Protein binding and biological evaluation of a polymer-anchored cobalt(III) complex containing a 2,2′-bipyridine ligand
Gopalaswamy Vignesha,
Rajendran Senthilkumar*ab,
Preethy Paulbc,
Vaiyapuri Subbarayan Periasamycd,
Mohammad Abdulkadher Akbarshac and
Sankaralingam Arunachalam*a
aDepartment of Chemistry, Bharathidasan University, Tiruchirappalli, 620 024, India. E-mail: biosen81@gmail.com
bDepartment of Biosciences, Cell biology, ÅboAkademi University, Tykistokatu 6B, FI-20520 Turku, Finland
cDepartment of Animal Science, Bharathidasan University, Tiruchirappalli, 620 024, India
dDepartment of Food and Nutrition, King Saud University, Saudi Arabia
First published on 23rd October 2014
Abstract
The molecular interaction of polymer-anchored cobalt(III) complex, cis-[Co(bpy)2(BPEI)Cl]Cl2·4H2O (bpy = 2,2′-bipyridine, BPEI = branched polyethyleneimine) with two plasma proteins, human serum albumin (HSA) and bovine serum albumin (BSA) using various spectrophotomeric techniques has been investigated. The steady-state and time-resolved fluorescence spectra clearly demonstrated the static quenching mechanism. The calculated thermodynamic parameters revealed that the interaction between the polymer–cobalt(III) complex and HSA/BSA was driven mainly by van der Waals forces and hydrogen bonds. The results observed from three dimensional fluorescence and circular dichorism (CD) spectral studies manifested the conformational changes of HSA/BSA upon addition of the polymer–cobalt(III) complex. Furthermore, the antimicrobial result showed that the polymer–cobalt(III) complex exhibits good antibacterial and antifungal activities against certain human pathogenic microorganisms. In addition, the antiproliferative properties of the polymer–cobalt(III) complex on the HEp-2 human larynx cancer cells were determined using the MTT assay. The mode of cell death induced by the complex following treatment was analyzed adopting specific staining techniques. MTT assay revealed that the viability of the cells thus treated was significantly decreased and the cells succumbed to apoptosis as well as necrosis as reflected in changes in the nuclear morphology and cytoplasmic features by AO & EB and Hoechst staining methods.
Introduction
Serum albumins are the most abundant plasma proteins and their interaction with drug molecules has attracted great interest in recent years.1,2 The most important physiological function of albumins is their involvement in the transport of numerous endogenous and exogenous compounds such as fatty acids, amino acids and drugs etc.3 The binding affinity of drug to serum albumin is pivotal in the design of new drugs. The studies on the binding of drugs to albumins are of great importance to obtain information on structural features that determine the therapeutic effect of drugs. Among the serum albumins, bovine serum albumin (BSA) and human serum albumin (HSA) are the most extensively studied plasma proteins in biophysical research. The major difference between these two proteins is in the number and positioning of tryptophan residues. Crystal structure analyses have revealed that HSA contains 585 amino acid residues with 17 tyrosyl residues and only one tryptophan (Trp) located at position 214 along the chain (subdomain IIA); whereas, BSA contains 582 amino acid residues with 20 tyrosyl residues and two tryptophans located at positions 134 and 212 and Trp-134 at the surface of the molecule.4Larynx or laryngeal cancer is a type of head and neck malignancy. In 2014, an estimated 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 630 new cases of larynx cancer are expected to be diagnosed.5 In turn, the number of chemotherapy agents for the treatment of larynx cancer approved by the food and drug administration (FDA) is very low, even with a dwindling trend over the past decade, demonstrating an urgent need for the development of new classes of compounds for treating larynx cancer.
630 new cases of larynx cancer are expected to be diagnosed.5 In turn, the number of chemotherapy agents for the treatment of larynx cancer approved by the food and drug administration (FDA) is very low, even with a dwindling trend over the past decade, demonstrating an urgent need for the development of new classes of compounds for treating larynx cancer.
Significant developments have occurred in the recent years in the field of biopolymers and biomaterials, especially investigations of pharmacologically active polymers as drugs or carriers for normal pharmaceutical agents6 have created much interest. High molecular-weight polyethylene glycol (PEG), in conjugation with therapeutic proteins or small molecules, has been used and found to enhance the therapeutic value of anticancer agents by offering extended life in circulation, lower toxicity, increased drug stability and solubility.7 The use of the polymer-supported drugs for chemotherapy has the potential to deliver the active metal chelates at concentrations below that at which depilating side effects, particularly acute kidney damage, become apparent. It has been noticed that polymer–drug conjugates are potential candidates for the selective delivery of anticancer agents to tumor tissue and only a very few reports are available indicating the use of polymer–metal complexes as anticancer drugs.8–10
A polymer–metal complex is composed of a polymeric ligand and metal ion in which the metal ion is attached to the polymer ligand by a coordination bond. Its synthesis represents an attempt to impart inorganic function to an organic polymer. Many synthetic polymer–metal complexes have been shown in metallo-enzymes such as oxidases, and hemoglobin, where a metal complex is the active site, and the macromolecular protein part plays significant biological roles, or even controls the reactivity of the metal complex.11 Thus, research on the polymer–metal complexes has attracted considerable attention in the recent years. Polyethyleneimine (PEI) has been known as an efficient gene carrier with the highest cationic charge density potential.12 PEI possesses a number of properties, which make it a noteworthy complexing agent. It also appears to be a suitable polymeric matrix for use as a carrier in drug delivery systems.13 Further, research in respect of polyethyleneimine(PEI)–metal complexes might throw light on the biological processes in which they participate and unravel avenues for their application as therapeutics.
Previously, we have reported the DNA binding and apoptosis inducing properties of some polyethyleneimine–cobalt(III) and copper(II) complexes against various cell lines.14–16 In the present work, our aim is to explore the molecular interaction of one of these polymer–cobalt(III) complexes (with varying degree of coordination of cobalt(III) chelate in the polymer chain) with plasma proteins and also the cytotoxic and antimicrobial activities of this polymer–cobalt(III) complex against human larynx cancer cell lines and certain human pathogenic microorganisms.
Experimental
Materials
Human serum albumin (HSA), bovine serum albumin (BSA) and polyethyleneimine were obtained from Sigma-Aldrich and used as such. The cobalt(II) chloride hexahydrate and 2,2′-bipyridine were obtained from Loba, India. The polymer-anchored cobalt(III) complex (Fig. 1) with different degree of coordination of cobalt(III) in the polymer chain were synthesized according to our earlier report.14 The degree of coordination (x), defined as the number of moles of cobalt(III) chelate per mole of the repeating unit (amine group) of polymeric ligand, has been determined by the procedure adopted earlier.14 The degree of coordination (x), thus, obtained for our polymer–cobalt(III) complex samples are 0.115, 0.95, 0.65.The antimicrobial screening studies were carried out at Periyar College of Pharmaceutical Sciences, Tiruchirappalli, India, and the bacteria and fungus species were obtained from National Chemical Laboratory (NCL), Pune, India. Ciprofloxacin and clotrimazole discs were purchased from HiMedia Laboratories Pvt. Ltd., Mumbai, India. The human larynx cancer cell lines, HEp-2, was obtained from National Center for Cell Science (NCCS), Pune, India.
Methods
Spectroscopy measurements
Fluorescence experiments were carried out on a JASCO FP6500 spectrofluorometer using 1 cm quartz cell and a thermostat bath. The concentrations of HSA and BSA were determined spectrophotometrically using the extinction coefficient of 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 M−1 cm−1 and 43
500 M−1 cm−1 and 43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 M−1 cm−1 respectively at 280 nm in Tris–HCl buffer solution (pH = 7.4) refrigerating at 4 °C. The emission was measured from 290 nm to 450 nm with an excitation wavelength of 280 nm. The excitation and emission slits were fixed at 5 nm and 3 nm, and scanning speed to 500 nm min−1. The experiments were carried out at three different temperatures of (291, 296 and 301 K). The three dimensional fluorescence spectra were measured under the following conditions: the emission wavelength was recorded between 250–500 nm, the initial excitation wavelength was set to 250 nm with increments of 5 nm, the number of scanning curves was 14, and other scanning parameters were the same as those for the fluorescence quenching experiments. The fluorescence data were corrected for the inner filter effect using the following equation:5
800 M−1 cm−1 respectively at 280 nm in Tris–HCl buffer solution (pH = 7.4) refrigerating at 4 °C. The emission was measured from 290 nm to 450 nm with an excitation wavelength of 280 nm. The excitation and emission slits were fixed at 5 nm and 3 nm, and scanning speed to 500 nm min−1. The experiments were carried out at three different temperatures of (291, 296 and 301 K). The three dimensional fluorescence spectra were measured under the following conditions: the emission wavelength was recorded between 250–500 nm, the initial excitation wavelength was set to 250 nm with increments of 5 nm, the number of scanning curves was 14, and other scanning parameters were the same as those for the fluorescence quenching experiments. The fluorescence data were corrected for the inner filter effect using the following equation:5| Fcor = Fobs × e(Aex+Aem)/2 | (1) |
Fluorescence lifetimes were determined from time-resolved intensity decay by the method of nanosecond time correlated single photon counting (TCSPC) spectrometer using a nanosecond diode (Jobin-yvon, IBH with LED with resolution less than 1 ns, 280 nm) as a light source at room temperature. The data analysis was carried out by the software provided by IBH (DAS-6). The concentration of HSA/BSA and polymer–cobalt(III) complexes used were of 3 × 10−6 mol dm−3 and 6 × 10−6 mol dm−3 respectively.
The CD spectra were recorded on a JASCO spectrophotometer at room temperature with a quartz cell having a path length of 0.2 mm. The concentration of both HSA/BSA and polymer–cobalt(III) complex samples were fixed at 3 × 10−6 mol dm−3. The spectra of HSA/BSA in the presence and absence of complexes were recorded in the range of 200–250 nm and from this measurement the content of α-helix of HSA/BSA was calculated.
Anti-microbial assay
The in vitro antimicrobial screening of the polymer–cobalt(III) complex (x = 0.115) was carried out to find the effect of the polymer on certain human pathogenic bacteria and a fungus adopting disc-diffusion method.17 This method is highly suited for assay against rapidly growing microorganisms, and the activities of the complex are expressed by measuring the diameter of the zone of inhibition. The formation of inhibition zones reveals the dynamic interaction between the test drug and the microbes. The complex was stored dry at room temperature and dissolved in dimethylsulfoxide (DMSO) (1%). Both the Gram-positive (Staphylococcus aureus, Bacillus subtilis) and Gram-negative (Escherichia coli, Pseudomonas aeruginosa) bacteria were grown in nutrient agar medium and incubated at 37 °C for 48 h followed by frequent subculturing to fresh medium and were used as test bacteria. The yeast Candida albicans grown in Sabouraud dextrose agar medium, incubated at 27 °C for 72 h, followed by periodic sub-culturing to fresh medium was used as the test fungus. The Petri-dishes were inoculated with a loop full of bacterial or fungal culture and spread throughout the Petri-dishes uniformly using a sterile glass spreader. To each disc 200 μL of the polymer–cobalt(III) complex (10 μg mL−1) or the reference antibiotic ciprofloxacin (1 μg per disc for bacteria) or clotrimazole (10 μg per disc for fungus) was added using a sterile micropipette. The plates were then incubated at 35 ± 2 °C for 24–48 h and 27 ± 1 °C for bacteria and fungus, respectively. The plates with disc containing the respective solvents served as the control. The inhibition was recorded by measuring the diameter of the inhibitory zone after the period of incubation. All the experiments were repeated thrice and the average values are presented.Cell culture
HEp-2 human larynx cancer cells were was cultured as a monolayer with Roswell Park Memorial Institute Medium (RPMI-1640), supplemented with 10% fetal bovine serum (FBS) and 100 μg mL−1 of streptomycin as antibiotics (Himedia, Mumbai, India), at 37 °C, in a humidified atmosphere of 5% CO2, in a CO2 incubator (Heraeus, Hanau, Germany).MTT assay
The polymer–cobalt(III) complex (x = 0.115) was dissolved in DMSO, diluted in the culture medium and used to treat Hep-2 cells with the complex in the concentration range of 25 to 225 μg mL−1 for a period of 24 h and 48 h. DMSO was used as the solvent control. A miniaturized viability assay using 3-(4,5-di-methylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) was carried out according to the method described by Mosmann.18 The gross morphological changes in HEp-2 cells were observed and photographed at 24 h and 48 h after treatment. The treated cells were then assayed by the addition of 20 μL of 5 mg mL−1 MTT in phosphate-buffered saline (PBS). The plates were wrapped with aluminum foil and incubated for 4 h at 37 °C. The purple formazan product was dissolved by addition of 100 μL of 100% DMSO to each well. The absorbance was monitored at 570 nm (measurement) and 630 nm (reference) using a 96 well plate reader (Bio-Rad, Hercules, CA, USA). Data were collected for four replicates each and used to calculate the respective means. The percentage inhibition was calculated, from this data, using the formula: percentage of inhibition = ((mean optical density of untreated cells (control) − mean optical density of complex treated cells)/mean optical density of untreated cells (control)) × 100.The IC50 value was determined as the concentration of the complex that is required to reduce the absorbance to half that of the control.
Hoechst 33258 staining19
The cell pathology was detected by staining the nuclear chromatin of trypsinized cells (4.0 × 104 per mL) with 1 μL of Hoechst 33258 (1 mg mL−1, aqueous) for 10 min at 37 °C. A drop of cell suspension was placed on a glass slide and a cover-slip was laid over to reduce light diffraction. At random 300 cells were observed in a fluorescent microscope (Carl Zeiss, Germany) fitted with a 377–355 nm filter and observed at 400× magnification, and the percentage of cells reflecting pathological changes was calculated. Data were collected for four replicates and used to calculate the mean and the standard deviation.Acridine orange (AO) and ethidium bromide (EB) staining
Acridine orange and ethidium bromide staining was performed as described.20 Twenty five microliters of cell suspension of each sample, containing 5 × 105 cells, was treated with AO and EB solution (1 part of 100 μg per mL AO and 1 part of 100 μg per mL EO in PBS) and examined in a fluorescent microscope (Carl Zeiss, Germany) using an UV filter (450–490 nm). Three hundred cells per sample were counted in tetraplicate for each dose point. Cells were scored as viable, apoptotic or necrotic as judged by the staining pattern, nuclear morphology and membrane integrity. The percentages of apoptotic and necrotic cells were then calculated. Morphological changes were also observed and photographed.Result and discussion
Analysis of fluorescence quenching of HSA/BSA by polymer–cobalt(III) complex
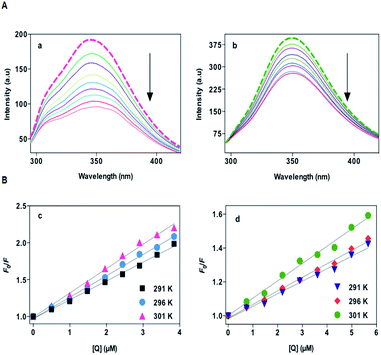
Proteins are considered to have intrinsic fluorescence mainly due to tryptophan (Trp), tyrosine (Try) and phenylalanine (Phe) residues. Among these the intrinsic fluorescence of HSA/BSA is mainly due to tryptophan, because phenylalanine has a very low quantum yield and the fluorescence of tyrosine is almost totally quenched, if it is ionized or near an amino group (or) tryptophan residues.21 A variety of molecular interactions can result in fluorescence quenching of the intrinsic fluorophores because of molecular rearrangement, energy transfer, ground state complex formation (or) collision quenching. The quenching mechanisms are usually classified as dynamic and static quenching, which can be distinguished by their differing dependence on temperature and viscosity or preferably by life time measurements. For dynamic quenching, when the temperature of the system rises, the effective collision times between molecules, the energy transfer efficiency, and fluorescence quenching constant also get increased. In contrast, the increase of temperature is likely to result in decreased stability of complexes, thus the value of the static quenching constant is expected to be smaller.22 Fig. 2A shows the fluorescence spectrum of HSA/BSA in the absence and presence of our polymer–cobalt(III) complex.It is observed that the fluorescence intensity of BSA/HSA decreases regularly with the increase in concentration of our polymer–cobalt(III) complex, which indicates that changes in the microenvironment have occurred around the single tryptophan residue Trp 214 within the hydrophobic pocket of sub-domain IIA of HSA/BSA.
To confirm the quenching mechanism the fluorescence quenching data was analyzed according to the Stern–Volmer equation23
| F0/F = 1 + Ksv[Q] = 1 + kqτ0[Q] | (2) |
| T (K) | x = 0.115 | x = 0.095 | x = 0.065 | |||
|---|---|---|---|---|---|---|
| kq (× 1012 M−1 s−1) | kq (× 1012 M−1 s−1) | kq (× 1012 M−1 s−1) | ||||
| HSA | BSA | HSA | BSA | HSA | BSA | |
| a x – degree of coordination. | ||||||
| 291 | 2.255 | 1.086 | 0.658 | 0.643 | 0.729 | 0.235 |
| 296 | 2.914 | 1.136 | 0.783 | 0.707 | 0.801 | 0.238 |
| 301 | 3.300 | 1.500 | 0.119 | 0.744 | 0.857 | 0.252 |
In order to confirm the static quenching process, life time measurements for HSA/BSA in the presence of polymer–cobalt(III) complex samples were carried out. Fig. 3 shows the fluorescence decay profiles of HSA/BSA in the absence and presence of polymer–cobalt(III) complex. There are two lifetimes for native HSA/BSA (τ1 and τ2) and the mean fluorescence lifetime was chosen as an important parameter for exploring the behaviour of HSA/BSA rather than placing too much attention on the magnitude of individual decay constants in the biexponential decay. The decay curves are well fitted by χ2 values, and residuals mean (average) fluorescence lifetime (τ) for biexponential iterative fitting was calculated from the decay times and the pre-exponential factor using the following relation:26
| 〈τ〉 = α1τ1 + α2τ2 | (3) |
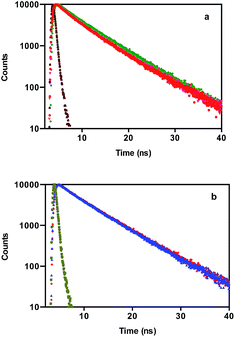 | ||
| Fig. 3 The time-resolved fluorescence decay of HSA (a) and BSA (b) in the absence and presence polymer–cobalt(III) complex. [HSA/BSA] = 3 × 10−6 M, [polymer–cobalt(III) complex] = 6.10 × 10−6 M. | ||
The life times (τ1 and τ2), relative amplitudes (α1 and α2) and χ2 values for the HSA/BSA–polymer–cobalt(III) complex samples are listed in Table 2.
| Samples | τ1 (ns) | τ2 (ns) | α1 (ns) | α2 (ns) | 〈τ〉 (ns) | χ2 |
|---|---|---|---|---|---|---|
| a x – degree of coordination. | ||||||
| HSA | 2.45 | 6.37 | 0.24 | 0.76 | 5.43 | 1.057 |
| HSA + x(0.115) | 2.45 | 6.37 | 0.26 | 0.74 | 5.35 | 0.988 |
| HSA | 2.94 | 6.37 | 0.23 | 0.77 | 5.58 | 1.125 |
| HSA + x(0.095) | 2.94 | 6.37 | 0.27 | 0.73 | 5.44 | 1.010 |
| HSA | 2.94 | 6.35 | 0.17 | 0.83 | 5.77 | 1.127 |
| HSA + x(0.065) | 2.64 | 6.30 | 0.14 | 0.86 | 5.79 | 1.221 |
| BSA | 2.88 | 6.47 | 0.11 | 0.89 | 6.47 | 1.177 |
| BSA + x(0.115) | 3.18 | 6.51 | 0.20 | 0.80 | 6.51 | 1.182 |
| BSA | 2.58 | 6.40 | 0.12 | 0.88 | 5.94 | 1.168 |
| BSA + x(0.095) | 2.49 | 6.32 | 0.11 | 0.89 | 5.90 | 1.166 |
| BSA | 2.94 | 6.45 | 0.15 | 0.85 | 5.92 | 1.081 |
| BSA + x(0.065) | 2.55 | 6.26 | 0.11 | 0.89 | 5.85 | 1.082 |
From Fig. 2 and Table 2, it is found that the curves fit well to a biexponential function with acceptable χ2 values. The results illustrate that the average lifetime of HSA/BSA did not change in the presence of polymer–cobalt(III) complex samples indicating static quenching as the mechanism.27
Binding constant and the number of binding sites
For the static quenching, when molecules bind independently to a set of equivalent sites on macromolecules, the binding constant (Kb) and the number of binding sites (n) can be determined by the following equation:28
log(F0 − F)/F = log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb + n Kb + n![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log[Q] log[Q]
| (4) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) F0 − F/F versus log[Q] (Fig. 4) could be used to determine Kb and n. The Kb and n at different temperatures are listed in Table 3.
F0 − F/F versus log[Q] (Fig. 4) could be used to determine Kb and n. The Kb and n at different temperatures are listed in Table 3.
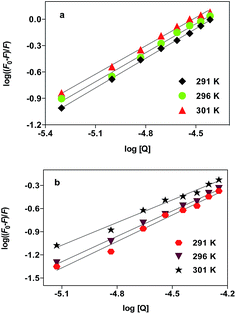 | ||
| Fig. 4 Plot of log[(F0 − F)/F] vs. log[Q] for HSA (a) /BSA (b) polymer–cobalt(III) complex (x = 0.115) system at different temperatures. | ||
| T (K) | x = 0.115 | x = 0.095 | x = 0.065 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Kb (× 105 M−1) | n | R | Kb (× 105 M−1) | n | R | Kb (× 105 M−1) | n | R | |
| a x – degree of coordination, R – correlation coefficient. | |||||||||
| HSA | |||||||||
| 291 | 0.880 | 1.12 | 0.999 | 0.122 | 1.11 | 0.994 | 0.011 | 0.82 | 0.984 |
| 296 | 0.572 | 1.06 | 0.996 | 0.111 | 1.07 | 0.994 | 0.037 | 0.92 | 0.997 |
| 301 | 0.351 | 1.02 | 0.998 | 0.104 | 1.02 | 0.996 | 0.146 | 1.05 | 0.992 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||||
| BSA | |||||||||
| 291 | 0.287 | 1.14 | 0.991 | 0.159 | 1.09 | 0.997 | 0.043 | 0.83 | 0.996 |
| 296 | 0.127 | 1.09 | 0.982 | 0.101 | 1.04 | 0.994 | 0.050 | 0.84 | 0.992 |
| 301 | 0.081 | 0.97 | 0.993 | 0.063 | 0.98 | 0.992 | 0.071 | 0.87 | 0.997 |
The calculated Kb values suggest that a moderate affinity existed between HSA/BSA and polymer–cobalt(III) complex, and the values of Kb decreased with increasing temperature indicating that the capacity of polymer–cobalt(III) complex (x = 0.115) binding to HSA/BSA was reduced. The increasing temperature results in the increasing diffusion coefficient and the reduction of stability of the polymer–cobalt(III) complex–HSA/BSA complex. The high linear correlation coefficient (R > 0.98) at different temperatures indicated that eqn (4) is valid. The polymer–cobalt(III) complex with higher degree of coordination shows higher binding constant values compared to those of the corresponding complexes with lower degree of coordination. This is due to the presence of a large number of high positively charged cobalt(III) complex units in the polymer chain, which cooperatively act to increase the overall binding ability of each polymer–cobalt(III) complex unit to HSA/BSA. The value of n are approximately equal to 1 suggesting that there is a one binding site on HSA/BSA for polymer–cobalt(III) complex samples.
Thermodynamic analysis
In general, binding forces between quenchers and macromolecules are non-covalent in nature. There are essentially four types of interaction forces that are involved in macromolecule–ligand interactions which are hydrogen bonding, van der Waals forces, electrostatic forces and hydrophobic forces. The thermodynamic parameters like enthalpy change, entropy change and free energy change of binding will reveal the binding mode of the interaction. On the basis of the sign of the thermodynamic parameters, the probable interaction forces involved in the binding can be elucidated.29 These are(1) ΔH° > 0 and ΔS° > 0, hydrophobic.
(2) ΔH° < 0 and ΔS° < 0, van der Waals interaction and hydrogen bonds.
(3) ΔH° < 0 and ΔS° > 0, electrostatic interaction.
If the enthalpy change (ΔH°) does not vary significantly over the temperature range studied, the enthalpy change (ΔH°), free-energy change (ΔG°) and the entropy change (ΔS°) can be calculated based on the van't Hoff and thermodynamic equations30
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K = ΔH°/2.303RT − ΔS°/2.303R K = ΔH°/2.303RT − ΔS°/2.303R
| (5) |
ΔG° = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K K
| (6) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K vs. 1/T (Fig. 5). The negative value of ΔH° and ΔS° (Table 4) suggest that the hydrogen bonds and/or van der Waals interaction played a major role in the binding of polymer–cobalt(III) complex to HSA/BSA and contributed to the stability of the complex.
K vs. 1/T (Fig. 5). The negative value of ΔH° and ΔS° (Table 4) suggest that the hydrogen bonds and/or van der Waals interaction played a major role in the binding of polymer–cobalt(III) complex to HSA/BSA and contributed to the stability of the complex.
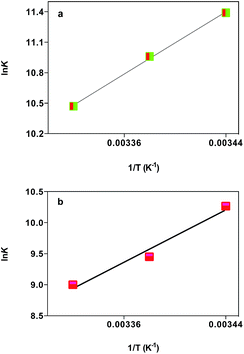 | ||
| Fig. 5 van't Hoff plot for the binding of polymer–cobalt(III) complex (x = 0.115) with HSA (a) and BSA (b). | ||
| T (K) | ΔH° (kJ mol−1) | ΔG° (kJ mol−1) | ΔS° (J mol−1 K−1) | |||
|---|---|---|---|---|---|---|
| HSA | BSA | HSA | BSA | HSA | BSA | |
| 291 | −27.55 | −24.84 | ||||
| 296 | −63.74 | −87.66 | −26.96 | −23.01 | −124.49 | −216.72 |
| 301 | −26.19 | −22.53 | ||||
Three dimensional fluorescence studies
Three dimensional fluorescence spectroscopy is one of the most recently developed methods for providing conformational and structural information of proteins. Also, it gives information regarding the fluorescence characteristics which can be acquired by simultaneously changing the excitation and emission of wavelengths.30 Fig. 6 shows the three dimensional fluorescence spectra of HSA/BSA in presence of polymer–cobalt(III) complex. It is observed that the fluorescence intensity of Rayleigh scattering peak (peak 1) increased in the presence of complex due to the increase in the macromolecule diameter. There is another one strong fluorescence peak 2 (λex < λem) which mainly reveals the spectral behavior of tryptophan and tyrosine residues. The maximum emission wavelength and fluorescence intensity of the residues closely correlates with the microenvironment polarity.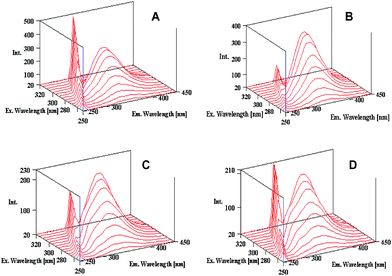 | ||
| Fig. 6 Three-dimensional fluorescence spectra of BSA (A), BSA–polymer–cobalt(III) complex (B), HSA (C), HSA–polymer–cobalt(III) (D). [HSA = BSA = polymer–cobalt(III) complex = 1 × 10−5 M]. | ||
As seen from Fig. 6, the intensity of the peak 2 decreased which indicates that the polarity of both the residues decreases and more amino residues of HSA/BSA were buried in the hydrophobic pockets. This result in combination with synchronous fluorescence results reveals that the complexes induce changes in the polarity of microenvironment and the conformation of the BSA/HSA.31
Circular dichroism
Circular dichroism (CD) spectroscopy is widely used to monitor the confirmation of the protein in solution.32 The CD spectrum of BSA/HSA exhibits two negative bands in the far UV region at 208 and 222 nm, which represent the typical α-helix structure of protein. The CD results are expressed in terms of mean residue ellipticity (MRE) in degree cm2 dmol−1 according to following equation32
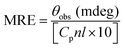 | (7) |
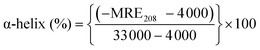 | (8) |
where Cp is the molar concentration of the protein, n is the number of amino acid residues and l is the path length. Fig. 7 illustrates the CD spectra of HSA/BSA in the presence of polymer–cobalt(III) complex. From this figure, it is observed that intensity of the band decreased without any shift in the presence of polymer–cobalt(III) complex. This unalteration of the position of band shows that even after binding of polymer–cobalt(III) complex, α-helix is still predominant conformation of HSA/BSA. In order to quantify the content of the α-helix of BSA/HSA, the changes in the intensity have been analyzed by the above equations and it is found that α-helix content of free HSA decreases from 62.42% to 51.96% (HSA–polymer–cobalt(III) complex) and of free BSA decreases from 50.06% to 32.74% (BSA–polymer–cobalt(III) complex). This reduction in the percentage of α-helix structure of the protein upon binding indicates that the polymer–cobalt(III) complex binds to main polypeptide chain of the proteins and destroy their hydrogen bonding network.
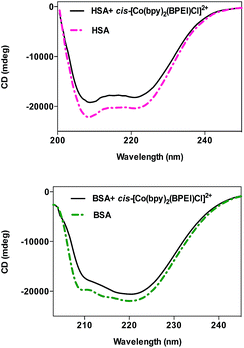 | ||
| Fig. 7 The CD spectra of HAS/BSA in the absence and presence of polymer–cobalt(III) complex (x = 0.115). | ||
Antimicrobial studies
The polymer–cobalt(III) complex (x = 0.115) was screened for in vitro anti-microbial activity against certain human pathogenic bacterial and a fungal species using disc diffusion method, and the results are summarized in Table 5. The complex was found to exhibit considerable activity against Gram-positive bacteria Staphylococcus aureus and Bacillus subtilis and the Gram-negative bacterium Escherichia coli though not as pronounced as the standard drugs, where as there was no effect on Pseudomonas aeruginosa The polymer–cobalt(III) complex was also very active against the fungus C. albicans. The polymer–cobalt(III) complex showed higher activity against Gram positive than Gram negative bacteria. This antimicrobial activity may be due to an efficient diffusion of the metal complexes into the bacterial/fungal cells and/or interaction with these organisms.33 It was reported by Nagababu et al.,34 that cobalt(III) complexes with bipyridine, 1,10-phenanathroline, dimethylimidazodrevcele and pyrazole complexes possess antibacterial activity. The cobalt(III) complexes phenanathroline/bipyridine and benzoylhydrazones inhibited the growth of E. coli cells and the activity was found to be concentration- and substituent-dependent.35 In another study the surfactant–cobalt(III) complexes with bipyridyl, phenanthroline and triethylenetetramine ligands showed antimicrobial activity against various microorganisms.36,37 Thus, cobalt(III) complexes in general, and our polymer–cobalt(III) complex in particular, may attract application as antimicrobial substances.| Test organisms | Diameter of zone inhibition (mm) | |
|---|---|---|
| cis-[Co(bpy)2(BPEI)Cl]Cl2·4H2O | Standard | |
| a Standard – ciprofloxacin for bacteria and clotrimazole for fungus. Solvent – DMSO (showed nil effect against the microorganisms under test). NS – not seen any activity. | ||
| Staphylococcus aureus | 10 | 16 |
| Bacillus subtillis | 12 | 17 |
| Escherichia coli | 11 | 16 |
| Pseudomonas auregenosa | NS | 17 |
| Candida albicans | 10 | 12 |
Anticancer studies
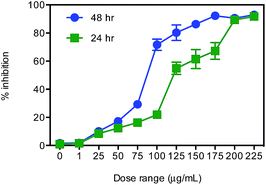
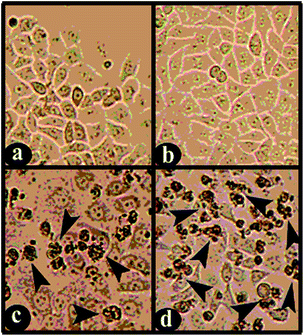
The cytotoxic property of the polymer–cobalt(III) complex (x = 0.115) was examined on cultured HEp-2 human larynx cancer cells by exposing cells for 24 and 48 h to the medium containing the complex at 25–225 μg mL−1 concentration (Fig. 8). Bright field microscopic analysis of control and treated cells revealed that the polymer–cobalt(III) complex inhibited the growth of larynx cancer cells to significant levels, in dose- and duration-dependent manner (Fig. 9). The cytotoxic activity was determined according to the dose values of the exposure of the complex required to reduce survival of the cells to 50% (IC50), compared to untreated cells. The IC50 values of the tested polymer–cobalt(III) (x = 0.115) is 110–125 μg mL−1 (24 h) and 75–100 μg mL−1 (48 h). The polymer–cobalt(III) complex showed highly effective cytotoxic activity against the larynx cancer cells, and the IC50 value of the complex was lesser for the 48 h treatment groups than for the 24 h treatment groups, revealing the dose and the duration-dependence.After treatment of HEp-2 cancer cells with polymer–cobalt(III) complex(x = 0.115) at the respective IC50 concentrations for 24 and 48 h, the cells were observed for the gross cytological changes. The observations revealed that the complex brought about cytological changes such as compaction, chromatin fragmentation, binucleation, cytoplasmic vacuolation, nuclear swelling, cytoplasmic blebbing and late apoptosis indication of chromatin condensation and dot-like chromatin (Fig. 10). These cytological changes indicated that the cells were committed to specific modes of cell death.
To find the mode of cell death, we used AO and EB staining and observed the cells in a fluorescent microscope. The cells were classified into four types according to the fluorescence emission and features of the chromatin. (i) Viable cells had uniformly green fluorescing nuclei with a highly organized structure (Fig. 11A, a and b). (ii) Early apoptotic cells (which still had intact membranes but had started undergoing DNA fragmentation) had green fluorescing nuclei, but peri-nuclear chromatin condensation was visible as bright green patches or fragments (Fig. 11A, c and d). (iii) Late apoptotic cells had orange to red fluorescing nuclei with condensed or fragmented chromatin (Fig. 11A, c and d). (iv) Necrotic cells had uniformly orange to red fluorescing nuclei with no indication of chromatin fragmentation and the cells were swollen to large size (Fig. 11A, c and d). The results suggest that polymer–cobalt(III) complex treatment caused cell death both through apoptosis and necrosis, but a higher percentage of necrotic cell death was observed compared to apoptotic cell death. Data collected from the manual counting of cells with normal and abnormal nuclear features are shown in the Fig. 11B. Cells with apoptotic as well as necrotic features increased in duration-dependent manner.
One of the mechanisms underlying the efficiency of the cobalt(III)complex in its cytotoxic property would be derived from its amphiphilic nature since the complex has the capacity to penetrate cell membrane easily, reduce the energy status in tumors and also bring about hypoxia state in the cancer cell microenvironment, factors which would influence the cytotoxic activity.38 Complexes of Co(III) are kinetically inert octahedral coordination complexes. This inertness is due to the d6 low spin electron configuration of trivalent cobalt.39 The biological consequence of kinetic inertness is that many Co(III) complexes will remain intact when added to a culture medium or injected into animals and should arrive at their cellular targets with their ligand configuration intact.40 A substantial amount of indirect evidence suggests that the induction of apoptosis in cancer cells is critical to their successive elimination by therapeutic agents.41 Second, since the complex has bipyridine ligand, it would bind efficiently with DNA and bring about double as well as single strand breaks which can initiate the events in cell death leading to apoptosis.14 Third, the complex can be capable of inducing hypoxic status in the cells.42 Hypoxia causes inhibition of oxidative phosphorylation and a switch to glycolytic metabolism, resulting in decreased levels of high-energy phosphates, increased lactic acid production, and lower intracellular pH.43–45 If ATP is rapidly depleted, necrosis will occur because of passive loss of trans-membrane ion gradients, followed by cell swelling which in turn can bring about necrotic changes in the cells.44,45 In as much as apoptosis is the principal molecular mechanism of drug-induced cell death required of chemotherapy for cancer, recently, necrosis is considered equally important in cancer therapy. Recent studies have suggested that in response to a given death stimulus, there is often a continuum of apoptosis and necrosis. Many insults induce apoptosis at lower doses and necrosis at higher doses. Thus, programmed cell necrosis is an important cell fate as apoptosis.46–48
Conclusions
The interaction of water soluble branched polyethyleneimine-anchored cobalt(III) complexes (with varying degree of coordination of cobalt(III) chelate containing bipyridine) with HSA and BSA was studied by absorption, fluorescence, lifetime measurement and circular dichromism spectroscopic techniques. The polymer–cobalt(III) complex strongly quenches the intrinsic fluorescence of HAS/BSA by static quenching mechanism through van der Waals force and/or hydrogen bonds. The three dimensional fluorescence and circular dichroism studies indicate that the binding has induced considerable amount of conformational changes in the protein. The antimicrobial result shows that that our polymer–cobalt(III) complex is in general capable of inhibiting the growth of bacteria and fungi to a fairly good extent. In bringing about induction of apoptosis at lower doses treated for a shorter duration and necrosis at higher doses treated for a longer duration, our polymercobalt(III) complex containing 2,2′-bipyridyl ligand has the potential to be developed as a cancer chemotherapeutic agent, provided serious side effects, if any, are eliminated in in vivo testing. Further study to clarify the molecular mechanisms of the polymer–cobalt(III) complex, is currently in progress and the finding will be the subject of future publications.Notes and references
- J. Zhang, S. Zhuang, C. Tong and W. Liu, J. Agric. Food Chem., 2013, 61, 7203 CrossRef CAS PubMed.
- M. Mathew, S. Sreedhanya, P. Manoj, C. T. Aravindakumar and U. Aravind, J. Phys. Chem. B, 2014, 118, 3832 CrossRef CAS PubMed.
- G. Fanali, A. di Masi, V. Trezza, M. Marino, M. Fasano and P. Ascenzi, Mol. Aspects Med., 2012, 33, 209 CrossRef CAS PubMed.
- T. Peters Jr, All about albumin: biochemistry, genetics and medical applications, Academic Press, San Diego, CA, 1996 Search PubMed.
- R. Siegel, J. Ma, Z. Zou and A. Jemal, Ca–Cancer J. Clin., 2014, 64, 9 CrossRef PubMed.
- J. J. Grodzinski, Polym. Adv. Technol., 2006, 17, 395 CrossRef.
- G. Pasut and F. M. Veronese, Adv. Drug Delivery Rev., 2009, 61, 1177 CrossRef CAS PubMed.
- M. R. Roner, K. R. Shahi, G. Barot, A. Battin and C. E. Carraher, J. Inorg. Organomet. Polym., 2009, 19, 410 CrossRef CAS.
- D. S. Raja, E. Ramachandran, N. S. Bhuvanesh and K. Natarajan, Eur. J. Med. Chem., 2013, 64, 148 CrossRef PubMed.
- M. Callari, J. R. Aldrich-Wright, P. L. de Souza and M. H. Stenzel, Prog. Polym. Sci., 2014, 39, 1614 CrossRef CAS PubMed.
- M. Kaneko and E. Tsuchida, J. Polym. Sci., 1981, 16, 397 CAS.
- A. von Zelewsky, L. Barbosa and C. W. Schlapfer, Coord. Chem. Rev., 1993, 123, 229 CrossRef CAS.
- C. H. Lee, Y. H. Ni, C. C. Chen and C. K. Chou, Biochim. Biophys. Acta, 2003, 1611, 55 CrossRef CAS.
- R. Senthil Kumar and S. Arunachalam, Polyhedron, 2006, 25, 3113 CrossRef PubMed.
- R. Senthil Kumar, V. S. Periasamy, C. P. Paul, A. Riyasdeen, S. Arunachalam and M. A. Akbarsha, Med. Chem. Res., 2011, 20, 726 CrossRef CAS.
- J. Lakshmipraba, S. Arunachalam, D. A. Gandi and T. Thirunalasundari, Eur. J. Med. Chem., 2011, 46, 3013 CrossRef CAS PubMed.
- W. Bauer, N. M. Kirby, J. C. Sherris and M. Turck, Am. J. Clin. Pathol., 1966, 45, 493 Search PubMed.
- T. Mosmann, J. Immunol. Methods, 1983, 65, 55 CrossRef CAS.
- G. P. Kasibhatla, D. Finucane, T. Brunner, E. B. Wetzel and D. R. Green, Cell: A laboratory manual culture and biochemical analysis of cells, CSHL Press, 2000 Search PubMed.
- D. L. Spector, in Cells: A laboratory manual, ed. R. D. Goldman and L. A. Leinwand, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 1998 Search PubMed.
- K. Osama, A. Zied, K. I. Othman and A. I. Shihi, J. Am. Chem. Soc., 2008, 130, 10793 CrossRef PubMed.
- F. Ding, G. Y. Zhao, S. C. Chen, F. Liu, Y. Sun and L. Zhang, J. Mol. Struct., 2009, 929, 159 CrossRef CAS PubMed.
- M. R. Eftink, Fluorescence quenching reactions: Probing biological macromolecular structures, in Biophysical and Biochemical aspects of fluorescence Spectroscopy, ed. T. G. Eewey, Plenum Press, New York, 1991, p. 1 Search PubMed.
- J. R. Lackowicz and G. Weber, Biochemistry, 1973, 12, 4161 CrossRef.
- W. R. Ware, J. Phys. Chem., 1962, 66, 455 CrossRef CAS.
- U. Anand, C. Jash and S. Mukherjee, J. Phys. Chem. B, 2010, 114, 15839 CrossRef CAS PubMed.
- X. Wu, J. Liu, H. Huang, W. Xue, X. Yao and J. Jin, Int. J. Biol. Macromol., 2011, 49, 343 CrossRef CAS PubMed.
- Y. J. Hu, Y. Liu, R. M. Dong and S. S. Qu, J. Photochem. Photobiol., A, 2006, 179, 324 CrossRef CAS PubMed.
- P. D. Ross and S. Subramanian, Biochemistry, 1981, 20, 3096 CrossRef CAS.
- X. Guo, L. Zhang, X. Sun, X. Han, C. Guo and P. Kang, J. Lumin., 2009, 923, 114 Search PubMed.
- V. D. Suryawanshi, P. V. Anbhule, A. H. Gore, S. R. Patil and G. B. Kolekar, Ind. Eng. Chem. Res., 2012, 51, 95 CrossRef CAS.
- J. N. Tian, J. Q. Liu, Z. D. Hu and X. G. Chen, Bioorg. Med. Chem., 2005, 13, 4124 CrossRef CAS PubMed.
- Z. H. Chohan, M. U. Hassan, K. M. Khan and C. T. Supuran, J. Enzyme Inhib. Med. Chem., 2005, 20, 183 CrossRef CAS PubMed.
- P. Nagababu, J. N. Latha, P. Pallavi, S. Harish and S. Satyanarayana, Can. J. Microbiol., 2006, 52, 1247 CrossRef CAS PubMed.
- S. Srinivasan, J. Annaraj and P. R. Athappan, J. Inorg. Biochem., 2005, 99, 876 CrossRef CAS PubMed.
- R. Senthil Kumar and S. Arunachalam, Biophys. Chem., 2008, 136, 136 CrossRef PubMed.
- R. Senthil Kumar, S. Arunachalam, V. S. Periasamy, C. P. Preethy, A. Riyasdeen and M. A. Akbarsha, J. Inorg. Biochem., 2009, 103, 117 CrossRef PubMed.
- D. C. Ware, B. D. Palmer, W. R. Wilson and W. A. Denny, J. Med. Chem., 1993, 36, 1839 CrossRef CAS.
- F. Cotton and G. Wilkinson, Advanced Inorganic Chemistry, Interscience, New York, 3rd edn, 1972, p. 883 Search PubMed.
- B. A. Teicher, M. J. Abrams, K. W. Rosbe and T. S. Herman, Cancer Res., 1990, 50, 6971 CAS.
- J. C. Reed, Am. J. Pathol., 2000, 157, 1415 CrossRef CAS.
- R. K. Martin, Exp. Biol. Med., 2006, 231, 117 Search PubMed.
- D. G. Allen and C. H. Orchard, Circ. Res., 1987, 60, 153 CrossRef CAS.
- D. G. Allen, J. A. Lee and G. L. Smith, J. Physiol., 1989, 410, 297 CAS.
- J. R. Neely and L. W. Grotyohann, Circ. Res., 1984, 55, 816 CrossRef CAS.
- L. M. Buja, M. L. Eigenbrod and E. H. Eigenbrodt, Arch. Pathol. Lab. Med., 1993, 117, 1208 CAS.
- G. Majno and S. Joris, Am. J. Pathol., 1995, 146, 3 CAS.
- W. Zong and C. B. Thompson, Genes Dev., 2006, 20, 1 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2014 |