Hot off the Press
In the Hot off the Press section of Molecular BioSystems members of the Editorial Board and their research groups highlight recent literature for the benefit of the community. This month the highlighted topics include the development of transcriptional networks, the use of systems biology in classifying cancers, and a new method for protein labelling in bacterial cells.
Protein labelling using alkyne–azide methodology
To be able to study protein interactions within the cell, one has to be able to visualise and track them within complex biological systems. Fluorescence is one of the most widely used screening techniques and there is a constant need for new and efficient methods for fluorescent protein labelling. Many methods so far rely on the use of green fluorescent protein (GFP) and its variants as fluorescent labels. However, the bulk of GFP and the need for genetic manipulations of the target protein limit the applicability of this method.Recently, David Tirrell and co-workers have described an elegant and simplified approach to protein labelling. They used histidine-tagged barstar as the model protein to which non-natural amino acids homopropargylglycine (Hpg) and ethynylphenylalanine (Eth) containing alkyne groups were incorporated by expression. Hpg and Eth serve effectively as Met and Phe analogues. Once the protein containing these amino acids is expressed, the E. coli cell cultures are allowed to react with an azide derivative of coumarin in the presence of Cu(I) catalyst (Fig. 1).
Confocal microscopy was employed to examine fluorescence and only the cells containing barstar protein with non-natural amino acids were fluorescent. Additionally, gel electrophoresis of crude lysate proved that labelling occurred selectively on newly synthesised proteins.
These results demonstrate that alkyne–azide coupling can be used as an effective method of protein labelling in bacterial cells and it will be interesting to see how it transfers to other enzymes, particularly those expressed in mammalian cells.
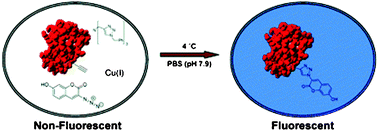 | ||
| Fig. 1 Reproduced with permission from J. Am. Chem. Soc., 2005, 127, 14150–14151. Copyright 2005 American Chemical Society. | ||
K. Beatty, F. Xie, Q. Wang and D. A. Tirrell, J. Am. Chem. Soc., 2005, 127, 14150–14151
Reviewed by: Ljiljana Fruk, Universität Dortmund, GermanyDevelopment of intricate transcriptional networks
Deciphering the transcriptional networks responsible for human development is a challenge whose answers will prove invaluable to understanding the basics of human development. Richard A. Young and colleagues have developed a strategy to construct such a network by focusing on the targets of transcription factors. Specifically, this group used human embryonic stem cells to study the targets of OCT4, SOX2, and NANOG.Using genome-wide mapping techniques, they found that these factors not only regulate two distinct gene sets, they also co-occupy many of these sites. One set of these genes is activated by the presence of these three factors and contributes to pluripotency and self-renewal of the cell. The second set of genes is repressed by these same three factors and these genes are linked to developmental processes.
By combining these results with the current literature, Young and colleagues pieced together a novel transcriptional network that begins to explain how stem cells regulate pluripotency, self-renewal, and differentiation.
By expanding this approach to other transcription factors, the network will be elaborated to gain a better understanding of how cells are regulated and how we as scientists can possibly control cell fate.
Laurie A. Boyer, Tong Ihn Lee, Megan F. Cole, Sarah E. Johnstone, Stuart S. Levine, Jacob P. Zucker, Matthew G. Guenther, Roshan M. Kumar, Heather L. Murray, Richard G. Jenner, David K. Gifford, Douglas A. Melton, Rudolf Jaenisch and Richard A. Young, Cell, 2005, 122, 947–956
Reviewed by: Melissa O’Neal, Division of Translational Research, University of Texas Southwestern Medical Center, USAQuantifying the polar wind
The positioning of chromosomes in the cell is determined by a balance of pulling forces developed by the kinetochore machinery and pushing polar ejection forces generated by motor proteins associated with the chromosome arms (chromokinesins). Determining the magnitude of such forces has been proven to be very difficult in vivo. Gary Brouhard and Alan Hunt have recently developed an interesting in vitro assay to measure the forces developed by chromokinesins.Manipulating a bead to which a microtubule is attached with optical tweezers, and bringing this construct into contact with isolated chromosomes, they determined the force that is generated by several chromokinesins. The determination and interpretation of the forces is not obvious, as they are on the limit of the resolution of the tweezers. Nevertheless, their measurements indicate that the force applied by chromokinesins is about 1 pN. This is quite low compared to the typical forces that other motor proteins develop. They suggest that many distributed low force generators may be more appropriate than fewer stronger motors, as these could damage the chromosomes (Fig. 2).
Such in vitro study provides important insight on the origin of the so-called “polar wind”, and are also important for quantifying the forces involved in mitosis.
Gary J. Brouhard and Alan J. Hunt, Proc. Natl. Acad. Sci. U.S.A., 2005, 102(39), 13903–13908
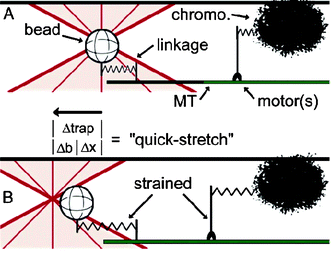 | ||
| Fig. 2 Reproduced with permission from Proc. Natl. Acad. Sci. U.S.A., 2005, 102(39), 13903–13908. Copyright 2005 National Academy of Sciences, U.S.A. | ||
A systems biology approach to classify poor-prognosis neuroendocrine cancers
There is a pressing need to identify new cancer biomarkers and mediators of tumorigenesis for better diagnosis and therapeutic strategies. To this end Jeff Gordon and colleagues have worked on an integrated systems biology approach to stratify neuroendocrine (NE) cancers (NEC) by prognosis.Their approach applies functional genomics and mass-spectrometry (MS)-based metabolomics to analyze transgenic mice that develop metastatic cancer from prostate NE cell lineage (CR2-TAg), NE cell lines derived from these mice, and human NE tumors of varying aggressiveness.
Gene expression profiles of primary prostate tumors and metastasis from CR2-TAg transgenic mice reveal glutamic acid decarboxylase (Gad1) to be a highly enriched enzyme in NE tumors. These cells also possess transcripts encoding biosynthetic enzymes for GABA via an alternative Gad1 independent pathway involving amiloride-binding protein 1 (ABP1). The ultimate metabolic outcome of these elevated transcripts was confirmed by MS studies, which identified elevated GABA levels in prostate NEC cells, primary tumors, and venous blood of CR2-TAg mice compared to normal controls.
In addition to its role in GABA biosynthesis, ABP-1, which is almost exclusively present in poor prognosis NE tumors, participates in the formation of imidazole-4-acetate. Electrophysiological experiments have revealed that like GABA, imidazole-4-acetate can activate GABAA receptors found on PNEC cells and neighboring normal prostate epithelial cells. These findings suggest that, in poor prognosis NECs, GABA signaling pathways may contribute to the pathological properties of neoplastic NE cells within a tumor microenvironment.
These metabolic pathways were also shown to be useful in distinguishing poor-prognosis metastatic NE tumors from benign NE tumors through the successful assignment of prognosis to various lung cancers that match their respective survival rates solely through analysis of gene expression datasets.
In summary, the Gordon group has shown that an integrated genomic and metabolomic approach is capable of identifying pathogenesis related metabolic pathways, which may serve as potential therapeutic targets.
J. E. Ippolito, J. Xu, S. Jain, K. Moulder, S. Mennerick, J. R. Crowley, R. R. Townsend, and J. I. Gordon, Proc. Natl. Acad. Sci. U.S.A., 2005, 102, 9901
Reviewed by: Kyle P. Chiang, The Scripps Research Institute, California, USAViral nanotemplates
The search for a successful and affordable method of nanoparticle patterning is one of the important tasks in the production of nanoscale devices. Some biological components, among them DNA and viruses, have been used as templates to produce nanopatterns. Recently, James Culver and colleagues combined the nucleic acid hybridisation principle with the ability of some viruses to act as a template to produce a structured assembly of viruses on silicon oxide wafers.Firstly, gold patterns were fabricated on wafers using photolithography. Then electrodeposition of amino polysaccharide chitosan was performed followed by glutaraldehyde activation and subsequent covalent conjugation of 24mer amino DNA. Such capture DNA was used to immobilise tobacco mosaic virus (TMV) through hybridisation of a part of the viral genome. Additionally TMV was genetically modified to contain cysteine groups in the protein coat. That allowed for fluorescent labeling of the viruses with fluorophore maleimides and subsequent screening of the TMV immobilised on the chips. SEM was used to check the orientation and the assembly of the virus templates and indeed, the patterns were easily recognisable (Fig. 3).
To prove that the TMV can act as a nanotemplate, gold and platinum clusters were successfully assembled onto the TMV surfaces and the authors anticipate that this strategy could be further utilised in the construction of nanodevices.
H. Yi, S. Nisar, S. Y. Lee, M. A. Powers, W. E. Bentley, G. F. Payne, R. Ghodssi, G. W. Rubloff, M. T. Harris, J. N. Culver, Nano Lett., 2005, 5 (10), 1931–1936.
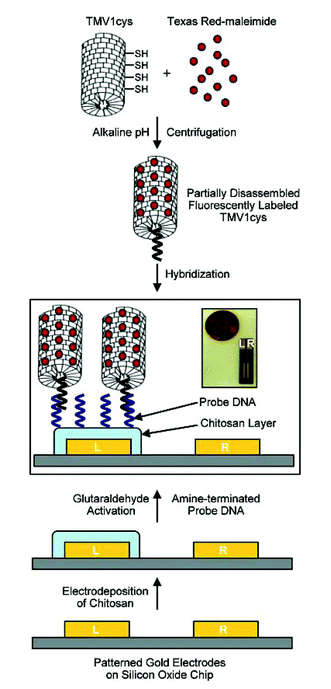 | ||
| Fig. 3 Reproduced with permission from Nano Lett., 2005, 5 (10), 1931–1936. Copyright 2005 American Chemical Society. | ||
| This journal is © The Royal Society of Chemistry 2005 |
