Selective mitochondrial accumulation of cytotoxic dinuclear polypyridyl ruthenium(II) complexes
Michelle J.
Pisani
a,
Daniel K.
Weber
b,
Kirsten
Heimann
c,
J. Grant
Collins
*a and
F. Richard
Keene
*b
aSchool of Physical, Environmental and Mathematical Sciences, University of New South Wales, Australian Defence Force Academy, Canberra, ACT 2600, Australia
bSchool of Pharmacy and Molecular Sciences, James Cook University, Townsville, QLD 4811, Australia. E-mail: richard.keene@jcu.edu.au; Fax: +61 7 4781 6078
cSchool of Marine and Tropical Biology, James Cook University, Townsville, QLD 4811, Australia
First published on 23rd April 2010
Abstract
The lipophilic ligand-bridged dinuclear cation Rubb16 is significantly cytotoxic and preferentially accumulates in the mitochondria of the L1210 murine leukemia cancer cell line.
It has been more than fifty years since Dwyer and coworkers showed that mononuclear tris(bidentate)ruthenium(II) complexes containing polypyridyl ligands have antibacterial activity.1 Subsequent studies have shown that this class of compounds also has significant anticancer potential,2–6 where enhanced cytotoxicity is correlated with increased lipophilicity. The mechanism of action for these complexes has generally been assumed to involve DNA interaction, although other mechanisms have been suggested.5,6 Over the last decade there has been growing interest in inert dinuclear polypyridyl ruthenium(II) complexes, particularly as nucleic acid binding agents.7,8 To date, dinuclear polypyridyl ruthenium(II) complexes have only exhibited modest levels of cytotoxicity;9,10 however, it is probable that their cytotoxic properties could be significantly improved by increasing the overall lipophilicity as observed for the dinuclear ruthenium(II) arene complexes reported by Mendoza-Ferri et al.11 We have been interested in the DNA binding ability of a series of dinuclear complexes where the metal centres are linked by a flexible bridge8 in which the lipophilicity can be systematically varied by increasing the number of methylene groups in the chain. In the present communication, we demonstrate that increasing the length of the alkane chain does significantly improve the cytotoxic properties. More interestingly, we show for the first time that dinuclear ruthenium(II) complexes can selectively accumulate in the mitochondria of live cancer cells and that for this series of complexes the level of accumulation is reflected in their cytotoxities.
We have studied a series of dinuclear ruthenium(II) complexes, ΔΔ/ΛΛ-[{Ru(phen)2}2{μ-bbn}]4+ {“Rubbn”; where phen = 1,10-phenanthroline; bbn = 1,n-bis[4(4′-methyl-2,2′-bipyridyl)]-nane (n = 2, 5, 7, 10, 12 or 16; Fig. 1}, which only differ in the number of methylene groups in the bridging ligand between the two metal centres. Rubbn (n = 2, 5, 7, 10) were obtained as their chloride salts as previously described,12 while the Rubb12 and Rubb16 analogues were stereoselectively synthesised by the same method, but using 1,12-bis[4(4′-methyl-2,2′-bipyridyl)]dodecane and 1,16-bis[4(4′-methyl-2,2′-bipyridyl)]hexadecane as the linking ligands, respectively. Confirmatory NMR, elemental analysis, circular dichroism and mass spectroscopy data were obtained. The cytotoxicity of this family of ruthenium complexes was analysed in the L1210 murine leukemia cancer cell line. The results are summarised in Table 1. The complex Rubb2 showed very low cytotoxicity, while that of Rubb7 and Rubb10 was relatively modest and similar to that reported for other inert dinuclear ruthenium(II) polypyridyl complexes:9,10 however, Rubb12 and Rubb16 exhibited significant cytotoxicity to the leukemia cell line. Interestingly, the ΔΔ isomers of the more active ruthenium complexes (Rubb12 and Rubb16) were slightly more cytotoxic than their ΛΛ counterparts.
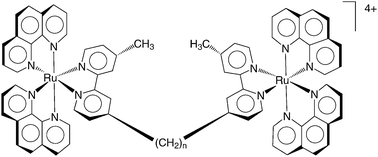 | ||
| Fig. 1 The chemical structure of Rubbn (where n = 2, 5, 7 10, 12 or 16 methylene groups in the linking chain). | ||
Cellular uptake of the Rubbn analogues can be compared using fluorescence microscopy.13 As shown in Fig. 2, all three Rubbn complexes (n = 5, 10, 16) entered the L1210 cells. No obvious decrease in fluorescence intensity was observed, indicating that no significant quenching took place as a consequence of the ruthenium complexes binding to intracellular components or being chemically degraded. Incubation with 50 μM Rubb5 resulted in luminescence inside the cell which was appreciably above the background, with sites of localisation visible. The luminescence intensity inside the cell increased dramatically with incubation of Rubb10 and Rubb16 at 50 μM, resulting in high fluorescence obscuring localisation patterns; however, incubation at 25 μM and 5 μM for Rubb10 and Rubb16, respectively, decreased the luminescence intensity to the point where organelle staining was visible (Fig. 2). Organelle staining for all metal complexes was suggestive of mitochondrial localisation.
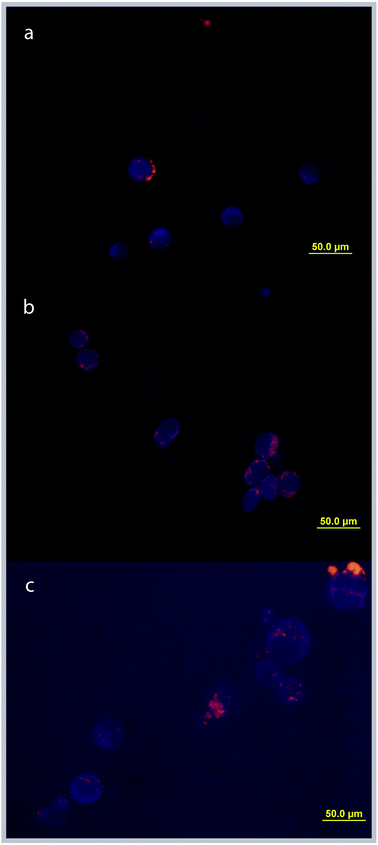 | ||
| Fig. 2 Fluorescence microscopy images of cellular uptake and organelle accumulation of Rubbn. Incubation concentrations are: (a) Rubb5 (50 μM), (b) Rubb10 (25 μM) and (c) Rubb16 (5 μM). | ||
Confocal microscopy has been shown to be a useful tool for studying the cellular localisation of luminescent ruthenium(II) compounds.14 As the results of fluorescence microscopy studies suggested that mitochondria were the cellular sites of uptake for the Rubbn complexes, confocal microscopy was used in conjunction with a mitochondrial tracking dye to identify organelle localisation sites within the cell. Localisation studies were undertaken by adding Mitotracker® Green FM (Invitrogen) to the cell suspension during the last 30 min of incubation to give a final incubation concentration of 100–200 nM. Live cells were washed, mounted and viewed at 60× (oil) magnification on a BIO-RAD Radiance 2000 laser-scanning confocal microscope. Both the metal complex (λex ∼ 450 nm, λem ∼ 610 nm) and Mitotracker (λex ∼ 490 nm, λem ∼ 516 nm) were excited using a blue argon laser (488 nm). After four hours incubation, the site(s) of accumulation of the metal complexes were visible. For Rubb5, the cellular staining indicated that the drug was significantly taken up by mitochondria (Fig. 3a); however other sites of accumulation were also observed. These sites are thought to be lysosomes and further studies are being undertaken to confirm this hypothesis. In contrast, all the Rubb10 and Rubb16 accumulated in the mitochondria, even at high complex concentrations, with no staining being observed in the cytoplasm or other organelles (Fig. 3b), consistent with observations at 5 μM.
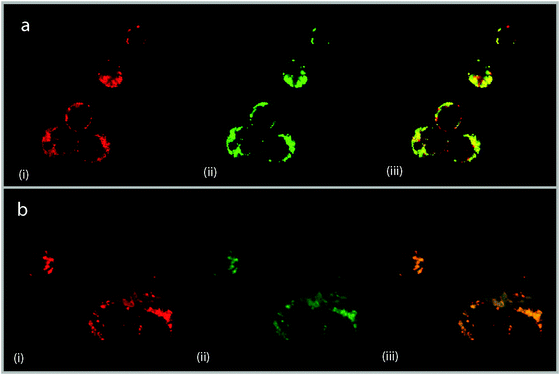 | ||
| Fig. 3 Confocal microscopy images of (a) Rubb5 (50 μM) and (b) Rubb16 (50 μM). Image (i) shows cellular staining of Rubbn, (ii) Mitotracker® Green FM staining, and (iii) a micrograph overlay showing sites of co-localisation. Areas stained yellow are indicative of drug localisation in mitochondria. Mitotracker® Green FM incubation concentrations were 200 nM and 100 nM for (a) and (b), respectively. | ||
Differential interference contrast light microscopy was used to determine cell characteristics in order to establish morphological changes induced by the addition of the metal complexes. The ruthenium complexes did induce several changes to the cell morphology, most notably the appearance of a grainy texture, blebbing of the plasma membrane, and irregular cell size and shape (Fig. 4). Cells incubated with Rubb16 exhibited a greater degree of morphological changes compared to Rubb5 and Rubb10, most significantly a 20% reduction in the average cell size compared to untreated cells (Table 2) and an increase in the incidence of membrane blebbing.
 | ||
| Fig. 4 Differential interference contrast light micrographs of L1210 murine leukemia cells (left) untreated and (right) treated with Rubb16 (5 μM) highlighting cell morphology changes induced by drug uptake; i.e. irregular cell shape and size, increased membrane blebbing and grainy texture. | ||
While Rubb7 and Rubb10 only showed modest cytotoxicity, similar to that reported for other inert dinuclear ruthenium(II) polypyridyl complexes,9,10 Rubb12 and Rubb16 exhibited significant cytotoxicity to the leukemia cell line. It is concluded that the additional methylene groups in Rubb12 and Rubb16 increase the lipophilicity of the metal complex, thereby allowing greater cellular accumulation. DNA binding is often considered as a dominant mechanism of biological activity; indeed, Gill et al.15 reported that a dinuclear ruthenium(II) polypyridyl complex can be used to image DNA in living cells. However other mechanisms, e.g. modification of cell membrane function and cell adhesion, have been proposed.5,6 Furthermore, while Puckett and Barton suggested some mitochondrial uptake, inter alia, for a mononuclear intercalating complex,14 this present study shows that the mitochondria are the predominate site of accumulation for our dinuclear complexes. Further experiments are under way to establish the precise mechanism of the cytotoxicity.
As the ruthenium complexes are localised in the mitochondria it is also concluded that they are acting as delocalised lipophilic cations. Owing to the fact that mitochondria play an important role in cell function—most notably energy generation, apoptosis, cell cycle control and calcium homeostasis16—it represents a key target for anticancer therapeutics. Consequently, lipophilic dinuclear ruthenium(II) complexes may have potential as anticancer agents.
Acknowledgements
This work was funded by the Australian Research Council. The L1210 murine leukemia cells were provided by the Peter McCallum Cancer Center, Melbourne, Australia and we thank Dr Carleen Cullinane for her cooperation and assistance. We thankfully acknowledge the technical assistance of Dr Kevin Blake (Advanced Analytical Centre, James Cook University) with the confocal microscopy experiments. We are also thankful to Professor Alan Baxter (Comparative Genomic Center, James Cook University) for the use of cell culture facilities, and Mr Martin Keene for assistance with the establishment of the cell line.References
- F. P. Dwyer, E. C. Gyarfas, W. P. Rogers and J. H. Koch, Nature, 1952, 170, 190 CrossRef CAS.
- F. P. Dwyer, E. Mayhew, E. M. F. Roe and A. Schulman, British. J. Cancer, 1965, 19, 195 CAS.
- S. Schäfer, I. Ott, R. Gust and W. S. Sheldrick, Eur. J. Inorg. Chem., 2007, 3034 CrossRef.
- O. Novakova, J. Kasparkova, O. Vrana, P. M. Vanvliet, J. Reedijk and V. Brabec, Biochemistry, 1995, 34, 12369 CrossRef CAS.
- U. Schatzschneider, J. Niesel, I. Ott, R. Gust, H. Alborzina and S. Wölfl, ChemMedChem, 2008, 3, 1104 CrossRef CAS.
- O. Zava, S. M. Zakeeruddin, C. Danelon, H. Vogel, M. Grätzel and P. J. Dyson, ChemBioChem, 2009, 10, 1796 CrossRef CAS.
- C. Metcalfe and J. A. Thomas, Chem. Soc. Rev., 2003, 32, 215 RSC.
- F. R. Keene, J. A. Smith and J. G. Collins, Coord. Chem. Rev., 2009, 253, 2021 CrossRef CAS.
- U. McDonnell, J. M. C. A. Kerchoffs, R. P. M. Castineiras, M. R. Hicks, A. C. G. Hotze, M. J. Hannon and A. Rodger, Dalton Trans., 2007, 667 Search PubMed.
- E. Corral, A. C. G. Hotze, H. den Dulk, A. Leczkowska, A. Rodger, M. J. Hannon and J. Reedjik, J. Biol. Inorg. Chem., 2009, 14, 439 CrossRef CAS.
- M.-G. Mendoza-Ferri, C. G. Hartinger, R. E. Eichinger, N. Stolyarova, K. Severin, M. A. Jakupec, A. A. Nazarov and B. K. Keppler, Organometallics, 2008, 27, 2405 CrossRef CAS.
- J. L. Morgan, C. B. Spillane, J. A. Smith, D. P Buck, J. G. Collins and F. R. Keene, Dalton Trans., 2007, 4333 RSC.
- L1210 cells (obtained from the Peter McCallum Cancer Center, Melbourne) were maintained in RPMI-1640 media (supplemented with 10% foetal calf serum (FCS), 1 mM L-glutamine, 1 mM penicillin/streptomycin, 1 mM HEPES buffer). Suspensions of L1210 cells were incubated in a 50 mL tissue culture flask in final concentrations of Rubbn (n = 5, 10, 16) of 50 μM for 4 h at 37 °C with 5% humidified CO2. Live cell suspensions were washed, mounted and viewed at 40× magnification on an Olympus BX51 fluorescence microscope fitted with the UMWIBA2 fluorescence filter cube (excitation filter 400–440 nm, emission filter 475 nm and a dichromatic mirror 455 nm) and the CCD-cooled digital camera Olympus DP70 for documentation.
- C. A. Puckett and J. K. Barton, J. Am. Chem. Soc., 2007, 129, 46 CrossRef CAS.
- M. R. Gill, J. Garcia-Lara, S. J. Foster, C. Smythe, G. Battaglia and J. A. Thomas, Nat. Chem., 2009, 1, 662 Search PubMed.
- A. S. Don and P. Hogg, Trends Mol. Med., 2004, 10, 372 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2010 |
