Direct observation of the wrapping/unwrapping of ssDNA around/from a SWCNT at the single-molecule level: towards tuning the binding mode and strength†
Abstract
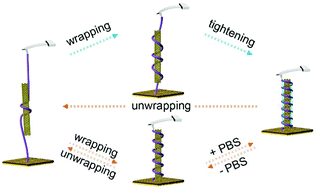
Complexation of single-stranded DNA (ssDNA) with a chiral single-walled carbon nanotube (SWCNT) exhibits surprising efficacy in CNT dispersion and sorting, optical sensing, and nanoelectronic device design. Studying the wrapping/unwrapping mechanism is challenging because an in situ method at the single-molecule level is required. Here, we developed a method based on single-molecule force spectroscopy to monitor the unwrapping/wrapping of ssDNA from/around a SWCNT. Our results reveal that the wrapping/unwrapping processes are reversible in water, and these processes occur in an equilibrium manner driven mainly by π–π interactions between DNA bases and CNTs. In phosphate buffered saline, the unwrapping process is loading rate-dependent, and ssDNA wrapping around a CNT undergoes two distinct stages dominated by both π–π interactions and hydrogen bonding. In addition, our results show that salts could further stabilize ssDNA/CNT complexes by blocking the electrostatic interactions between adjacent DNA segments and by catalyzing the formation of hydrogen bonds between DNA bases. The stability of ssDNA/CNT is dependent on the DNA sequence and CNT chirality. These results deepen our understanding of ssDNA–CNT interactions and provide effective means to tune the binding mode and strength.


 Please wait while we load your content...
Please wait while we load your content...