossRef CAS PubMed.
- J. Winter, S. Jung, S. Keller, R. I. Gregory and S. Diederichs, Nat. Cell Biol., 2009, 11, 228–234 CrossRef CAS PubMed.
- C. B. Yoo and P. A. Jones, Nat. Rev. Drug Discovery, 2006, 5, 37–50 CrossRef CAS PubMed.
- M. A. Parasramka, E. Ho, D. E. Williams and R. H. Dashwood, Mol. Carcinog., 2012, 51, 213–230 CrossRef CAS PubMed.
- S. I. Ivashuta, J. S. Petrick, S. E. Heisel, Y. Zhang, L. Guo, T. L. Reynolds, J. F. Rice, E. Allen and J. K. Roberts, Food Chem. Toxicol., 2009, 47, 353–360 CrossRef CAS PubMed.
- S. A. Ross and C. D. Davis, Adv. Nutr., 2011, 2, 472–485 CrossRef CAS PubMed.
- D. Sayed and M. Abdellatif, Physiol. Rev., 2011, 91, 827–887 CrossRef CAS PubMed.
- S. Maciotta, M. Meregalli and Y. Torrente, Front Cell Neurosci., 2013, 7, 265 Search PubMed.
- A. S. Ali, S. Ali, A. Ahmad, B. Bao, P. A. Philip and F. H. Sarkar, Obes. Rev., 2011, 12, 1050–1062 CrossRef CAS PubMed.
- T. Melkman-Zehavi, R. Oren, S. Kredo-Russo, T. Shapira, A. D. Mandelbaum, N. Rivkin, T. Nir, K. A. Lennox, M. A. Behlke, Y. Dor and E. Hornstein, EMBO J., 2011, 30, 835–845 CrossRef CAS PubMed.
- C. Fernandez-Hernando, Y. Suarez, K. J. Rayner and K. J. Moore, Curr. Opin. Lipidol., 2011, 22, 86–92 CrossRef CAS PubMed.
- A. Cimmino, G. A. Calin, M. Fabbri, M. V. Iorio, M. Ferracin, M. Shimizu, S. E. Wojcik, R. I. Aqeilan, S. Zupo, M. Dono, L. Rassenti, H. Alder, S. Volinia, C. G. Liu, T. J. Kipps, M. Negrini and C. M. Croce, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 13944–13949 CrossRef CAS PubMed.
- G. Korkmaz, C. le Sage, K. A. Tekirdag, R. Agami and D. Gozuacik, Autophagy, 2012, 8, 165–176 CrossRef CAS PubMed.
- H. Zhu, H. Wu, X. Liu, B. Li, Y. Chen, X. Ren, C. G. Liu and J. M. Yang, Autophagy, 2009, 5, 816–823 CrossRef CAS.
- A. Bouchie, Nat. Biotechnol., 2013, 31, 577 CrossRef CAS PubMed.
- L. Garcia-Segura, M. Perez-Andrade and J. Miranda-Rios, J. Nutrigenet Nutrigenomics, 2013, 6, 16–31 CrossRef CAS PubMed.
- C. Boesch-Saadatmandi, A. Loboda, A. E. Wagner, A. Stachurska, A. Jozkowicz, J. Dulak, F. Doring, S. Wolffram and G. Rimbach, J. Nutr. Biochem., 2011, 22, 293–299 CrossRef CAS PubMed.
- A. E. Wagner, C. Boesch-Saadatmandi, J. Dose, G. Schultheiss and G. Rimbach, J. Cell Mol. Med., 2012, 16, 836–843 CrossRef CAS PubMed.
- S. Gaedicke, X. Zhang, C. Schmelzer, Y. Lou, F. Doering, J. Frank and G. Rimbach, FEBS Lett., 2008, 582, 3542–3546 CrossRef CAS PubMed.
- L. Zhang, D. Hou, X. Chen, D. Li, L. Zhu, Y. Zhang, J. Li, Z. Bian, X. Liang, X. Cai, Y. Yin, C. Wang, T. Zhang, D. Zhu, D. Zhang, J. Xu, Q. Chen, Y. Ba, J. Liu, Q. Wang, J. Chen, J. Wang, M. Wang, Q. Zhang, J. Zhang, K. Zen and C. Y. Zhang, Cell Res., 2012, 22, 107–126 CrossRef CAS PubMed.
- Z. Zhou, X. Li, J. Liu, L. Dong, Q. Chen, J. Liu, H. Kong, Q. Zhang, X. Qi, D. Hou, L. Zhang, G. Zhang, Y. Liu, Y. Zhang, J. Li, J. Wang, X. Chen, H. Wang, J. Zhang, H. Chen, K. Zen and C. Y. Zhang, Cell Res., 2015, 25, 39–49 CrossRef CAS PubMed.
- K. W. Witwer, J. Nutr., 2014, 144, 1880–1881 CrossRef CAS PubMed.
- Y. Zhang, B. E. Wiggins, C. Lawrence, J. Petrick, S. Ivashuta and G. Heck, BMC Genomics, 2012, 13, 381 CrossRef CAS PubMed.
- K. M. Howard, et al., J. Agric. Food Chem., 2015, 63, 588–592 CrossRef CAS PubMed.
- B. Dickinson, Y. Zhang, J. S. Petrick, G. Heck, S. Ivashuta and W. S. Marshall, Nat. Biotechnol., 2013, 31, 965–967 CrossRef CAS PubMed.
- J. W. Snow, A. E. Hale, S. K. Isaacs, A. L. Baggish and S. Y. Chan, RNA Biol., 2013, 10, 1107–1116 CrossRef CAS PubMed.
- S. R. Baier, C. Nguyen, F. Xie, J. R. Wood and J. Zempleni, J. Nutr., 2014, 144, 1495–1500 CrossRef CAS PubMed.
- A. Lukasik and P. Zielenkiewicz, PLoS One, 2014, 9, e99963 Search PubMed.
- B. C. Melnik, S. M. John and G. Schmitz, J. Transl. Med., 2014, 12, 43 CrossRef PubMed.
- K. D. Taganov, M. P. Boldin, K. J. Chang and D. Baltimore, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 12481–12486 CrossRef CAS PubMed.
- R. M. O'Connell, K. D. Taganov, M. P. Boldin, G. Cheng and D. Baltimore, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 1604–1609 CrossRef PubMed.
- S. Grasedieck, A. Sorrentino, C. Langer, C. Buske, H. Dohner, D. Mertens and F. Kuchenbauer, Blood, 2013, 121, 4977–4984 CrossRef CAS PubMed.
- Q. Zhou, M. Li, X. Wang, Q. Li, T. Wang, Q. Zhu, X. Zhou, X. Wang, X. Gao and X. Li, Int. J. Biol. Sci., 2012, 8, 118–123 CrossRef CAS.
- N. Kosaka, H. Izumi, K. Sekine and T. Ochiya, Silence, 2010, 1, 7 CrossRef PubMed.
- S. R. Baier, F. Xie and J. Zempleni, J. Nutr., 2014, 144, 1882 Search PubMed.
- L. Qin, Y. Chen, X. Liu, S. Ye, K. Yu, Z. Huang, J. Yu, X. Zhou, H. Chen and D. Mo, PLoS One, 2013, 8, e72418 CAS.
- T. Li, R. Wu, Y. Zhang and D. Zhu, BMC Genomics, 2011, 12, 186 CrossRef CAS PubMed.
- F. Meng, H. Liu, K. Wang, L. Liu, S. Wang, Y. Zhao, J. Yin and Y. Li, BMC Plant Biol., 2013, 13, 140 CrossRef PubMed.
- M. Hackenberg, P. J. Huang, C. Y. Huang, B. J. Shi, P. Gustafson and P. Langridge, DNA Res., 2013, 20, 109–125 CrossRef CAS PubMed.
- M. Kang, Q. Zhao, D. Zhu and J. Yu, BMC Genomics, 2012, 13, 360 CrossRef CAS PubMed.
- D. Huang, C. Koh, J. A. Feurtado, E. W. Tsang and A. J. Cutler, BMC Genomics, 2013, 14, 140 CrossRef CAS PubMed.
- W. Jin, J. R. Grant, P. Stothard, S. S. Moore and L. L. Guan, BMC Mol. Biol., 2009, 10, 90 CrossRef PubMed.
- X. Chen, C. Gao, H. Li, L. Huang, Q. Sun, Y. Dong, C. Tian, S. Gao, H. Dong, D. Guan, X. Hu, S. Zhao, L. Li, L. Zhu, Q. Yan, J. Zhang, K. Zen and C. Y. Zhang, Cell Res., 2010, 20, 1128–1137 CrossRef CAS PubMed.
- J. Wang, X. Yang, H. Xu, X. Chi, M. Zhang and X. Hou, Gene, 2012, 505, 300–308 CrossRef CAS PubMed.
- T. Peng, Q. Lv, J. Zhang, J. Li, Y. Du and Q. Zhao, J. Exp. Bot., 2011, 62, 4943–4954 CrossRef CAS PubMed.
- R. Xia, H. Zhu, Y. Q. An, E. P. Beers and Z. Liu, Genome Biol., 2012, 13, R47 CrossRef CAS PubMed.
- Q. Xu, Y. Liu, A. Zhu, X. Wu, J. Ye, K. Yu, W. Guo and X. Deng, BMC Genomics, 2010, 11, 246 CrossRef PubMed.
- W. Yang, X. Liu, J. Zhang, J. Feng, C. Li and J. Chen, Mol. Biol. Rep., 2010, 37, 3081–3087 CrossRef CAS PubMed.
- J. Zuo, B. Zhu, D. Fu, Y. Zhu, Y. Ma, L. Chi, Z. Ju, Y. Wang, B. Zhai and Y. Luo, BMC Genomics, 2012, 13, 7 CrossRef CAS PubMed.
- V. Pantaleo, G. Szittya, S. Moxon, L. Miozzi, V. Moulton, T. Dalmay and J. Burgyan, Plant J., 2010, 62, 960–976 CAS.
| This journal is © The Royal Society of Chemistry 2015 |