Development of a method for bacteria and virus recovery from heating, ventilation, and air conditioning (HVAC) filters
James E.
Farnsworth
*ab,
Sagar M.
Goyal
c,
Seung
Won Kim
d,
Thomas H.
Kuehn
b,
Peter C.
Raynor
d,
M. A.
Ramakrishnan
c,
Senthilvelan
Anantharaman
c and
Weihua
Tang
b
aEngineering Department, TSI Incorporated, 500 Cardigan Road, Saint Paul, MN 55126-3903, USA
bEnvironmental Division, Department of Mechanical Engineering, University of Minnesota, 111 Church Street SE, Minneapolis, MN 55455-0111, USA
cDepartment of Veterinary Population Medicine, University of Minnesota, 1333 Gortner Avenue, Saint Paul, MN 55108-1098, USA
dDivision of Environmental Health Sciences, School of Public Health, University of Minnesota, 420 Delaware St SE, Minneapolis, MN 55455-0341, USA
First published on 6th September 2006
Abstract
The aim of the work presented here is to study the effectiveness of building air handling units (AHUs) in serving as high volume sampling devices for airborne bacteria and viruses. An HVAC test facility constructed according to ASHRAE Standard 52.2-1999 was used for the controlled loading of HVAC filter media with aerosolized bacteria and virus. Nonpathogenic Bacillus subtilis var. niger was chosen as a surrogate for Bacillus anthracis. Three animal viruses; transmissible gastroenteritis virus (TGEV), avian pneumovirus (APV), and fowlpox virus were chosen as surrogates for three human viruses; SARS coronavirus, respiratory syncytial virus, and smallpox virus; respectively. These bacteria and viruses were nebulized in separate tests and injected into the test duct of the test facility upstream of a MERV 14 filter. SKC Biosamplers upstream and downstream of the test filter served as reference samplers. The collection efficiency of the filter media was calculated to be 96.5 ± 1.5% for B. subtilis, however no collection efficiency was measured for the viruses as no live virus was ever recovered from the downstream samplers. Filter samples were cut from the test filter and eluted by hand-shaking. An extraction efficiency of 105 ± 19% was calculated for B. subtilis. The viruses were extracted at much lower efficiencies (0.7–20%). Our results indicate that the airborne concentration of spore-forming bacteria in building AHUs may be determined by analyzing the material collected on HVAC filter media, however culture-based analytical techniques are impractical for virus recovery. Molecular-based identification techniques such as PCR could be used.
1. Introduction
Indoor and outdoor air quality concerns have received increased public attention in recent years with a focus on biocontaminants. Bioaerosols such as pollen and fungus have been linked to allergic responses and Sick Building Syndrome. In addition, disease-causing bacteria such as Mycobacterium tuberculosis and Streptococcus pneumoniae and viruses such as smallpox, chickenpox, and influenza viruses are documented to have been transmitted via the airborne route.1Routine sampling of airborne microorganisms is often performed indoors to monitor air quality. Sampling may be conducted in hospitals to assess infection risk, in the workplace to characterize air quality, or in response to a specific threat such as the potential release of a pathogen such as Bacillus anthracis or the SARS (Severe Acute Respiratory Syndrome) coronavirus. Routine biosampling may also be conducted to monitor outdoor air. Currently the federally funded BioWatch program, an early warning system designed to detect a biothreat agent within 36 hours of release, provides coverage in over 30 metropolitan areas across the United States.2,3
Sampling of bioaerosols can be performed by impaction onto growth media, impingement into liquid, or capture by filtration. The most common samplers in each category are used at flow rates between 2 and 22 L min−1 using a vacuum pump or personal pump. While impaction and impingement devices are more commonly used for bioaerosol sampling due to their ability to preserve microbial viability and often serve as reference samplers in evaluating other sampling devices,4–6 sampling by filtration is simple, portable, relatively inexpensive, and compatible with a number of analysis methods.
Indoor air quality concerns have provided the motivation behind most of the studies performed to date dealing with microorganisms on HVAC filter media. Kemp et al. studied the loading of HVAC filter media by outdoor air and performed fractional efficiency measurements to determine filter effectiveness against airborne microorganisms.7 A similar study was done by Foarde et al. who measured bioaerosol removal efficiencies of a room air cleaner with HVAC filter media.8 Kowalski et al. offered a theoretical solution to determining bioaerosol collection efficiency of HVAC and HEPA filter media.9 Other studies have been performed to examine the survival of microorganisms on HVAC filter media as functions of time,10 temperature,11 and relative humidity.11–13
There has recently been interest in exploiting building air-handling units (AHUs) as high volume bioaerosol filtration samplers.14 Existing HVAC filters, operating for a period of weeks or months, capture the bioaerosols present in the local air. If the captured microorganisms can be removed from the filters and separated from the crustal material, one may be able to determine the composition of the microbial populations in the air. The source of the bioaerosol could be investigated by removing HVAC filters from several AHUs within the same building; if the fresh-to-recirculated air ratio for each AHU is known, one could determine which bioaerosols were most likely from sources within the building and which were from sources outside the building. Although this same information could be gathered by employing a portable bioaerosol sampling device such as an AGI-30 or Andersen impactor, AHUs offer the advantage of handling flow rates that are typically four orders of magnitude greater than most portable sampling devices. Microorganisms present in concentrations as low as 0.01 colony-forming units (CFUs) per m3 can be detected using AHUs.14
This method of sampling is versatile and nonintrusive. Although obviously not a method that would provide early alarm of a biological attack, high volume sampling using HVAC media could determine the prevalence of natural background pathogens and pathogens’ near neighbors in ambient air. The results of such research would be useful for biosensor development in minimizing false positives.
In this study we present the methodology for studying HVAC filter loading and subsequent microorganism removal. We first discuss the test facility used to challenge the filter media and the suite of tests conducted for the validation of the facility, and then we present the filter loading and microorganism recovery and enumeration processes through which we determined the recovery efficiencies of four microorganisms: Bacillus subtilis var. niger, transmissible gastroenteritis virus (TGEV), avian pneumovirus (APV), and fowlpox virus (FPV). The results from this study can be used to predict the relative size of the populations of these species if present in the supply air of a building AHU, excluding time-dependent deactivation effects.
2. Materials
2.1 Test facility
The loading tests were conducted in a filter test facility in the Department of Mechanical Engineering at the University of Minnesota. The duct design conforms to ASHRAE Standard 52.2-1999, which delineates appropriate methodology for the testing of ventilation particulate air filters and other air cleaning devices.15 The facility is a closed-loop wind tunnel through which air can be continuously recirculated (Fig. 1). The test duct is constructed of stainless steel. The test section of the duct has a 61 cm × 61 cm cross section, just large enough to accommodate a standard AHU bag filter or pleated filter. The entire duct is composed of individual duct sections measuring between 90 and 150 cm, each section resting on frames equipped with wheels so sections may be easily added or removed. Rubber gaskets between sections prevent leakage. Vital components of the test duct include a centrifugal fan, a HEPA filter bank, an aerosol injection port, a pair of mixing baffles, a pair of upstream and downstream isokinetic sampling probes, and the test filter. For both pairs of sampling probes, the downstream probe geometry was identical to the upstream probe geometry so that the fraction of aerosol loss in each sampling tube could be assumed equal. The test filter is held in place with a removable frame, sealed on both the duct side and the filter side with adhesive rubber sealing strips to prevent leakage. The air flow rate, dry bulb temperature, and relative humidity of the air in the test duct can be controlled and monitored. The test section of the duct operates at a negative pressure to prevent bioaerosol from escaping into the room.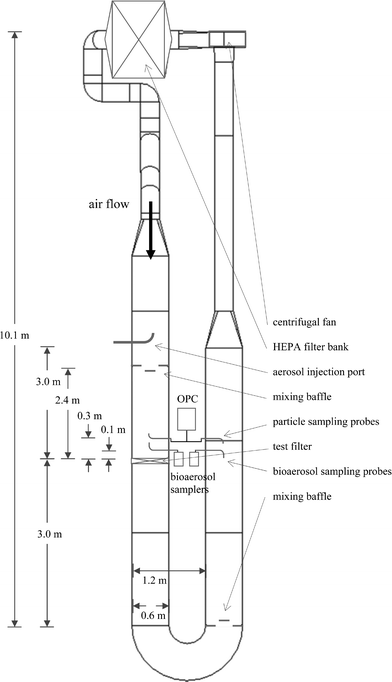 | ||
| Fig. 1 Layout of the test duct in the HVAC test facility. | ||
As a preventative measure, the laboratory that houses the test duct is equipped with an exhaust fan. The fan exhausts through a HEPA filter to the outdoors. The exhaust fan, when operating, depressurizes the laboratory by 25 Pa with respect to the adjacent spaces as recommended by the American Institute of Architects,16 ensuring containment of any bioaerosol that might escape from the test duct into the room.
2.2 Facility characterization
A number of qualification tests were performed on the test duct to validate the operation of the test facility. The qualification tests included a velocity uniformity test, an aerosol concentration uniformity test, a downstream mixing test, a test duct air leakage test, and a correlation test. All characterization test protocols conformed to ASHRAE Standard 52.2-1999 requirements. All tests were performed at test duct flow rates of 240 L s−1 and 940 L s−1 with the exception of the correlation test, which was performed only at 940 L s−1.2.3 Microorganisms
One bacterium and three viruses were selected for the loading tests. The microorganisms chosen were selected based on (1) nonpathogenicity, (2) availability, and (3) their relation to known pathogens. See Table 1 for additional details concerning the bacteria and viruses chosen.| Microorganism | Size | Source | Related pathogen |
|---|---|---|---|
| Bacteria | |||
| Bacillus subtilis var. niger | 0.9–1.2 μm |
Life Sciences Division (MT-L),
U.S. Army Dugway Proving Ground, Dugway, UT |
Bacillus anthracis |
| Viruses | |||
| Transmissible gastroenteritis virus (TGEV) | 50–85 nm |
American Type Culture Collection,
Manassas, VA, ATCC # VR-76 |
SARS coronavirus |
| Avian pneumovirus (APV) | 150–200 nm |
Clinical isolate from the U of M
Veterinary Diagnostics Laboratory, # MN 2a17 |
Respiratory syncytial virus (RSV) |
| Fowlpox virus (FPV) | 200–300 nm |
American Type Culture Collection,
Manassas, VA, ATCC # VR-249 |
Smallpox |
Bacillus subtilis var. niger was chosen as a representative spore-forming bacterium and a surrogate of Bacillus anthracis. B. subtilis is nonpathogenic and is widely used due to its ubiquity and hardiness, and has been used in various studies by investigators studying microorganism recovery efficiency or filter media capture efficiency.8,13,18,19 Two other bacteria, Mannheimia haemolytica and Yersinia ruckeri, were considered as representative vegetative bacteria and surrogates for pathogenic Pasteurella multocida and Yersinia pestis, respectively. However in a feasibility study, the loss of viability due to the stress caused by the nebulization and/or sampling processes prohibited us from having confidence in our upstream reference samplers for these species, and thus the filters were not challenged with these two bacteria.14
The viruses chosen were transmissible gastroenteritis virus (TGEV), avian pneumovirus (APV), and fowlpox virus (FPV). These animal viruses are nonpathogenic to humans and can be propagated and titrated in vitro in commonly available cell cultures.
2.4 Filters
The filters used in all the loading tests were Viledon Mini 95-2/44 filters (Freudenberg Nonwovens L. P., Hopkinsville, KY). These are MERV 14 filters and are typical of pleated media used in building AHUs. Technical data on these filters are provided in Table 2.| Dimensions | 61 cm × 61 cm × 5 cm |
| Pleats | 114 |
| Total media area | 4.8 m2 |
| Rated airflow | 5.6 × 104 L min−1 |
| Collection efficiency (1 μm) | 95% |
3. Methods
3.1. Bacteria
A new, unused Viledon filter was placed in the filter test frame and installed in the duct. B. subtilis was prepared at a concentration of (4.1 ± 0.8) × 108 CFUs mL−1 in deionized and filtered (DIF) water, and was nebulized using a Retec Aerosol Generator (Retec Development Laboratory, Portland, OR), having a flow rate of 1.5–3.0 L min−1 and a liquid aerosolization rate of approximately 60 μL min−1. The aerosol was mixed with 20 L min−1 dilution air in a drying column before passing through a 10 mCi Kr-85 charge neutralizer and being introduced into the test duct. The average air flow rate Qduct, temperature, and relative humidity during the test were 230 ± 3 L s−1, 23.0 ± 0.4 °C, and 14 ± 1%, respectively. The test duration Δttest was 30 min.Biosamplers (SKC Inc., Eighty Four, PA) are samplers similar to AGI-30s but reduce stress losses by impinging at an angle.20,21 These were used as reference samplers to determine the upstream and downstream bioaerosol concentrations. Coincident upstream and downstream samples were collected once during the test. The aerosol sampling rate Qimpinger was 12.5 L min−1 and the sampling duration Δtimpinger was 10 min. The Biosampler solution was 20 mL of phosphate buffered saline (PBS).
The particle concentration in the test duct was monitored upstream and downstream of the test filter using an 8-channel HIAC/Royco Model 5230 (Pacific Scientific Instruments, Elgin, IL) optical particle counter (OPC) sampling at 1 cfm. The aerosol size range of the OPC was 0.3–4.0 μm. Alternating 1-min upstream and downstream samples were taken at 1.5 min intervals over the course of the test. The OPC was used primarily for real-time monitoring to verify stability of the aerosol concentration vs. time but was also used as a secondary means of calculating the bioaerosol capture efficiency of the test filter.
The average upstream and downstream particle concentrations remained stable throughout the 30 min test with the CVs calculated to be 6% (n = 10) and 4% (n = 10), respectively. It was later verified that these CV were indicative not only of particle concentration stability but of bacterial concentration stability: running the identical test except filling the nebulizer with only DIF water revealed that nearly all of the airborne particles (99.96% upstream and 90% downstream) in the test duct consisted of bacteria.
Immediately following nebulization, the test duct was shut off and the test filter was removed. Using sterilized scissors, four 2.5 cm × 2.5 cm samples were cut from the filter at random locations. The filter samples were eluted in 10 mL of 0.02% Tween 80 by hand-shaking (∼500 shakes). Serial 2-fold dilutions were prepared from each of the four elutions, and two 50 μL aliquots from each dilution were inoculated on agar plates. The plates were incubated for 24 hours at 35 °C to culture the bacteria. CFUs formed on the plates, and the total number of CFUs from each filter section were determined. The total number of CFUs on the test filter, Nfilter, was calculated by multiplying the dilution ratio by the total number of CFUs counted and dividing it by the fraction of total filter area eluted.
This same diluting and plating process was repeated to culture bacteria from the Biosampler solution so that the total number of CFUs in the impinger solution, Nimpinger, could be determined. Nimpinger was calculated by multiplying the dilution ratio by the total number of CFUs counted and dividing it by the fraction of total impinger solution used.
3.2 Virus
Nine loading tests were conducted, three for each of the three viruses. Viruses suspended in maintenance medium at a concentration of ∼105 TCID50 (50% tissue culture infective dose) were nebulized using a 500 ELSD nebulizer (Alltech, Deerfield, IL). This nebulizer was chosen due to its fast aerosolization rate. It was capable of nebulizing 15 mL of liquid per minute, approximately 250 times the rate of the Retec nebulizer, minimizing time-dependent deactivation of the virus during the test. The nebulizer did not always run at full capacity however due to occasional clogging of the nebulizer nozzle. As a result, Δttest varied from 11–62 min (Table 3).| Virus | Trial | Δttest/min | Q duct/L s−1 | Dry bulb temp /°C | Relative humidity (%) | # AGI-30 samples |
|---|---|---|---|---|---|---|
| Transmissible gastroenteritis virus | ||||||
| 1 | 60 | 228 ± 2 | 24.9 ± 0.4 | 37 ± 2 | One | |
| 2 | 62 | 224 ± 4 | 22.3 ± 0.6 | N/A | One | |
| 3 | 16 | 232 ± 4 | 24.4 ± 0.4 | N/A | Two | |
| Avian pneumovirus | ||||||
| 1 | 32 | 228 ± 2 | 24.4 ± 0.4 | N/A | Three | |
| 2 | 20 | 228 ± 2 | 26.1 ± 0.4 | N/A | Three | |
| 3 | 13 | 218 ± 4 | 24.0 ± 0.4 | 9 ± 4 | Three | |
| Fowlpox virus | ||||||
| 1 | 12 | 496 ± 2 | 19.7 ± 0.4 | 79 ± 1 | One | |
| 2 | 11 | 490 ± 2 | 21.0 ± 0.5 | N/A | One | |
| 3 | 11 | 496 ± 2 | 21.4 ± 0.4 | N/A | One | |
The nebulized virus passed through a drying column and a 10 mCi Kr-85 charge neutralizer before combining with 25 L min−1 dilution air and being introduced into the test duct. Up to 200 mL of liquid was aerosolized per test, however less than 10 percent of it reached the test duct air stream. The majority of the aerosol gathered on the walls of the drying column and dripped into the sump below.
AGI-30 all-glass impinger samplers (Ace Glass, Inc., Vineland, NJ) were used as reference samplers.5,6 Coincident upstream and downstream samples were collected 1–3 times during each test. The aerosol sampling rate QAGI was 12.5 L min−1 and the sampling duration ΔtAGI was ≤10 min. The AGI sampler solution was 20 mL of MEM (Cellgro, MediaTech, Inc., Holly Hill, FL) and antibiotics. See Table 3 for environmental data pertaining to each test and AGI-30 sampling data.
Prior to each test, the background particle concentration in the test duct was determined using the HIAC/Royco OPC. Usually five, but no less than three, 1-min upstream and downstream samples were taken to determine the background concentration. In all 9 trials, the background particle concentration at the upstream and downstream sampling locations was less than 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 particles ft−3. This is approximately 100 times less than typical indoor room air concentration for the size range of the OPC. During each test, the particle concentration was monitored to verify output of the atomizer and to determine when the virus solution had run out. 1-min samples were taken at 1.5 min intervals.
000 particles ft−3. This is approximately 100 times less than typical indoor room air concentration for the size range of the OPC. During each test, the particle concentration was monitored to verify output of the atomizer and to determine when the virus solution had run out. 1-min samples were taken at 1.5 min intervals.
Following complete nebulization of the virus solution, the test duct was shut off and the test filter was removed.
The AGI-30 sampler solutions were serially diluted on 96-well tissue culture plates on which a monolayer of the host cells had been grown. The host cells used for the TGEV tests were swine testis (ST) cells; APV, African green monkey kidney (Vero) cells; fowlpox, chicken embryo fiberblast (CEF) cells. These cells were grown and maintained in MEM with Earle’s salt (Media Tech, Herndon, VA) containing antibiotics (150 IU mL−1 penicillin, 150 μg mL−1 streptomycin, 50 μg mL−1 neomycin, and 1 μg mL−1 fungizone), 8% fetal calf serum, and Edamin-S as additive (growth medium).
Six serial log base 10 step dilutions (0–5) were prepared from each impinger solution on the tissue culture plates. The plates were incubated at 37 °C for five days. Virus was detected by observing the wells for cytopathic effect (CPE) by direct microscopy. Plates for which CPE could not be verified by this method were retested by the indirect fluorescent antibody technique (IFAT), which involves binding viral proteins to fluorescein dye-labeled antibodies and viewing the wells under UV light. For each of the AGI-30 samples the TCID50 was calculated, and Nimpinger was determined from the dilution ratio.
The filter was divided into sixteen subquadrants (four sections wide by four sections high). One 3 cm × 8 cm filter sample was removed from each section of the filter and placed in 50 mL tubes for elution, four in a tube. The virus was eluted using 5 mL of 3% beef extract (BE) in 0.05 M glycine, pH 8.5 and vortexed for 20–30 s, three times in 1 min intervals. The filter samples were then removed, and the pH of the eluate was adjusted to 7.5 to prevent virus deactivation using 1 M HCl. The eluates were serially diluted in MEM on a 96-well tissue culture plate on which a monolayer of the host cells had been grown in the manner described previously. Six serial log base 10 step dilutions (1–6) were prepared. For each eluate the TCID50 was calculated, and Nfilter was determined.
Methodology differed slightly for the fowlpox tests. The samples cut from the filter were 3 cm × 4 cm instead of 3 cm × 8 cm. The filter samples were pooled into two groups instead of four. 10 mL of BE was used per tube for elution instead of 5 mL.
3.3. Data analysis
Although sampler type and culturing methodology differed between bacteria and viruses, the same formula was used for the calculation of collection and recovery efficiencies. The upstream and downstream bioaerosol concentrations (Cup and Cdown) were calculated according to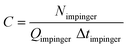 | (1) |
 | (2) |
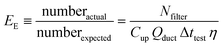 | (3) |
4. Results and discussion
4.1 Bacteria
The size distribution of the upstream and downstream samples obtained by the OPC is shown in Fig. 2 and can be compared to the size distribution of two types of B. subtilis cells obtained by An et al. shown in Fig. 3.23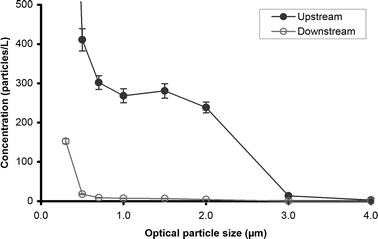 | ||
| Fig. 2 Upstream and downstream particle size distribution in test duct during B. subtilis test. | ||
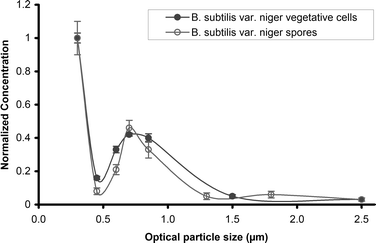 | ||
| Fig. 3 Upstream concentration of B. subtilis vegetative cells and spores, redrawn from An et al.23 | ||
Our size distribution is similar to the An et al. plot, although the peak near 1 μm corresponding to the approximate aerodynamic spore diameter of B. subtilis is less pronounced and is shifted to the right. The difference between these two size distributions is probably due to a larger amount of residual particles in our nebulized solution such as cell membranes and bacterial slime.24,25 An et al. minimized residual cell fragments and slime by washing the organisms three times with water by centrifugation prior to nebulization, whereas no washing was performed prior to our test.
The upstream and downstream bioaerosol concentrations calculated from the Biosampler data were 1500 ± 200 CFU L−1 and 50 ± 20 CFU L−1, respectively, thus the bioaerosol collection efficiency η of the test filter was 96.5 ± 1.5 percent.
A secondary calculation of bioaerosol collection efficiency was made using the OPC data. Assuming that the B. subtilis spore size is a function of local relative humidity as per Johnson et al.,26
| dp(μm) = 0.94 + 0.003 RH(%) | (4) |
N filter was calculated to be (7.1 ± 0.8) × 108 CFU, or approximately 1.5 × 104 CFU cm−2. Using eqn (3) it was determined that bacteria were recovered from the test filter with an efficiency of 105 ± 19 percent. This efficiency is similar to that seen during preliminary testing in which B. subtilis was recovered from used and unused filter media with efficiencies of 108 ± 7 percent and 109 ± 10 percent, respectively, when known amounts of bacterial suspension were pipetted onto the filter media and allowed to dry.14
It should be noted that efficiencies of >100% are not necessarily cause for alarm. All extraction efficiencies reported here are within one standard deviation of 100%, and as the averages were based on sample sizes of 4 or less, it is conceivable that a greater sample size would yield an efficiency of less than 100%. Alternatively, >100% efficiency may indicate that the microorganism was more efficiently extracted from the HVAC filter media than the reference sampler. Lin et al.21 report a relative recovery of 90–95% for B. subtilis from a Biosampler at a flow rate of 12.5 L min−1, therefore margin exists for the possibility that the HVAC filter media outperformed the reference sampler in this case.
4.2 Virus
The filter collection efficiency was not calculated for any of the virus tests because no virus was recovered from the downstream AGI-30 samplers. The virus collection efficiency was therefore estimated. Since ambient virus is rarely found alone in the form of a single virion but within some biological nuclei of much greater size (∼1 μm),27,28 the virus collection efficiency was estimated to be near the nominal filter collection efficiency. A collection efficiency of 100 percent was assumed. See Table 4 for recovery efficiencies as well as Cup and Nfilter for each trial.| Virus | Trial | C up/TCID50 L−1 | N filter/TCID50 | E E | ||
|---|---|---|---|---|---|---|
| AGI 1 | AGI 2 | AGI 3 | ||||
| Transmissible gastroenteritis virus | ||||||
| 1 | 5 | — | — | 1 × 105 | 2% | |
| 2 | 5 | — | — | 1 × 106 | 20% | |
| 3 | 41 | 33 | — | 1 × 105 | 1% | |
| Avian pneumovirus | ||||||
| 1 | 28 | 28 | 28 | 1 × 105 | 0.7% | |
| 2 | 10 | 5 | 15 | 1 × 105 | 3% | |
| 3 | 2 | 3 | 25 | 3 × 104 | 2% | |
| Fowlpox virus | ||||||
| 1 | 28 | — | — | 2 × 105 | 1% | |
| 2 | 0 | — | — | 1 × 105 | N/A | |
| 3 | 0 | — | — | 0 | 0% | |
A much smaller percentage of virus was recovered in these tests than in the TGEV tests we conducted previously using a smaller test apparatus (35 ± 45%), although identical elution and culturing methods were used. The most significant difference between the two sets of tests was face velocity. The face velocity during the preliminary tests was approximately 0.1 m s−1, while the face velocity in the tests described here was around 0.6 m s−1.
5. Conclusions
The loading of Viledon filter media with bacteria and virus aerosols at the Test Facility served as a pilot run for future filter loading tests. The ability to reliably measure the collection efficiency of the test filter using the facility’s reference samplers and to determine the recovery efficiency of bacteria and virus species using standard elution and enumeration methods was demonstrated.The measured collection efficiency for B. subtilis (97.6 ± 0.2 percent) can be compared to the findings of Foarde et al. who measured a collection efficiency of 96.7 ± 0.5 percent using single-stage Andersen samplers when challenging a self-contained Amway room air cleaner with B. subtilis.8 Our bioaerosol collection efficiencies are also near the particle collection efficiency of the filter media as given by the manufacturer (95% at 1 μm, 2.5 m s−1).
Our extraction efficiency (105 ± 19 percent) is comparable to the findings of other studies that have reported extraction efficiencies for B. subtilis using other filter media types. Burton et al. reported extraction efficiencies of 101–123 percent from polytetrafluoroethylene and mixed cellulose ester porous filters when using vortexing and mechanical shaking for elution,19 and Wang et al. report an EE of 96–98 percent from polycarbonate filters when using vortexing and ultrasonic agitation for elution.22 Our results indicate that B. subtilis can be removed from HVAC media with little loss of culturability. This implies that, neglecting time-dependent culturability losses or microbial proliferation, the airborne concentration of spore-forming bacteria upstream of the HVAC filters in building AHUs may be determined by eluting viable microorganisms from the filter media. An extensive field study was conducted under this hypothesis, and the results are currently being reviewed by authors of this paper.
TGEV, APV, and FPV aerosols were recovered from the test filter with efficiencies of 8 ± 11%, 2 ± 1%, and 0.5 ± 0.7%, respectively. As mentioned previously, feasibility testing of TGEV using a smaller-scale test duct with a slower air flow yielded higher extraction efficiencies (35 ± 45%). Although elution conditions were identical between the two sets of tests, there was a difference in the length of time between loading and elution. Whereas elution was performed on-site during the feasibility study, the lab work for these tests was not performed at the HVAC Test Facility. We estimate that up to two hours may have elapsed between loading and elution for these tests due to transport and drive time, while elution during the feasibility study most likely occurred within 10 minutes of removal of the filter from the test duct. Literature on virus survival indicates that this difference may have reduced our ability to recover live virus from the filter samples. Studies on virus survival indicate that the SARS coronavirus may persist only 12 hours on wood and cotton cloth,29 the human coronavirus persists only 3–6 hours on sterile sponges,30 and that respiratory syncytial virus persists just 2.5 hours on cloth gowns and 1 hour on paper towels.31 Relative humidity may have also influenced the survival of the virus, however there is no generalized model to predict microorganism deactivation with respect to relative humidity. Some viruses such as the human coronavirus (229E) survive better at high humidities;32 others, particularly those with lipids in their outer coat such as APV and TGEV, survive best at low humidities;33 still others such as the pigeon pox virus are unaffected by humidity.34 In general, virus half-lives are on the order of hours.35 We conclude that the use of AHUs as high volume sampling devices for live viruses is impractical if culture-based identification methods are used since most of the virus present on the HVAC filters would deactivate within a day. Molecular-based techniques such as the polymerase chain reaction (PCR) technique could be used to identify virus nucleic acid on HVAC filter media. The elution methodology presented here is compatible with such techniques.
Acknowledgements
This study was funded by the U.S. Department of Homeland Security (W91CRB-04-C-0035) through the Technical Support Working Group (TSWG) Task DC-2012A.References
- T. H. Kuehn, J. Sol. Energy Eng., 2003, 125, 366–371 CrossRef.
- A. D. Shea and A. L. Lister, The BioWatch program: detection of bioterrorism, 2003, Congressional Research Service Report No. RL 32152 Search PubMed.
- A. Emory and F. Light, EPA needs to fulfill its designated responsibilities to ensure effective BioWatch program, Environmental Protection Agency Evaluation Report No. 2005-P-00012, 2005 Search PubMed.
- M. P. Schafer, J. E. Fernback and M. K. Ernst, Aerosol Sci. Technol., 1999, 30, 161–173 CAS.
- C. S. Li, M. L. Hao, W. H. Lin, C. W. Chang and C. S. Wang, Aerosol Sci. Technol., 1999, 30, 100–108 CAS.
- B. Z. Predicala, J. E. Urban, R. G. Maghirang, S. B. Jerez and R. D. Goodband, Curr. Microbiol., 2002, 44, 136–140 CrossRef CAS.
- S. J. Kemp, T. H. Kuehn, D. Y. H. Pui, D. Velsey and A. J. Streifel, ASHRAE Trans., 1995, 101(1), 305–316 Search PubMed.
- K. K. Foarde, J. T. Janley, D. S. Ensor and P. Roessler, Aerosol Sci. Technol., 1999, 30, 223–234 CAS.
- W. J. Kowalski, W. P. Bahnfleth and T. S. Whittam, ASHRAE Trans., 1999, 105(2), 4–17 Search PubMed.
- P. C. Kemp, H. G. Neumeister-Kemp, G. Lysek and F. Murray, Atmos. Environ., 2001, 35, 4739–4749 CrossRef CAS.
- M. Möritz, H. Peters, B. Nipko and H. Rüden, Int. J. Hyg. Environ. Health, 2001, 203, 401–409 Search PubMed.
- S. J. Kemp, T. H. Kuehn, D. Y. H. Pui, D. Velsey and A. J. Streifel, ASHRAE Trans., 1995, 101(2), 228–238 Search PubMed.
- R. Maus, A. Goppelsröder and H. Umhauer, Atmos. Environ., 2001, 35, 105–113 CrossRef CAS.
- J. E. Farnsworth, Masters Thesis, University of Minnesota, 2005.
- ANSI/ASHRAE Standard 52.2-1999, Method of Testing General Ventilation Air-Cleaning Devices, ANSI/ASHRAE, Atlanta, 1999 Search PubMed.
- Guidelines for Design and Construction of Hospital and Health Care Facilities, AIA, Washington, DC, 2001 Search PubMed.
- S. M. Goyal, S. Chiang, A. Dar, K. V. Nagaraja, D. P. Shaw, D. A. Halvorson and V. Kapur, J. Vet. Diagn. Invest., 2001, 12, 116–168 Search PubMed.
- C. S. Li and Y. C. Lin, Sci. Total Environ., 2001, 278, 231–237 CrossRef CAS.
- N. C. Burton, A. Adhikari, S. A. Grinshpun, R. Hornung and T. Reponen, J. Environ. Monit., 2005, 7, 475 RSC.
- K. Willeke, X. J. Lin and S. A. Grinshpun, Aerosol Sci. Technol., 1998, 28, 439–456 CAS.
- X. J. Lin, T. Reponen, K. Willeke, Z. Wang, S. A. Grinshpun and M. Trunov, Aerosol Sci. Technol., 2000, 32, 184–196 CAS.
- Z. Wang, T. Reponen, S. A. Grinshpun, R. L. Górny and K. Willeke, J. Aerosol Sci., 2001, 32, 661–674 CrossRef CAS.
- H. R. An, G. Mainelis and M. Yao, Indoor Air, 2004, 14, 385–393 CrossRef CAS.
- Y. Qian, K. Willeke, V. Ulevicius, S. A. Grinshpun and J. Donnelly, Atmos. Environ., 1995, 29, 1123–1129 CrossRef CAS.
- S. Terzieva, J. Donnelly, V. Ulevicius, S. A. Grinshpun, K. Willeke, G. N. Stelma and K. P. Brenner, Appl. Environ. Microbiol., 1996, 62, 2264–2272 CAS.
- D. L. Johnson, T. A. Pearce and N. A. Esmen, Aerosol Sci. Technol., 1999, 30, 202–210 CAS.
- F. E. Buckland and D. A. S. Tyrell, Nature, 1962, 195, 1063–1064 CrossRef CAS.
- M. K. Ijaz, Y. G. Karim, S. A. Sattar and C. M. Johnson-Lussenburg, J. Virol. Methods, 1987, 18, 87–106 CrossRef CAS.
- World Health Organization, First data on stability and resistance of SARS coronavirus compiled by members of WHO laboratory network, 2003, http://www.who.int/csr/sars/survival_2003_05_04/en/ Search PubMed.
- J. Sizun, M. W. N. Yu and P. J. Talbot, J. Hosp. Infect., 2000, 46, 55–60 CrossRef CAS.
- C. B. Hall, R. G. Douglas and J. M. Geiman, J. Infect. Dis., 1980, 141, 98–102 CAS.
- M. K. Ijaz, A. H. Brunner, S. A. Sattar, R. C. Nair and C. M. Johnson-Lussenburg, J. Gen. Virol., 1985, 66, 2743–2748 Search PubMed.
- A. J. Mohr, in Modeling the environmental fate of microorganisms, ed. C. J. Hurst, ASM Press, Washington, DC, 1991, pp. 160–190 Search PubMed.
- S. J. Webb, Can. J. Microbiol., 1967, 13, 733–742 CrossRef CAS.
- M. D. Sobsey and J. S. Meschke, Proceedings of the World Health Organization International SARS Symposium, Rome, 2003 Search PubMed.
| This journal is © The Royal Society of Chemistry 2006 |
