Magnetic mesoporous silica incorporated with TiO2 for selective and rapid capture of peptides†
Gong
Cheng
abc,
Yong
Wang
a,
Zhi-Gang
Wang
bc,
Xiao-Jing
Sui
abc,
Ji-Lin
Zhang
*b and
Jia-Zuan
Ni
*ab
aKey Laboratory of Marine Bioresources and Ecology, College of Life Science, Shenzhen University, Shenzhen 518060, China. E-mail: jzni@ciac.ac.cn
bState Key Laboratory of Rare Earth Resource Utilization, Changchun Institute of Applied Chemistry, Chinese Academy of Sciences, Changchun 130022, China. E-mail: zjl@ciac.ac.cn
cUniversity of Chinese Academy of Sciences, Beijing 100049, PR China
First published on 8th January 2014
Abstract
Exploration of valuable information from peptide biomarkers in biological fluids for early diagnosis and elucidation of disease has drawn increasing attention in clinical biomedicine. In this work, a novel affinity probe consisting of magnetic core and mesoporous silica shell incorporated with TiO2 has been constructed and developed for selective enrichment of endogenous peptides taking advantage of the stronger interaction of TiO2 to carboxyl groups of the peptides and the size-exclusion effect of the ordered mesopores under mild enriching conditions. Application effectiveness of the porous structured affinity probe with large specific surface area (216.4 m2 g−1), narrow pore size distribution (3.0 nm) and high saturation magnetization (49.8 emu g−1) was investigated using standard peptides ([Pyr1]-Apelin-13), digest of bovine serum albumin (BSA) proteins, human urine and serum. The signals of peptides can be amplified more than 100 times after enrichment using the affinity probe, and 44 target peptides with sequence coverage of 59% can be captured and identified from 5 nM BSA digest due to plentiful affinity sites of the affinity probe. Furthermore, the affinity probe can selectively capture peptides from the mixture of peptides and proteins with the molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 in virtue of the size-exclusion effect of ordered nanopores. More importantly, the affinity probe is also effective for selective capture of peptides from human urine and serum with high sensitivity. It is expected that this work would be beneficial to rapid capture of potential peptide biomarkers from complex biological samples.
1000 in virtue of the size-exclusion effect of ordered nanopores. More importantly, the affinity probe is also effective for selective capture of peptides from human urine and serum with high sensitivity. It is expected that this work would be beneficial to rapid capture of potential peptide biomarkers from complex biological samples.
Introduction
Early diagnosis is of great importance for disease control and prevention. Disease biomarkers are the molecular signature of a disease in a specific physiological state,1,2 which could provide valuable information for detection of many diseases and elucidation of pathological variations. Research shows that many low-molecule weight endogenous peptides in the body fluids or tissues could be biomarkers with higher clinical sensitivity and specificity.3–5 Therefore, detection of the disease biomarkers in the body fluids or tissues is extremely important for early diagnosis and control of the diseases.6–8 Many efforts in proteomics have been made to solve this problem in recent years. However, how to directly detect these endogenous peptides in complex biological samples is still a great challenge for analysis of mass spectrometry (MS).9,10 There are two major reasons for this. First, MS detection of the extremely low concentration of disease-relevant biomarkers that enter circulation at an early stage of a disease may be masked by other high-abundant proteins (e.g. albumin and immunoglobulins).11,12 Second, the high levels of small molecules such as salts, surfactants and other contaminants in samples often interfere with the MS signals,13,14 resulting in low MS intensity and signal to noise (S/N) ratios of the detected biomarkers. Thus, it is critical to purify and separate these peptide biomarkers prior to MS analysis.Although solvent precipitation is easy to operate and remove most of high-abundant proteins,15 the target peptides need to be further diluted and could suffer from the loss during the removal of proteins. Centrifugal ultrafiltration is a useful technology with an accurate molecular weight cutoff for the separation of proteins and peptides.16,17 However, it is still embarrassed by the concentration of small molecules and salts, thereby leading to poor mass spectrum. Recently, some affinity materials such as immunoaffinity beads,18,19 modified hydrogels20,21 and porous silica22,23 have been developed for selective capture of low molecular weight peptide biomarkers. Especially, the mesoporous silica has attracted particular attentions in enrichment of endogenous peptides due to its remarkable stability (e.g. to be able to effectively prevent Fe3O4 cores from corrosion of the surrounding media),24,25 excellent water dispersibility and unique selectivity for extraction of peptides rather than proteins by the size-exclusion effect of the mesopores.26 What is more, mesoporous silica-coated magnetic microspheres have also been explored to address this deficiency of centrifugal separation process which is a time-consuming and inconvenient procedure.27–29 Nevertheless, owing to the inherently insufficient affinity interactions between silica and target peptides, some peptides are not enriched. Although functionalization of pore walls with organic groups has been developed to address this issue,30,31 this may lead to blockage of the pores and reduction of target biomarker loading.
Up to now, various nanostructures32–34 were explored for selective capture of some specific peptides, such as cysteine-containing peptides,35 phosphopeptides28,36 and glycopeptides.37 Especially, the magnetic nanostructures are of great interest for application in separation of target biomolecules due to their unique magnetic response, high separation efficiency, and convenient operation.38–40 For example, various titania-coated magnetic microspheres were used for enrichment of phosphopeptides by taking advantage of the particular affinity of TiO2 to phosphate groups under the optimized experimental conditions (e.g. trifluoroacetic acid for low pH (2–3) control to prevent non-specific binding of non-phosphorylated peptides by protonation of their carboxyl groups and acetonitrile at levels of 50–80% to prevent hydrophobic interactions with the sorbent).41–44 However, considerable non-specific binding still cannot be avoided.45 This implies that the TiO2 also has strong interaction to non-phosphopeptides via chelating the carboxyl groups of the target peptides, although it is a demerit for the selectivity of phosphopeptide enrichment. However, few of reports focused on developing the titania-based functional materials for enrichment of multiple peptides rather than phosphopeptides. Herein, magnetic mesoporous silica microspheres incorporated with TiO2 (denoted as MMS) have been developed as an affinity probe for selective and direct capture of peptides from a complex biosample under mild enriching conditions. The MMS affinity probe can selectively capture the endogenous peptides rather than the proteins from the complex biological samples by taking advantage of the better interaction of the MMS affinity probe and the size-exclusion effect of its hierarchical mesopores, as illustrated in Scheme 1.
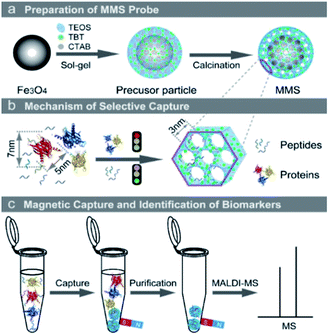 | ||
| Scheme 1 (a) Synthesis of the MMS; (b) mechanism of selective capture of the peptides; and (c) selective capture, rapid magnetic isolation and MS identification of the target peptides using the MMS. | ||
Meanwhile, the high specific surface area of the porous structure and the specific affinity of the metal oxide can further increase capture efficiency and MS detection sensitivity of target peptides. Compared with the conventional filtration, the centrifugation and the gravitational separation, the magnetic bioseparation technique with high efficiency and low cost can realize fast, simple, and effective isolation of the biomarkers. It can be expected that this work could provide new insights for the design of affinity probe for extracting and separating peptide biomarkers from complex biosamples in biomedical application.
Experimental
Chemicals and materials
Ferric chloride hexahydrate (FeCl3·6H2O), Ethylene glycol (EG), Sodium acetate (NaOAc) and Ammonium hydroxide (NH3·H2O) were obtained from Beijing Chemical Regent Co. Ltd. (Beijing, China). Tetrabutyl titanate (TBT), hexamethylenetetramine (HMT), ammonium bicarbonate (NH4HCO3), acetonitrile (ACN), tetraethoxysilane (TEOS), cetyltrimethylammonium bromide (CTAB), trifluoroacetic acid (TFA), 2,5-dihydroxybenzoicacid (2,5-DHB) were purchased from Aladdin (Shanghai, China). The standard peptides [Pyr1]-Apelin-13 (MW = 1533.8; sequence: Pyr-RPRLSHKGPMPF-OH) were obtained from Sangon Biotech (St. Shanghai, China). Bovine serum albumin (BSA) and trypsin (from bovine pancreas, TPCK treated) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Human urine and serum samples were obtained from a healthy volunteer. All the chemical agents were used without further purification.Preparation of Fe3O4 microspheres
The magnetic particles were prepared via a reported solvothermal method with modification.46 Briefly, 10.25 g of FeCl3·6H2O and 10.00 g of NaOAc were dissolved in 200 mL of EG under vigorous stirring, followed by the addition of 2.5 g PEG (MW = 2000) and sonication for 1 h to form a homogeneous yellow solution. 90 mL of the solution was transferred into a Teflon-lined stainless-steel autoclave (120 mL). The autoclave was sealed and heated at 200 °C for 12 h, and then naturally cooled to room temperature. The resulting black Fe3O4 particles were collected with the help of a permanent magnet, followed by washing several times with ethanol and deionized water, and finally dried at 60 °C overnight.Preparation of MMS microspheres
MMS microspheres were prepared via a Stöber sol–gel process. Typically, 100 mg as-prepared Fe3O4 microspheres and 600 mg CTAB were ultrasonically dispersed in a solution containing 80 mL ethanol, 160 mL water and 2.4 mL concentrated ammonia (28 wt%). Then, 0.428 mL TEOS and 0.033 TBT were dissolved into 4 mL EtOH with ultrasonic vibration, and the obtained solution was added into the above homogenous particle dispersion under ultrasonic vibration. Subsequently, the dispersion was mechanically stirred for 2 h at room temperature and then reaction 6 h at 70 °C. Finally, the resulting particles were separated using a permanent magnet and washed with deionized water and ethanol for several times, followed by drying overnight at 60 °C. The CTAB embedded in the silica layer was removed to form mesoporous shell by calcination at 550 °C for 5 h.Preparation of tryptic digests of proteins
BSA was dissolved in 50 mM NH4HCO3 solution, and then trypsin was added into the solution with a molar ratio of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (protein to trypsin) at 37 °C for 16 h. Finally, the obtained tryptic digests were diluted to the target concentrations for selective capture experiments of peptides and detection analysis of MALDI-TOF MS.
1 (protein to trypsin) at 37 °C for 16 h. Finally, the obtained tryptic digests were diluted to the target concentrations for selective capture experiments of peptides and detection analysis of MALDI-TOF MS.
Selective capture of peptides from model samples
To investigate validity of the MMS affinity probe for enrichment of trace peptides, the following experiments were carried out. For a simple peptide sample: 500 μL (5 nM) of standard peptide ([Pyr1]-Apelin-13) solution was mixed with 5 μL of 10 mg mL−1 well-dispersed MMS microsphere suspension and then shaken for 5 min. Subsequently, the particles trapped target peptides were collected and isolated from the mixture with the help of a permanent magnet. After that, the trapped peptides were eluted with 5 μL of 0.5% TFA and 80% ACN solution, and the supernatant for MS detection was collected after magnetic separation. For a complex peptide sample: 500 μL of BSA digestion (5 nM) was mixed with 5 μL of 10 mg mL−1 well-dispersed MMS microsphere suspension and then shaken for 5 min. Subsequently, the particles trapped target peptides were collected and isolated from the mixture with the help of a permanent magnet. After that, the enriched peptides were eluted with 5 μL of 0.5% TFA and 80% ACN solution, and the supernatant for MS detection was collected after magnetic separation. For comparison, a commercial ZipTipC18 pipette tip and MCM-41 silica microspheres were also used to capture target peptides from the above model samples using the similar procedure except substituting centrifugal treatment for magnetic separation.To evaluate selectivity of the MMS affinity probe for enrichment of trace peptides taking advantage of the strong interaction of TiO2 to carboxyl groups of the peptides and the size-exclusion effect of the ordered mesopores under mild enriching conditions, 500 μL biomolecule solution containing the above standard peptide (5 nM) and BSA protein (5 μM) was mixed with 5 μL of 10 mg mL−1 well-dispersed MMS microsphere suspension and then shaken for 5 min. Subsequently, the particles captured target peptides were isolated from the mixture with the help of a permanent magnet, and the particles were rinsed with water two times before collected for MS detection.
Selective capture of peptides from real biological samples
For enriching peptides from the human urine, 1000 μL urine was mixed with 10 μL of 10 mg mL−1 well-dispersed MMS suspension and then shaken for 5 min. Subsequently, the particles trapped target peptides were collected by magnetic separation and washed with water for two times. Then, the MMS affinity particles captured peptides were eluted with 5 μL of 0.5% TFA and 80% ACN solution, and the supernatant for MS detection was collected after magnetic separation.For enriching peptides from the human serum, 10 μL of pristine serum was diluted with 200 μL of water to form a serum solution. 10 μL of 10 mg mL−1 well-dispersed MMS suspension was added into the serum solution, and then shaken for 5 min. Subsequently, the particles trapped target peptides were collected by magnetic separation and washed with water for two times. The composites enriched target peptides were collected by magnetic separation. The same procedure as the above one was employed for preparation of the sample for MS detection.
MALDI-TOF MS analysis and search parameters
Peptides or proteins obtained from the above experiments were mixed with 2 μL of 20 mg mL−1 DHB (in 50% ACN containing 1% H3PO4 aqueous solution, v/v) by pipetting, respectively. Then, 0.7 μL of the mixture was deposited onto the MALDI target. MALDI-TOF MS experiments were performed in positive ion mode with 400 laser shots accumulation on an AB SCIEX MALDI-TOF/TOF 5800 mass spectrometer (Foster City, CA, USA).Mass spectra were submitted to MASCOT (http://www.matrixscience.com/) for database search and identification of corresponding peptides employing the following search parameter settings. Database: SwissProt; enzyme: trypsin; maximum of missed cleavages: 1; taxonomy: mammalia (mammals); peptide tolerance: 0.05 Da; mass values: monoisotopic.
Measurements and characterizations
Scanning electron microscopy (SEM) images were obtained on a field emission scanning electron microscope (FESEM, S4800, Hitachi) equipped with an energy-dispersive X-ray analysis system (EDXA, JEOLJXA-840). Transmission electron microscopy (TEM) and high-resolution TEM (HRTEM) images were taken with a FEI Tecnai G2 S-Twin transmission electron microscope operated at 200 kV. Powder X-ray diffraction (XRD) data were collected on a Bruker D8FOCUS X-ray diffractometer using Cu Kα radiation (λ = 1.5406 Å) at a scanning rate of 15° min−1 and the detective range from 10 to 80 degree. Fourier transform infrared spectra were determined on a BRUKER Vertex 70 FTIR spectrometer over a potassium bromide pellet and then scanned from 400 to 4000 cm−1 at a resolution of 2 cm−1. Nitrogen adsorption isotherms were measured at a liquid nitrogen temperature (77 K) with a Micromeritcs ASAP 2010M apparatus. The specific surface area was determined by the Brunauer–Emmett–Teller (BET) method. The total pore volume was evaluated by the t-plot method, and pore size distribution was analyzed with the supplied BJH software package from the adsorption branches of the isotherms. Magnetization measurement was carried out with a superconducting quantum interface device (SQUID) magnetometer (Quantum Design MPMS XL) at 300 K.Results and discussion
Synthesis and characterization of the MMS microspheres
As illustrated in Scheme 1a, the core–shell MMS microspheres were synthesized by a modified one-pot Stöber sol–gel method using TEOS and TBT as the precursors and CTAB molecules as template agent of mesopores. After removal of CTAB templates by calcination process, the magnetic mesoporous silica microspheres incorporated with TiO2 were obtained. It should be noted that the mesopores in the affinity shell allow the small size of peptides to easily enter the nanopore channels while prevent the large proteins to enter ones in virtue of their size-exclusion effect, and the porous structure with high specific surface area would contribute to the enrichment efficiency. More importantly, the modified silica layer would provide additional TiO2 active sites for peptide affinity in comparison with the pure silica. In addition, the magnetic microspheres can be easily and rapidly separated and collected by an external magnetic field, which would facilitate the procedure of purification.The SEM images (Fig. S1a and b†) indicate that the precursor Fe3O4 microspheres with coarse surface consist of numerous aggregative small Fe3O4 nanoparticles and they have an average diameter of about 200 nm. After the Fe3O4 microspheres were coated with silica shell incorporated with TiO2, the obtained MMS microspheres become slightly larger in diameter and show smooth surface (Fig. 1a and b), indicating the formation of the uniform affinity shell. The TEM images (Fig. 1c) further demonstrate that they are core–shell structure composed of dark magnetic core and grey affinity shell, while many mesopores can be apparently observed in the affinity shell of about 20 nm thickness through the high-resolution TEM images (Fig. 1d). It should be pointed out that the shallow nanopores in the thin affinity layer would not only provide high specific surface area and realize the size-exclusion effect for selective capture of peptide biomarkers, but also facilitate entering and releasing of the target peptides, thereby avoiding the “shadow effect”47 which always appeared in the porous structures with long and small channels.
The elemental composition, crystal structure and surface property of the MMS microspheres can be verified by energy-dispersive X-ray spectrum (EDX), powder X-ray diffraction (XRD), and Fourier-transform infrared (FTIR) spectroscopic analysis, as shown in Fig. 2. Compared with EDX spectrum of the Fe3O4 particles, new elements of Si and Ti except the common Fe and O elements appear in the EDX spectrum of the MMS microspheres (Fig. 2a), indicating formation of the silica shell incorporated with TiO2. The XRD pattern of the Fe3O4 microspheres in Fig. 2b indicates that all diffraction peaks in the XRD pattern of Fe3O4 microspheres are consistent with the cubic Fe3O4 structure (JCPDS card no. 75-0449). By comparison, the XRD pattern of the MMS microspheres (Fig. 2b) has almost same diffraction peaks as those of the Fe3O4 microspheres does except a broad band in the range of 20–30°, indicating that the diffraction peaks root in magnetic cores and the affinity shell is amorphous in nature. Nevertheless, it should be pointed out that the Fe3O4 phase has been entirely or partially transformed into γ-Fe2O3 phase after calcination at 550 °C for 2 h in air under protection of the affinity shell coating and it is difficult to distinguish between Fe3O4 phase and γ-Fe2O3 phase by means of XRD due to their very similar structures. However, the MMS microspheres still have enough magnetism to realize magnetic isolation, which is our focus of attention rather than its crystal structure. The FTIR spectra of the Fe3O4 and the MMS microspheres in Fig. 2c indicate that the characteristic absorption peaks at 587 and 636 cm−1 in the two microspheres are due to stretching vibration of Fe–O from the iron oxide, while the characteristic absorption peaks at around 1074 cm−1 and 951 cm−1 in the FTIR spectrum of the MMS microspheres belong to stretching vibrations of Si–O–Si and Ti–O–Si, respectively.48,49 Above results further confirm the formation of the MMS microspheres.
 | ||
| Fig. 2 EDX spectra (a), XRD patterns (b), and FTIR spectra (c) of the Fe3O4 and the MMS affinity microspheres. | ||
The porous structure with high specific surface areas and appropriate pore sizes plays important roles in selectivity and efficiency of affinity probes for capture of peptides. Therefore, the porous structure of the MMS microspheres was characterized using nitrogen adsorption–desorption isotherm. As shown in Fig. 3, N2 adsorption–desorption isotherm of the MMS microspheres displays a type-IV curve with small hysteresis according to the IUPAC classification,50 indicating mesoporous structure of the MMS microspheres. The average Brunauer–Emmett–Teller (BET) specific surface area, the total pore volume, and the average pore diameter of the microspheres are calculated to be 216.4 m2 g−1, 0.164 cm3 g−1 and 3.0 nm, respectively. The pore-size distribution derived from the adsorption branch using the Barrett–Joyner–Halenda (BJH) method (Fig. 3, inset) shows a narrow peak in the mesoporous range, showing a hierarchical porous structure. It should be noted that the mesopores with narrow distribution would contribute to selective capture of the peptides while cut off the high-abundant proteins with larger MW (molecular weight).
 | ||
| Fig. 3 N2 adsorption–desorption isotherm and pore size distribution (inset) of the MMS affinity microspheres. | ||
The magnetic properties of the Fe3O4 particles and the MMS affinity microspheres were examined at 300 K via a superconducting quantum interface device (SQUID) magnetometer. As revealed in the hysteresis loops (Fig. 4b), the microspheres show a strong magnetism at room temperature. The saturation magnetization (Ms) values of Fe3O4 particles and MMS microspheres are 72.4 and 49.8 emu g−1, respectively. The Ms decrease of the MMS microspheres may be attributed to the introduction of nonmagnetic affinity shell on the surface of the magnetic Fe3O4 particles. However, it still has enough strong magnetism for the practical magnetic separation.
 | ||
| Fig. 4 The magnetic hysteresis loops of the Fe3O4 and the MMS affinity microspheres, and dispersion and separation process of the MMS microspheres (inset). | ||
As shown in Fig. 4b inset, the MMS microspheres can be separated from the mixture within 30 s by applying a permanent magnet, while they can be easily redispersed by shaking after removing the magnet. The excellent magnetic responsivity and redispersibility of the MMS affinity microspheres are beneficial to simplify the operation process of the purification and be free of the laborious centrifugation in bioseparation application.
To prove validity of the MMS for enrichment of trace peptides, a standard peptide ([Pyr1]-Apelin-13, MW = 1533.8; sequence: Pyr-RPRLSHKGPMPF-OH) (5 nM) and a diluted BSA tryptic digest (5 nM) were employed as a simple/complex peptide sample, respectively. Fig. 5 gives the MALDI-TOF mass spectra of the [Pyr1]-Apelin-13 (5 nM) without any treatment, and after enrichment with the MMS, the ZipTipC18 pipette tip and the normal mesoporous silica microspheres, respectively.
For direct detection, no effective peptide signal can be observed (Fig. 5a) due to the extremely low concentration of the target peptide; while after enrichment using the MMS, the peptide (MW = 1533.8) with high intensity and signal to noise (S/N) ratio and its methionine-oxidized peptide (MW = 1549.8) can be detected (Fig. 5b). For comparison, the commercial affinity product (a ZipTipC18 pipette tip) and normal mesoporous silica microspheres were also used to enrich the standard peptides. Although the ZipTipC18 can effectively capture the standard peptide, the MS intensities and S/N ratio of the peptides are still very low (Fig. 5c). In contrast, the normal mesoporous silica microspheres for the peptide enrichment have higher MS intensity and S/N ratio than that of the commercial ZipTipC18, but they all are much lower than that of the MMS affinity probe.
Fig. 6 gives the MALDI-TOF mass spectra of the diluted BSA tryptic digest (5 nM) without any treatment, and after enrichment with the MMS, the ZipTipC18 pipette tip and the normal mesoporous silica microspheres, respectively. Only five peptides with low intensity and S/N ratio can be detected through direct MS detection (Fig. 6a) due to the low concentration of the BSA digest and interference of the salts used in digestion. However, after enrichment using the MMS affinity probe, 44 peptides with enhanced MS intensity, high S/N ratio and a sequence coverage of 59% can be detected (Fig. 6b), which is much better than that in previous reports.29 Table S1† lists the detail informations of captured peptides from database search. Contrastively, less matched peptides (number: 10, sequence coverage: 16%, see Table S1†) with lower intensity and S/N ratio were observed (Fig. 6c) after the solution of diluted BSA digest was treated by a ZipTipC18 pipette tip. For the enrichment using normal mesoporous silica microspheres, 21 peptides from BSA tryptic digest with sequence coverage of 28% can be detected (see Table S1†). The above results indicate that the MMS affinity probe can effectively enrich the target peptides.
To demonstrate selectivity of the MMS for enrichment of trace peptides taking advantage of the size-exclusion effect in virtue of the mesopore size, a mixture of a standard peptide ([Pyr1]-Apelin-13, MW = 1533.8 Da) and a protein (BSA, MW = 66 KDa) with a molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 was chosen to verify the cutoff mechanism. The BSA molecules with size about 5–7 nm (ref. 51) would be excluded from the mesopores with the size of 3.0 nm because the size of BSA molecule is much larger than the pore size MMS affinity probe (Scheme 1b). Fig. 7a and b show the direct MS detection of the mixture of peptides and proteins. The target peptides cannot be detected owing to the low concentration and interference of large amounts of proteins (Fig. 7a), while the BSA proteins can be apparently detected with strong intensity and S/N ratio (Fig. 7b). After affinity capture of the target peptides using the MMS probe and magnetic separation, the supernatant was subjected to MS analysis. As shown in Fig. 7c and d, similar results can be observed, indicating the proteins were not captured by the MMS affinity probe. However, after captured by the MMS, the standard peptide with high intensity and S/N ratio can be detected (Fig. 7e). Interestingly, besides the standard peptides, many new peptides with lower intensities can also be observed, which may be attributed to the trace degradation products of BSA proteins in storage. Contrastively, no BSA proteins were detected (Fig. 7f). Therefore, it can be concluded that the MMS affinity probe selectively and effectively captures the peptides but excludes the large proteins by the size-exclusion effect of the unique pore structure.
1000 was chosen to verify the cutoff mechanism. The BSA molecules with size about 5–7 nm (ref. 51) would be excluded from the mesopores with the size of 3.0 nm because the size of BSA molecule is much larger than the pore size MMS affinity probe (Scheme 1b). Fig. 7a and b show the direct MS detection of the mixture of peptides and proteins. The target peptides cannot be detected owing to the low concentration and interference of large amounts of proteins (Fig. 7a), while the BSA proteins can be apparently detected with strong intensity and S/N ratio (Fig. 7b). After affinity capture of the target peptides using the MMS probe and magnetic separation, the supernatant was subjected to MS analysis. As shown in Fig. 7c and d, similar results can be observed, indicating the proteins were not captured by the MMS affinity probe. However, after captured by the MMS, the standard peptide with high intensity and S/N ratio can be detected (Fig. 7e). Interestingly, besides the standard peptides, many new peptides with lower intensities can also be observed, which may be attributed to the trace degradation products of BSA proteins in storage. Contrastively, no BSA proteins were detected (Fig. 7f). Therefore, it can be concluded that the MMS affinity probe selectively and effectively captures the peptides but excludes the large proteins by the size-exclusion effect of the unique pore structure.
To further evaluate the effectiveness of the MMS affinity probe for capture of peptides from real biological samples, human urine and human serum were employed as real biological samples. Human urine is an ideal candidate for use in clinical diagnostics and it contains some endogenous peptides correlating well with the pathophysiology, which could serve as biomarkers to diagnose, monitor and manage a number of diseases.52,53 For direct MS detection of human urine, as shown in Fig. 8a, no effective MS signals can be identified because the peptides in human urine are low-concentration and the urine always contains high levels of salts and contaminants, which would severely interfere with the detection. However, after treatment of the urine using the MMS affinity probe, many peptides can be evidently detected with high intensity and S/N ratio (Fig. 8b).
Human serum, as another important body fluid, contains an archive of information due to the presence of various endogenous peptides released by diseased tissue, and serum proteomics has gained considerable interest for the disease biomarker discovery. Unfortunately, it is difficult to directly detect MS signals of the peptide biomarkers due to their extremely low concentration and strong interference from high-abundance serum proteins, high levels of salts and other interferential substances in human serum.54Fig. 8c gives the mass spectrum of the direct detection of human serum using MALDI-TOF MS. As expected, almost no effective peptides can be detected because of the above-mentioned reasons. We use the MMS affinity probe to extract the peptides from the complex serum sample by taking advantage of the larger specific surface area, unique metal oxide affinity and size-exclusion effect. As shown in Fig. 8d, numerous peptides were apparently detected with high intensity and S/N ratio after treatment by the MMS affinity probe. These results demonstrate that the MMS probe can be used for selective capture, efficient enrichment and rapid separation of peptides from complex biological samples.
Conclusions
Novel magnetic microspheres with mesoporous silica shell incorporated with TiO2 were prepared via a simple sol–gel method, and as the MMS affinity probe they are applied for rapid capture and identification of target peptides. By taking advantage of the size-exlusion effect of ordered pores, the MMS affinity probe can extract selectively the small size peptides, while excluding the larger size proteins. The porous structure with high specific surface area and additional TiO2 affinity sites can promote efficiency of enrichment of target peptides, which is much superior to those of the commercial products (ZipTipC18 pipette tip) and normal mesoporous silica microspheres under the present experimental conditions. The magnetic core would contribute to the fast magnetic separation and reduce the sample loss, while the thin affinity shell with shallow pores would avoid the “shadow effect”, contributing to fast adsorption and release of target peptides. Therefore, this work may provide a powerful tool for capture and discovery of peptide biomarkers in biofluids, which is high potential in the early diagnosis and disease management in clinical utility.Acknowledgements
This work was supported by the National Natural Science Foundation of China (NNSFC) (Grant no. 21171161).Notes and references
- A. Dove, Science, 2010, 329, 1373 CrossRef.
- N. Kosaka, H. Iguchi and T. Ochiya, Cancer Sci., 2010, 101, 2087–2092 CrossRef PubMed.
- A. Doerr, Nat. Methods, 2008, 5, 858–859 CrossRef.
- M. Swami, Nat. Rev. Cancer, 2010, 10, 597 CrossRef.
- R. Terracciano, F. Casadonte, L. Pasqua, P. Candeloro, E. D. Fabrizio, A. Urbani and R. Savino, Talanta, 2010, 80, 1532–1538 CrossRef PubMed.
- J. Fan, Y. Huang, I. Finoulst, H.-J. Wu, Z. Deng, R. Xu, X. Xia, M. Ferrari, H. Shen and Y. Hu, Cancer Lett., 2012, 334, 202–210 CrossRef PubMed.
- S. M. Hanash, S. J. Pitteri and V. M. Faca, Nature, 2008, 452, 571–579 CrossRef PubMed.
- Y. Hu, A. Bouamrani, E. Tasciotti, L. Li, X. Liu and M. Ferrari, ACS Nano, 2009, 4, 439–451 CrossRef PubMed.
- B. F. Cravatt, G. M. Simon and J. R. Y. Iii, Nature, 2007, 450, 991–1000 CrossRef PubMed.
- A. K. Callesen, J. S. Madsen, W. Vach, T. A. Kruse, O. Mogensen and O. N. Jensen, Proteomics, 2009, 9, 1428–1441 CrossRef PubMed.
- W.-J. Qian, D. T. Kaleta, B. O. Petritis, H. Jiang, T. Liu, X. Zhang, H. M. Mottaz, S. M. Varnum, D. G. Camp, L. Huang, X. Fang, W.-W. Zhang and R. D. Smith, Mol. Cell. Proteomics, 2008, 7, 1963–1973 Search PubMed.
- D. M. Good, V. Thongboonkerd, J. Novak, J.-L. Bascands, J. P. Schanstra, J. J. Coon, A. Dominiczak and H. Mischak, J. Proteome Res., 2007, 6, 4549–4555 CrossRef PubMed.
- H. Qin, P. Gao, F. Wang, L. Zhao, J. Zhu, A. Wang, T. Zhang, R. A. Wu and H. Zou, Angew. Chem., Int. Ed., 2011, 50, 12218–12221 CrossRef PubMed.
- M. Gao, D. Qi, P. Zhang, C. Deng and X. Zhang, Expert Rev. Proteomics, 2010, 7, 665–678 CrossRef PubMed.
- Y. Xu, J. T. Mehl, R. Bakhtiar and E. J. Woolf, Anal. Chem., 2010, 82, 6877–6886 CrossRef PubMed.
- L. Firdaous, P. Dhulster, J. Amiot, A. Gaudreau, D. Lecouturier, R. Kapel, F. Lutin, L.-P. Vézina and L. Bazinet, J. Membr. Sci., 2009, 329, 60–67 CrossRef PubMed.
- L. Civiero, R. Vancraenenbroeck, E. Belluzzi, A. Beilina, E. Lobbestael, L. Reyniers, F. Gao, I. Micetic, M. D. Maeyer, L. Bubacco, V. Baekelandt, M. R. Cookson, E. Greggio and J.-M. Taymans, PLoS One, 2012, 7, e43472 Search PubMed.
- B. L. Ackermann and M. J. Berna, Expert Rev. Proteomics, 2007, 4, 175–186 CrossRef PubMed.
- J. R. Whiteaker, L. Zhao, L. Anderson and A. G. Paulovich, Mol. Cell. Proteomics, 2010, 9, 184–196 Search PubMed.
- A. Luchini, D. H. Geho, B. Bishop, D. Tran, C. Xia, R. L. Dufour, C. D. Jones, V. Espina, A. Patanarut, W. Zhou, M. M. Ross, A. Tessitore, E. F. Petricoin and L. A. Liotta, Nano Lett., 2007, 8, 350–361 CrossRef PubMed.
- C. Longo, A. Patanarut, T. George, B. Bishop, W. Zhou, C. Fredolini, M. M. Ross, V. Espina, G. Pellacani, E. F. Petricoin, III, L. A. Liotta and A. Luchini, PLoS One, 2009, 4, e4763 Search PubMed.
- M. F. Lopez, A. Mikulskis, S. Kuzdzal, E. Golenko, E. F. Petricoin, L. A. Liotta, W. F. Patton, G. R. Whiteley, K. Rosenblatt, P. Gurnani, A. Nandi, S. Neill, S. Cullen, M. O'Gorman, D. Sarracino, C. Lynch, A. Johnson, W. Mckenzie and D. Fishman, Clin. Chem., 2007, 53, 1067–1074 Search PubMed.
- R. Terracciano, M. Preianò, G. P. Palladino, G. E. Carpagnano, M. P. F. Barbaro, G. Pelaia, R. Savino and R. Maselli, Proteomics, 2011, 11, 3402–3414 CrossRef PubMed.
- J. Zhu, S. Wei, I. Y. Lee, S. Park, J. Willis, N. Haldolaarachchige, D. P. Young, Z. Luo and Z. Guo, RSC Adv., 2012, 2, 1136–1143 RSC.
- J. Zhu, S. Wei, N. Haldolaarachchige, D. P. Young and Z. Guo, J. Phys. Chem. C, 2011, 115, 15304–15310 Search PubMed.
- R. Tian, H. Zhang, M. Ye, X. Jiang, L. Hu, X. Li, X. Bao and H. Zou, Angew. Chem., Int. Ed. Engl., 2007, 46, 962–965 CrossRef PubMed.
- S. Liu, Y. Li, C. Deng, Y. Mao, X. Zhang and P. Yang, Proteomics, 2011, 11, 4503–4513 CrossRef PubMed.
- G. Cheng, Y. L. Liu, J. L. Zhang, D. H. Sun and J. Z. Ni, Anal. Bioanal. Chem., 2012, 404, 763–770 CrossRef PubMed.
- S. Liu, H. Chen, X. Lu, C. Deng, X. Zhang and P. Yang, Angew. Chem., Int. Ed. Engl., 2010, 49, 7557–7561 CrossRef PubMed.
- K. Böddi, A. Takátsy, S. Szabó, L. Markó, L. Márk, I. Wittmann, R. Ohmacht, G. Montskó, R. M. Vallant, T. Ringer, R. Bakry, C. W. Huck, G. K. Bonn and Z. Szabó, J. Sep. Sci., 2009, 32, 295–308 CrossRef PubMed.
- L. Zhang, S. Wu, C. Li and Y. Qihua, Chem. Commun., 2012, 48, 4190–4192 RSC.
- M. Atakay, Ö. Çelikbıçak and B. Salih, Anal. Chem., 2012, 84, 2713–2720 CrossRef PubMed.
- G. Cheng, Z. G. Wang, Y. L. Liu, J. L. Zhang, D. H. Sun and J. Z. Ni, Chem. Commun., 2012, 48, 10240–10242 RSC.
- F. Wang, Z. Shi, F. Hu, Z. Xia and L. Wang, Anal. Bioanal. Chem., 2012, 405, 1683–1693 CrossRef PubMed.
- J. Dai, J. Wang, Y. Zhang, Z. Lu, B. Yang, X. Li, Y. Cai and X. Qian, Anal. Chem., 2005, 77, 7594–7604 CrossRef PubMed.
- Z.-G. Wang, G. Cheng, Y.-L. Liu, J.-L. Zhang, D.-H. Sun and J.-Z. Ni, Small, 2012, 8, 3456–3464 CrossRef PubMed.
- Y. Qu, J. Liu, K. Yang, Z. Liang, L. Zhang and Y. Zhang, Chem. – Eur. J., 2012, 18, 9056–9062 CrossRef CAS PubMed.
- D. Qi, Y. Mao, J. Lu, C. Deng and X. Zhang, J. Chromatogr. A, 2010, 1217, 2606–2617 CrossRef CAS PubMed.
- H.-Y. Lin, W.-Y. Chen and Y.-C. Chen, Anal. Bioanal. Chem., 2009, 394, 2129–2136 CrossRef CAS PubMed.
- G. Cheng, Y.-L. Liu, Z.-G. Wang, J.-L. Zhang, D.-H. Sun and J.-Z. Ni, J. Mater. Chem., 2012, 22, 21998–22004 RSC.
- J. H. Wu, X. S. Li, Y. Zhao, Q. Gao, L. Guo and Y. Q. Feng, Chem. Commun., 2010, 46, 9031–9033 RSC.
- C. T. Chen and Y. C. Chen, Anal. Chem., 2005, 77, 5912–5919 CrossRef CAS PubMed.
- L. Hu, H. Zhou, Y. Li, S. Sun, L. Guo, M. Ye, X. Tian, J. Gu, S. Yang and H. Zou, Anal. Chem., 2009, 81, 94–104 CrossRef CAS PubMed.
- W. Ma, Y. Zhang, L. Li, L. You, P. Zhang, J. Li, M. Yu, J. Guo, H. J. Lu and C. Wang, ACS Nano, 2012, 6, 3179–3188 CrossRef CAS PubMed.
- A. Leitner, TrAC, Trends Anal. Chem., 2010, 29, 177–185 CrossRef CAS PubMed.
- G. Cheng, J. L. Zhang, Y. L. Liu, D. H. Sun and J. Z. Ni, Chem. Commun., 2011, 47, 5732–5734 RSC.
- B. Xu, L. Zhou, F. Wang, H. Qin, J. Zhu and H. Zou, Chem. Commun., 2012, 48, 1802–1804 RSC.
- J. Jarupatrakorn and T. D. Tilley, J. Am. Chem. Soc., 2002, 124, 8380–8388 CrossRef CAS PubMed.
- V. Lafond, P. H. Mutin and A. Vioux, Chem. Mater., 2004, 16, 5380–5386 CrossRef CAS.
- R. Pierotti and J. Rouquerol, Pure Appl. Chem., 1985, 57, 603–619 Search PubMed.
- S. Hudson, J. Cooney and E. Magner, Angew. Chem., Int. Ed., 2008, 47, 8582–8594 CrossRef CAS PubMed.
- J. Casado-Vela, T. G. del Pulgar, A. Cebrián, N. Álvarez-Ayerza and J. C. Lacal, Expert Rev. Proteomics, 2011, 8, 347–360 CrossRef CAS PubMed.
- Y.-T. Chen, H.-W. Chen, D. Domanski, D. S. Smith, K.-H. Liang, C.-C. Wu, C.-L. Chen, T. Chung, M.-C. Chen, Y.-S. Chang, C. E. Parker, C. H. Borchers and J.-S. Yu, J. Proteomics, 2012, 75, 3529–3545 CrossRef CAS PubMed.
- S. Ray, P. J. Reddy, R. Jain, K. Gollapalli, A. Moiyadi and S. Srivastava, Proteomics, 2011, 11, 2139–2161 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c3ra46887a |
| This journal is © The Royal Society of Chemistry 2014 |





