Lanthanide-doped nanoparticles for specific recognition of toll-like receptor (TLR) in human neutrophils†
Abstract
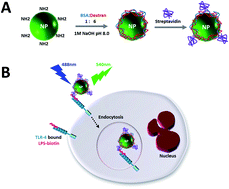
KYF4:Yb,Er nanoparticles were fabricated and functionalized with streptavidin for the recognition of toll-like receptors (TLRs) on human neutrophil through biotinylated lipopolysaccharide (biotin–LPS). X-Ray diffraction, Infrared spectroscopy, Transmission Electron Microscopy and Agarose Gel Electrophoresis were used to characterize the as-prepared and functionalized nanoparticles. Confocal Scanning Laser Microscopy was applied to study the TLR specific recognition using KYF4:Yb,Er and the uptake of the nanoparticles functionalized with BSA/Dextran–streptavidin in the presence of biotin–LPS by human neutrophils under normal (37 °C) and cell stressing conditions (4 °C). Confocal microscopy studies showed that the uptake of the functionalized KYF4:Yb,Er nanoparticles occurs faster and with a higher rate at 37 °C compared to the uptake at 4 °C, indicating that the uptake mechanism is an energy dependent process. Research reported in this work provides relevant guidance for the development of lanthanide doped KYF4:Yb,Er nanoparticles as specific intracellular probes, allowing control of the nanoparticle–cell interactions by tuning the surface properties. Furthermore, our results would pave the way to the use of lanthanide-doped materials for imaging cellular processes such as uptake of nanoparticles and recognition of receptors involved in immune responses.

 Please wait while we load your content...
Please wait while we load your content...