High-throughput co-former screening and structural elucidation using resonant acoustic mixing and 3D electron diffraction
Jacob
Danks
ab,
Daniel N.
Rainer
 c,
Ahmed S.
Hamza
c,
Ahmed S.
Hamza
 a,
Simon J.
Coles
a,
Simon J.
Coles
 c,
Anthony B.
Carter
c,
Anthony B.
Carter
 a and
Joseph E. G.
Benson
a and
Joseph E. G.
Benson
 *a
*a
aProcess and Analytical Chemistry, Pharmaron UK Ltd., Hoddesdon, UK. E-mail: joseph.benson@pharmaron-uk.com
bSchool of Chemistry, University of Lincoln, LN6 7DJ, UK
cSchool of Chemistry and Chemical Engineering, University of Southampton, SO17 1BJ, UK
First published on 17th March 2026
Abstract
Integrating 3D electron diffraction (3D ED) analysis into the screening workflow of multicomponent systems facilitates direct structural determination from high-throughput resonant acoustic mixing (RAM). This screening method enables the investigation of a wide experimental landscape, including various co-formers and solvents, within a short timeframe. Consequently, it allows for the rapid and efficient generation of numerous potential novel materials. Hit identification from the screening process is conducted through high-throughput X-ray powder diffraction (XRPD) and Raman microscopy, followed by structural determination of selected novel phases by 3D ED. This workflow has been demonstrated using sulfasalazine as an example active pharmaceutical ingredient (API), encompassing a broad range of crystallisation conditions. Multiple distinct diffractograms were observed, leading to the discovery and structural characterisation of two novel multicomponent systems of sulfasalazine by 3D ED, one of which shows evidence of tautomerisation within a single crystal.
Many APIs exist as crystalline materials1 and may exhibit polymorphism, meaning they can exist in many different crystalline forms. An understanding of this polymorphic landscape and associated physical properties is key to the development of a medicine.2 Where APIs have undesirable physical characteristics, multicomponent systems (including salts and co-crystals) can provide a route to improved solid-state properties, such as solubility or bioavailability.3
In the pharmaceutical industry, cost and time of development are important considerations. High throughput experimentation enables a wide range of experimental solid form production conditions to be screened quickly using minimal material resources.4 Common solution-based techniques include solvent evaporation, cooling crystallisation5 and slurry/reactive crystallisation.6 Mechanochemistry7 and melt-crystallisation8 techniques are widespread for solid-based experimentation and remove the effect of solvent,9 enabling a wider crystallisation landscape to be explored.
Mechanochemistry uses mechanical energy supplied to the system through collisions or grinding to generate multicomponent systems. This can be performed manually or through automated processes such as ball milling10 or, more recently, resonant acoustic mixing (RAM).11 RAM uses oscillation at the resonant frequency with relatively large displacement to transfer energy efficiently to the system. As well as the efficient mixing potential, other applications have been investigated. Ende et al.12 showcased the ability of RAM for co-crystal formation. More recent work carried out by Nagapudi et al.13 demonstrated the use of RAM for high throughput 96-well plate screening and showed that the addition of a minimal amount of a liquid additive prior to mixing improved the success of multicomponent system formation. In this study, we demonstrate a high throughput RAM workflow with sulfasalazine and a selection of co-formers, using XRPD and Raman for hit prioritisation and 3D ED for structural elucidation of two novel multicomponent systems.
Sulfasalazine (SSZ) is an antirheumatic drug used in the treatment of rheumatoid arthritis and ulcerative colitis.14,15 It is a biopharmaceutical classification system (BCS) class IV molecule and therefore, in the pharmaceutical industry, salt or cocrystal formation would be considered to improve its low solubility and permeability. SSZ has two tautomeric conformations referred to as the imide (monoclinic) and the amide (triclinic) forms.16,17 The change in the pyridine functionality on the SSZ molecule between the two tautomeric forms affects the potential for cocrystal formation at this site.
Structural elucidation is a key step in the process of patenting of novel drug molecules. Typically, large crystals are grown for analysis by single-crystal X-ray diffraction (SCXRD),18 but this can be time consuming, poses challenges for complex multicomponent systems and is not necessarily representative of as-manufactured solid forms. Whilst structural elucidation is possible directly from the XRPD diffractograms obtained from screening,19 this is both labour and time intensive, requiring larger quantities of high purity material to generate high quality powder diffraction data, and ab initio structure solution is difficult and prone to failure. High throughput XRPD for potential novel phase identification coupled with 3D ED is quicker and requires less high purity material, whilst both techniques can be performed from a single RAM experiment. Our study showcases the ability of 3D ED to successfully determine the structures of two novel SSZ multicomponent systems obtained directly from RAM during high throughput screening.
Co-crystals of SSZ have been reported with theobromine (Bala et al.20), trimethoprim and nicotinamide (Elbakush21). In 2023, Huang et al. investigated the crystal landscape of SSZ using slurry, evaporation and liquid-assisted grinding (LAG) methods with various salt and cocrystal formers.22 They identified six crystal structures by SCXRD, which comprised both salts and co-crystals; various other co-formers showed distinct diffractograms, however, single-crystal structures were not obtained. The study performed by Babu et al.23 showed that the most statistically likely synthons to form were carboxylic acid–pyridine interactions.24,25
Co-formers (Table 1) were screened in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
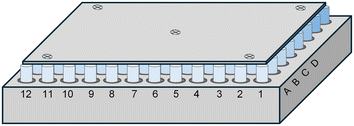
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometric ratio with SSZ in four organic solvents in a 96-well plate loaded with shell vials (Fig. 1). Parameters used for the mixing were derived from those investigated by Nagapudi et al.13 The solid components were combined in an equimolar ratio and pre-mixed at 30 g (acceleration due to gravity) for 5 minutes. The appropriate solvent (0.5 mL g−1 SSZ) was added and a second mixing was performed at 60 g for 2 hours. The resulting solids were analysed by XRPD and confocal Raman spectroscopy on a 96-well Kapton plate and by thermal methods.
1 stoichiometric ratio with SSZ in four organic solvents in a 96-well plate loaded with shell vials (Fig. 1). Parameters used for the mixing were derived from those investigated by Nagapudi et al.13 The solid components were combined in an equimolar ratio and pre-mixed at 30 g (acceleration due to gravity) for 5 minutes. The appropriate solvent (0.5 mL g−1 SSZ) was added and a second mixing was performed at 60 g for 2 hours. The resulting solids were analysed by XRPD and confocal Raman spectroscopy on a 96-well Kapton plate and by thermal methods.
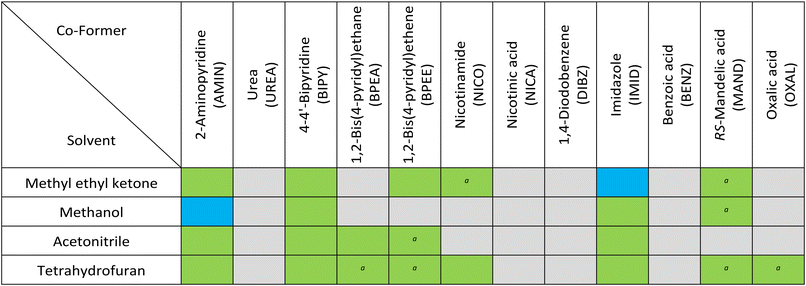
| a Some unreacted SSZ or co-former observed. |
|---|
|
Distinct crystalline diffractograms (i.e. patterns that did not completely match SSZ tautomers or co-former) were identified in 48% of conditions screened (Table 1), showcasing the effectiveness of the technique (data in SI). Additionally, there was a considerable time saving compared to alternate methods such as solvent evaporation. XRPD diffractograms of multicomponent systems SSZ-NICO and SSZ-BPEE are, to our knowledge, reported for the first time (Fig. S5). Two hits from aminopyridine and imidazole (SSZ-AMIN and SSZ-IMID), were chosen for further study based on crystallinity and distinction from input materials. An SSZ-IMID system was previously reported as an acetonitrile solvate,22 but no anhydrous form has been reported. Similarly, an XRPD pattern corresponding to the SSZ-AMIN system was previously reported22 but does not appear to match our powder data. Both were suitable for structural elucidation by 3D ED without further manipulation.
SSZ-AMIN: the XRPD diffractogram was distinct from AMIN and both tautomers of SSZ. DSC showed a single endothermic event with onset of 196.2 °C, indicating that the obtained material was phase pure. The endothermic event was observed between the melts of the two solid components (SSZ: 260.6 °C; AMIN: 57.0 °C), which was further evidence of formation of a multicomponent system.26 This was supported by the presence of new peaks (1138 cm−1 and 1435 cm−1) in the Raman spectrum resulting from the formation of the SSZ carboxylate anion and its interaction with the AMIN pyridinium cation.
SSZ-IMID: the XRPD diffractogram was distinct from IMID and both tautomers of SSZ. DSC showed an endothermic melting event with onset of 184.9 °C. There was a small endothermic event at lower temperature which may be attributed to unreacted IMID. The melt occurred between the melts of the two solid components (SSZ: 260.6 °C; IMID: 89.0 °C) which again was evidence of formation of a multicomponent system. This was supported by the presence of new peaks (1135 cm−1 and 1430 cm−1) in the Raman spectrum resulting from, as with SSZ-AMIN, the formation of the SSZ carboxylate anion and its interaction with the IMID imidazolium cation. Data for both systems is included in the SI.
Structure determination for both samples could be performed in a straight-forward manner. Vacuum and beam damage encountered in electron diffraction were reduced by performing the experiment at 175(5) K. In both cases, data collected from several particles could be indexed into the same unit cell and were subsequently merged into a single dataset.
The structure of SSZ-AMIN was determined in a monoclinic space group P2/n 19.50, 6.29, 19.70, 90, 106.13, 90, Z′ = 1. The carboxylic acid is deprotonated in favour of the co-former amino-pyridine, now in its protonated form as the aminopyridinium cation. This assignment is supported by the C–O bond lengths of 1.251(9) and 1.277(8) Å, for C18–O3 and C18–O4, respectively, as well as the difference map, which indicated the position of the proton to be in close proximity to the pyridine-N of the co-former. Indeed, freely refining the proton (both position and isotropic displacement parameter) yielded a stable refinement and a N5–H5b bond length of 0.92(5) Å. The second important hydrogen bonding interaction occurs on the other end of the SSZ molecule, where its pyridine N (N1) is also protonated (N1–H1: 1.03(4) Å), derived from the analogous observations as above. H1 is hydrogen-bonded to the sulfonimide nitrogen (N2) of the next molecule. A competitive refinement of the proton bound to either of the two nitrogen atoms (N1 or N2) was also attempted, resulting in the occupancy on N1 converging to unity. The simulated powder pattern was consistent with the experimental data (Fig. 2), which was confirmed by Rietveld refinement (see SI).
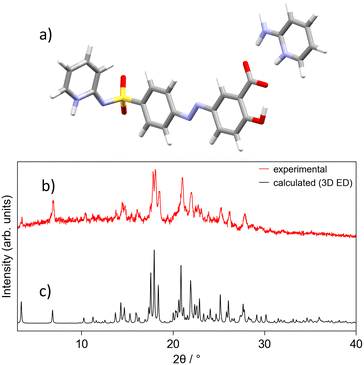 | ||
| Fig. 2 SSZ-AMIN structural data: a) 3D ED asymmetric unit; b) experimental powder pattern (298 K); c) simulated powder pattern (175(5) K). | ||
SSZ-IMID, on the other hand, crystalised in a triclinic lattice, P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) 8.12, 10.37, 26.28, 80.41, 86.62, 88.20, Z′ = 2. The carboxylic acid of SSZ is deprotonated for both molecules in the asymmetric unit and engaged in hydrogen bonding with the co-former imidazole. Once again, the simulated powder pattern agrees well with experimental data (Fig. 3), which was confirmed by Rietveld refinement (see SI). Procedurally, the same approach for locating hydrogen atoms/protons was taken as for SSZ-AMIN. The carboxylic acid groups for both molecules in the asymmetric unit are deprotonated and hydrogen-bonded to the imidazolium cations. Interestingly, the hydrogen atoms involved in the sulfonamide–pyridine interaction indicate that this crystal structure exhibits imide/amide tautomerism. Competitively refining the hydrogen positions for N1/N2 resulted in the same outcome as the SSZ-AMIN sample, with a complete protonation to form the pyridinium cation on N1. However, the same strategy led to ambiguity in the second molecule in the asymmetric unit, where the location of the proton appeared to be split between nitrogen N5 (pyridine/-ium) and N6 (sulfonamide/-imide). Refinement of the occupancies resulted in about 42% and 58% for H5 and H6, respectively. As the dataset used for structure determination of SSZ-IMID was obtained by merging data from 6 individual collections to overcome issues with the low completeness, it is possible that both forms are present, yet each crystal comprising only one of the forms. Thus, the obtained structure model was used as a starting model and the occupancies for the hydrogen atoms in question refined against each data set (the corresponding CIF files are in the SI). The results show that the individual crystals, and therefore the sample as a whole, are of one or the other tautomer (see SI, Table S3). Surprisingly, one of the data collections shows strong evidence for a roughly 45/55 split of the hydrogen bonded to the nitrogen atoms N5 and N6, respectively, suggesting a tautomeric behaviour within a single-crystal, to our knowledge this is the first example of such behaviour in a multicomponent system.
8.12, 10.37, 26.28, 80.41, 86.62, 88.20, Z′ = 2. The carboxylic acid of SSZ is deprotonated for both molecules in the asymmetric unit and engaged in hydrogen bonding with the co-former imidazole. Once again, the simulated powder pattern agrees well with experimental data (Fig. 3), which was confirmed by Rietveld refinement (see SI). Procedurally, the same approach for locating hydrogen atoms/protons was taken as for SSZ-AMIN. The carboxylic acid groups for both molecules in the asymmetric unit are deprotonated and hydrogen-bonded to the imidazolium cations. Interestingly, the hydrogen atoms involved in the sulfonamide–pyridine interaction indicate that this crystal structure exhibits imide/amide tautomerism. Competitively refining the hydrogen positions for N1/N2 resulted in the same outcome as the SSZ-AMIN sample, with a complete protonation to form the pyridinium cation on N1. However, the same strategy led to ambiguity in the second molecule in the asymmetric unit, where the location of the proton appeared to be split between nitrogen N5 (pyridine/-ium) and N6 (sulfonamide/-imide). Refinement of the occupancies resulted in about 42% and 58% for H5 and H6, respectively. As the dataset used for structure determination of SSZ-IMID was obtained by merging data from 6 individual collections to overcome issues with the low completeness, it is possible that both forms are present, yet each crystal comprising only one of the forms. Thus, the obtained structure model was used as a starting model and the occupancies for the hydrogen atoms in question refined against each data set (the corresponding CIF files are in the SI). The results show that the individual crystals, and therefore the sample as a whole, are of one or the other tautomer (see SI, Table S3). Surprisingly, one of the data collections shows strong evidence for a roughly 45/55 split of the hydrogen bonded to the nitrogen atoms N5 and N6, respectively, suggesting a tautomeric behaviour within a single-crystal, to our knowledge this is the first example of such behaviour in a multicomponent system.
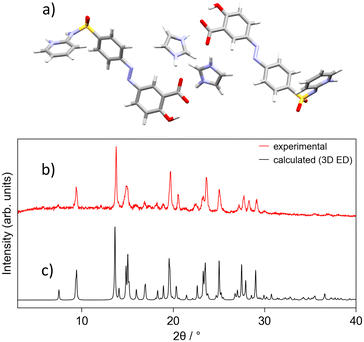 | ||
| Fig. 3 SSZ-IMID structural data: a) 3D ED asymmetric unit; b) experimental powder pattern (298 K); c) simulated powder pattern (175(5) K). | ||
Comparing the crystal structure models with each other, as well as with literature data, produces several noteworthy observations. Sulfasalazine itself has been reported in its amide form in space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) and in its imide form where crystals exhibit the monoclinic space group P2(1)/c.17 In both cases, the main body of the molecule from the carboxylic acid to the sulfur atom is planar with a positive torsion angle between this part and the pyridyl group (N2–S1–C6–C11). This torsion angle, however, differs significantly between the two forms, with an almost perpendicular arrangement in the triclinic lattice (86.4°) and a much more acute angle of about 44° in the monoclinic form. The crystal structure for SSZ-AMIN is similarly planar in the bulk part and a positive torsion angle, yet being much larger with about 79°. A similar geometry has also been observed by Huang et al. for a SSZ salt with 4,4′-trimethylenepyridine (amide, triclinic) and a cocrystal with 1,2-bis(4-pyridyl)ethane (imide, triclinic).22
and in its imide form where crystals exhibit the monoclinic space group P2(1)/c.17 In both cases, the main body of the molecule from the carboxylic acid to the sulfur atom is planar with a positive torsion angle between this part and the pyridyl group (N2–S1–C6–C11). This torsion angle, however, differs significantly between the two forms, with an almost perpendicular arrangement in the triclinic lattice (86.4°) and a much more acute angle of about 44° in the monoclinic form. The crystal structure for SSZ-AMIN is similarly planar in the bulk part and a positive torsion angle, yet being much larger with about 79°. A similar geometry has also been observed by Huang et al. for a SSZ salt with 4,4′-trimethylenepyridine (amide, triclinic) and a cocrystal with 1,2-bis(4-pyridyl)ethane (imide, triclinic).22
In contrast to the previously mentioned structures, SSZ-IMID contains 2 molecules in the asymmetric unit. The torsion angles for both molecules are negative (about −138° and −129°) and intriguingly, the main body of the second independent molecule of SSZ-IMID is not planar but slightly distorted with a torsion angle across between the two benzene rings (C28–C27–C30–C35) of 144°. This is a stark contrast to the other molecule in the asymmetric unit, which has a torsion angle of −179°. Some recently discovered salts and cocrystals also show an analogous distortion, a similar example being an imidazole salt albeit solvated with acetonitrile.22
The outcomes of the 3D ED study showcase one of the great strengths of the method, being positioned between single particle analysis (as generally adopted in single-crystal diffraction techniques such as single-crystal XRD), and a bulk diffraction measurement such as powder XRD. Equally, the importance of meticulous data analysis for 3D ED structures is highlighted, as the often-performed merging process may obscure underlying structural intricacies of the sample being investigated.
In conclusion, we report novel structures of two multicomponent sulfasalazine systems. Both were obtained via high-throughput resonant acoustic mixing experiments and characterised without further manipulation using powder X-ray diffraction, Raman spectroscopy, and 3D electron diffraction. 3D ED screening is shown to readily provide not only rapid, unambiguous structure solution on a nanoparticle-by-nanoparticle basis, but also reveal structural insights only possible with atomic resolution structures. The distribution of electron diffractometers is currently limited but growing quickly, so we anticipate the workflow has the potential to significantly increase efficiency in the pharmaceutical industry as the barrier to entry decreases.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). See DOI: https://doi.org/10.1039/d5ce01188d.Raw data for all the 3D ED experiments have been deposited with the 3D ED Zenodo community and can be accessed at https://doi.org/10.5281/zenodo.14653435.
CCDC 2417286 and 2417287 contain the supplementary crystallographic data for this paper.27a,b
Acknowledgements
Jacob Danks would like to thank the University of Lincoln for its support during his industrial placement at Pharmaron. The authors would like to thank Alexandra Shipilova for help preparing figures. D. N. R. and S. J. C. thank the EPSRC for funding (EP/X014444/1, A National Electron Diffraction Facility for Nanomaterial Structural Studies).Notes and references
- S. R. Vippagunta, H. G. Brittain and D. J. W. Grant, Adv. Drug Delivery Rev., 2001, 48(1), 3–26 Search PubMed.
- S. L. Morissette, S. Soukasene, D. Levinson, M. J. Cima and Ö. Almarsson, Proc. Natl. Acad. Sci. U. S. A., 2003, 100(5), 2180–2184 CrossRef.
- D. Gupta, D. Bhatia, V. Dave, V. Sutariya and S. V. Gupta, Molecules, 2018, 23(7), 1719 CrossRef PubMed.
- S. L. Morissette, Ö. Almarsson, M. L. Peterson, J. F. Remenar, M. J. Read, A. V. Lemmo, S. Ellis, M. J. Cima and C. R. Gardner, Adv. Drug Delivery Rev., 2004, 56(3), 275–300 CrossRef PubMed.
- J. P. Metherall, R. C. Carroll, S. J. Coles, M. J. Hall and M. R. Probert, Chem. Soc. Rev., 2023, 52, 1995–2010 RSC.
- M. M. Haskins and M. J. Zaworotko, Cryst. Growth Des., 2021, 21, 4141–4150 CrossRef PubMed.
- L. M. Martínez, J. Cruz-Angeles, M. Vázquez-Dávila, E. Martínez, P. Cabada, C. Navarrete-Bernal and F. Cortez, Pharmaceutics, 2022, 14(10), 2003 CrossRef PubMed.
- S. Jia, Z. Gao, N. Tian, Z. Li, J. Gong, J. Wang and S. Rohani, Chem. Eng. Res. Des., 2021, 166, 268–280 CrossRef.
- S. Chewle, F. Emmerling and M. Weber, Crystals, 2020, 10(12), 1107 CrossRef.
- M. Solares-Briones, G. Coyote-Dotor, J. C. Páez-Franco, M. R. Zermeño-Ortega, C. M. de la O. Contreras, D. Canseco-González, A. Avila-Sorrosa, D. Morales-Morales and J. M. Germán-Acacio, Pharmaceutics, 2021, 13(6), 790 CrossRef PubMed.
- A. L. Michalchuk, K. S. Hope, S. R. Kennedy, M. V. Blanco, E. V. Boldyreva and C. R. Pulham, Chem. Commun., 2018, 54, 4033–4036 RSC.
- D. J. am Ende, S. R. Anderson and J. S. Salan, Org. Process Res. Dev., 2014, 18(2), 331–341 CrossRef.
- K. Nagapudi, E. Y. Umanzor and C. Masui, Int. J. Pharm., 2017, 521(1–2), 337–345 CrossRef PubMed.
- M. E. Suarez-Almazor, E. Belseck, B. Shea, P. Tugwell and G. A. Wells, Cochrane Database of Systematic Reviews, 1998, Issue 2, Art. No.: CD000958 Search PubMed.
- D. Fletcher, S. Patel and K. Motaparthi, Cureus, 2023, 15(4), e37210 Search PubMed.
- A. J. Blake, X. Lin, M. Schröder, C. Wilson and R. X. Yuan, Acta Crystallogr., Sect. C: Cryst. Struct. Commun., 2004, 60, o226–o228 CrossRef PubMed.
- L. A. Filip, M. R. Caira, S. I. Fărcaş and M. T. Bojiţa, Acta Crystallogr., Sect. C: Cryst. Struct. Commun., 2001, 57, 435–436 CrossRef PubMed.
- E. Tedesco, D. Giron and S. Pfeffer, CrystEngComm, 2002, 4, 393–400 RSC.
- W. I. F. David and K. Shankland, Acta Crystallogr., Sect. A: Found. Crystallogr., 2008, 64, 52–64 CrossRef PubMed.
- M. Bala, M. K. Gautam and R. Chadha, Innovations Pharm. Pharmacother., 2017, 5(3), 154–158 Search PubMed.
- R. E. Elbakush, A Crystal Engineering Study of Selected Sulfa Drugs and Trimethoprim, University of Western Cape: South Africa, 2014 Search PubMed.
- S. Huang, V. K. R. Cheemarla, D. Tiana and S. E. Lawrence, Cryst. Growth Des., 2023, 23(8), 5446–5461 CrossRef.
- N. J. Babu, L. S. Reddy and A. Nangia, Mol. Pharmaceutics, 2007, 4(3), 417–434 CrossRef PubMed.
- J. A. McMahon, J. A. Bis, P. Vishweshwar, T. R. Shattock, O. L. McLaughlin and M. J. Zaworotko, Z. Kristallogr. – Cryst. Mater., 2005, 220(4), 340–350 CrossRef.
- F. H. Allen, W. D. Samuel Motherwell, P. R. Raithby, G. P. Shields and R. Taylor, New J. Chem., 1999, 23, 25–34 RSC.
- A. Alhadid, L. Kefalianakis, A. Wendler, S. Nasrallah, C. Jandl, S. M. Kronawitter, G. Kieslich and M. Minceva, Cryst. Growth Des., 2024, 24(11), 4770–4780 CrossRef.
- (a) CCDC 2417286: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2m4cz4; (b) CCDC 2417287: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2m4d06.
| This journal is © The Royal Society of Chemistry 2026 |