Open Access Article
Open Access ArticleTackling environmental and health issues via crystal engineering of metal complexes
Laura
Contini
 ,
Fabrizia
Grepioni
,
Fabrizia
Grepioni
 * and
Dario
Braga
* and
Dario
Braga

Dipartimento di Chimica “Giacomo Ciamician”, Università di Bologna, Via Gobetti 85, 41029, Bologna, Italy. E-mail: fabrizia.grepioni@unibo.it
First published on 12th February 2026
Abstract
This highlight reports results obtained by the group of Molecular Crystal Engineering at the University of Bologna by applying crystal engineering strategies to tackle two fundamental contemporary challenges of mankind, namely the antimicrobial resistance developed by pathogens as a consequence of misuse/overuse of antibiotics and, from a relatively distant but connected area, the need to inhibit urea degradation by soil enzymes to improve agricultural produce while decreasing environmental pollution. These two targets have been pursued by preparing and characterizing a series of co-crystals obtained by combining active organic molecules (antimicrobials, fertilizers, enzyme inhibitors) with coordination compounds of metals such as Zn, Cu, Ga, Ag, and Bi. Most compounds described in the following have been obtained by mechanochemically activated reactions between organic active molecules and coordination compounds, a method that also eliminates or reduces significantly the use of solvents. The characterization of the products has been carried out with a combination of solid-state techniques like X-ray diffraction, calorimetry, and thermogravimetry. The properties of all compounds have also been tested via collaborative efforts with expert research groups in Canada, USA, Germany and Italy to verify if the co-crystals obtained could, respectively, improve the activity of known organic antimicrobial agents or inhibit the activity of soil enzymes such as urease and ammonia monooxygenase responsible for urea degradation.
1 Introduction
Crystal engineering seeks to create materials suited to specific applications by assembling in the solid state, via supramolecular interactions, molecular or ionic building blocks possessing defined physico-chemical-biological attributes.1–3 Within this framework, the preparation of co-crystals—i.e., the solid-state association of two or more components, each possessing crystalline phases under standard conditions—represents a crystal-engineering synthetic approach, in which reactants are brought together to yield crystalline materials with their own distinct characteristics and properties.4–9 In such a way, co-crystallisation enables the incorporation, within a single crystalline phase, of an active ingredient, such as a pharmaceutical or a fertilizer, together with a chaperone or carrier, in order to enhance physico-chemical behaviour or impart new properties relative to those of the parent components.10–13 Consequently, when applied to materials destined for human and animal health or the food sector, co-crystal design has emerged as one of the most powerful synthetic strategies for addressing current challenges.4,7,11,13–16This highlight focuses on crystal-engineering approaches involving coordination compounds. Although crystal engineering originated within the organic chemistry field, it has since expanded across the full breadth of chemical science and beyond.17,18 The interplay between the physico-chemical features of the building blocks—whether neutral or charged, organic or inorganic—and their periodic arrangement in the crystal ultimately determines the collective properties of the resulting materials.19,20 The same reasoning applies to the synthesis of coordination compounds, coordination polymers, and metal–organic frameworks (MOFs).21–24 In coordination chemistry the possible combinations of ligand geometry and topology with metal-centre coordination environments are virtually limitless,25 as is their impact across numerous fields of application.26–37 The attribution of the 2025 Nobel Prize for Chemistry to R. Robson, S. Kitagawa and O. Yaghi for their work on coordination polymers and metal–organic frameworks is the most evident recognition of such impact.38
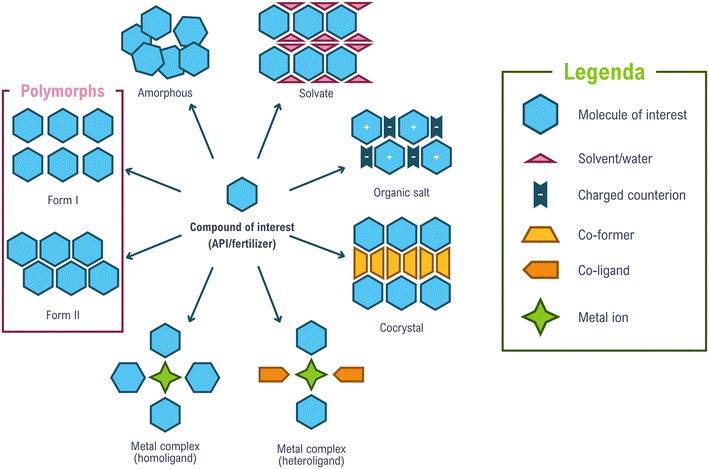
Here, we consider two specific domains in which crystal engineering has been applied to coordination chemistry. Before doing so, it is helpful to revisit the topological distinctions between molecular crystals and coordination polymers. Taking molecular crystals as a starting point (Fig. 1), an active molecule may crystallize as a single-entity phase, i.e., a crystal composed solely of the molecule of interest, or it may associate with a solvent to give a solvate, with a counterion to form a salt, or with a coformer (a substance solid under ambient conditions) to generate a co-crystal. The co-crystal may itself form a solvate or a salt, and these may in turn also be solvated.5,11,20,39 Such diversity defines the landscape of crystal forms that crystal engineering investigates. Each crystal form bears, within its supramolecular architecture, certain physico-chemical features of the active molecule, modulated by interactions with solvent, ions, and/or other molecules. These molecule-based crystal forms may find use across an extensive range of applications.
Within the context of this highlight, the term “molecule” is employed broadly, encompassing coordination compounds in which ligands—typically organic molecules—coordinate metal centres to yield species with distinct properties. However, we also include coordination polymers and networks, arising from through-space extension in one dimension (linear polymers), two dimensions (layered coordination frameworks), or three dimensions (metal–organic frameworks) of the structural nodes of the crystal architecture.40–42 These topologies may likewise afford solvates or capture/release guest molecules, thereby enabling a broad palette of properties and applications.43–46 A detailed treatment of these classes is beyond the scope of the present article.
In what follows, we focus on two contemporary challenges that can be addressed through crystal-engineering strategies applied to coordination compounds: antimicrobial resistance and the enhancement of fertilizer efficiency. Antimicrobial resistance (AMR) emerges when bacteria, viruses, fungi, and parasites cease to respond to antimicrobial agents, rendering treatments ineffective and infections increasingly difficult—or impossible—to cure.47,48 Loss of fertilizer efficiency, by contrast, primarily results from the rapid enzymatic hydrolysis of nitrogen-based fertilizers such as urea, catalysed by soil enzymes and microorganisms.49–54 These transformations accelerate nitrogen volatilization as ammonia and nitrous oxides, contributing to air pollution and decreasing the nitrogen available to plants.
Given the global relevance of these two issues, it is unsurprising that numerous research groups – clearly reflected in the references cited herein – are actively pursuing new approaches to safeguard human and animal health and to ensure sustainable agricultural production for a growing world population. Before presenting our recent findings, we briefly summarize the methods used for preparing, characterizing, and assessing the products of co-crystal syntheses.
2 Methods of preparation and characterization
The products of a crystal engineering synthesis are, generally, in the form of crystalline material. In addition to traditional solution crystallization methods, mechanochemical mixing of components has emerged as one of the most innovative approaches for preparing both molecular co-crystals as well as coordination compounds.55–58 These solvent-free or solvent-minimized techniques represent a greener and more cost-effective approach, enabling the synthesis of co-crystals and coordination compounds without having to worry about differences in solubilities of the components.59–61 Mastering the ability to produce co-crystals and coordination compounds without resorting to solution crystallization has been a significant step forward in the field,62 allowing molecular solids to be combined without concerns about solubility differences and enables the development of new solid-state reaction pathways.63The mechanochemical approach not only permits the preparation of otherwise not accessible organic and metal organic co-crystals,55,62,64 but also enables the study of the effects of grinding and comminution on crystal polymorphism and the uptake or release of water in the formation of hydrates. Importantly, reaction outcomes can be controlled by adjusting milling conditions, including solvent quantity, milling duration, jar material in ball milling, and grinding media.63,65–67
Since crystalline materials are primarily involved, diffraction techniques play a dominant role in the investigation of a mechanochemical preparation. X-ray powder diffraction is the preferred method for identifying and characterizing products, whether in pure form or mixed with unreacted starting materials. In cases where powders can be recrystallized into crystals of adequate size, one can resort to single-crystal X-ray diffraction for a thorough analysis of the crystal structure. Even when single-crystals are not available, advancements in structure determination from powder diffraction68 and electron diffraction,69 along with complementary techniques such as solid-state NMR spectroscopy,70,71 and in situ methods for monitoring mechanochemical reactions,72–74 have significantly accelerated the progress in crystal engineering. X-ray diffraction methods need to be accompanied by a variety of solid-state analysis techniques, to fully understand the behaviour of the crystalline material in terms of thermal stability, gain–loss of solvent/humidity, phase transitions, etc., need to be assessed via differential scanning calorimetry (DSC), thermogravimetric analysis (TGA), often accompanied by dynamic vapor sorption (DVS), and hot-stage microscopy (HSM). Spectroscopic methods, such as Fourier-transform infrared (FTIR), Raman spectroscopy, and solid-state NMR are also often indispensable to fully characterize the crystal engineering product.
3 Response to antimicrobial resistance
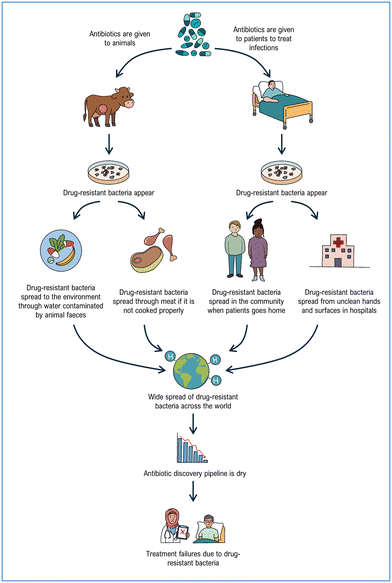
Antimicrobial resistance (AMR) refers to the condition in which bacteria, viruses, parasites, and fungi cease to respond to therapeutic agents.75–78 The WHO now identifies AMR as a “silent pandemic”, owing to the fact that pathogens are acquiring resistance to existing antimicrobials at a pace that far outstrips the introduction of new drugs to the market.79–81 This situation has become an urgent global concern that demands a coordinated, interdisciplinary response encompassing reduced unnecessary utilization of antibiotics, improved infection prevention and control, and continuous surveillance of AMR trends (Fig. 2).82–84 Meanwhile, the clinical development pipeline remains inadequate to counter the escalating challenge posed by AMR.85 Current strategies involve either the development of new, non-traditional antimicrobial agents, such as monoclonal antibodies,86 bacteriophages,87,88 and antimicrobial peptides,89 or the revitalization of “exhausted” drugs. Clearly, the latter approach is well suited for crystal engineering, as evidenced by the substantial number of research groups engaged in designing and evaluating co-crystals of active antimicrobials with appropriately selected coformers.11,13,90,91 Modifying the properties of existing drugs by improving solubility, permeability, dissolution rate, and related parameters, or generating novel co-drugs via pharmaceutical synthon strategies continues to motivate both academic and industrial efforts.92 The growing portfolio of patents on pharmaceutical co-crystals, several of which have reached the market, further attests to the impact of crystal engineering in this area.93Extending co-crystallization strategies to metal complexes offers an additional path to expand the toolbox against AMR. Metals have served as antimicrobial agents long before the discovery of antibiotics or their widespread use in human, animal, and plant health.94–97 Indeed, crystal-engineering methodologies employing metal complexes and coordination networks with antimicrobial properties have gained considerable traction as new means to combat AMR, including the sterilization of surfaces and medical devices compromised by biofilm formation.98–103
This approach capitalizes on the fact that traditional organic antibiotics typically act via a single mechanism, thereby facilitating the emergence of resistance. Coordination to a metal centre can enhance molecular reactivity, as redox behaviour, reactive oxygen species (ROS) generation, and interference with quorum sensing or biofilm formation among the key effects.94,96,104 Metal ions may also modulate ligand uptake, solubility, or stability. The overarching objective is to achieve multimodal antibacterial action, whereby the drug–metal complex retains the parent antibiotic mechanism while acquiring additional modes of activity conferred by the metal.105–108
Over the last decade, we have applied co-crystallization—via solution and mechanochemical routes—to generate new compounds combining established APIs with metal complexes. Table 1 lists the compounds that were synthesized, isolated, fully characterized, and evaluated for antimicrobial efficacy.
| Active molecule/ligand | Metal salt/complex, co-crystal | Reference |
|---|---|---|
| Proflavinium cation (HPF+) | ZnCl3(HPF) | 109, 110 |
| [HPF]2[ZnCl4]·H2O | 109, 110 | |
| [HPF]3[Ga(ox)3]·4H2O | 111 | |
| Proflavine (PF) | PF·CuCl | 112 |
| PF·AgNO3 | 112 | |
| Oxalate anion (ox−) | K3[Ga(ox)3]·3H2O | 111 |
| K4[Ga2(ox)4(μ-OH)2]·2H2O | 111 | |
| Ciprofloxacin (CIP) | CIP·QUE | 113 |
| CIP·CAR4 | 114 | |
| CIP·THY2 | 114 | |
| Levofloxacin (LEV) | LEV2·HES | 113 |
| LEV·MYR | 113 | |
| LEV·QUE | 113 | |
| Cephalexin (CPX) | CPX·THY·2.5H2O | 115 |
| Cephradine (CFD) | CFD·THY·2.5H2O | 115 |
| Cefaclor (CFC) | CFC·THY·4H2O | 115 |
| Kojic acid (HKA) and kojiate anion (KA−) | [Ag(HKA)(NO3)]·H2O | 116 |
| [Cu(KA)2] | 116 | |
| [Zn(KA)2] | 116 | |
| [Ga(KA)2(OH2)2][NO3]·H2O | 116 | |
| Ethylenediaminetetraacetic acid (HEDTA3−) | [Bi(HEDTA)]·2H2O | 117 |
| [Bi(HEDTA)] | 117 | |
| [Bi2(HEDTA)2(μ-DL-His)2]·6H2O | 117 | |
| [Bi(HEDTA)]·Cyt·2H2O | 117 | |
| Metronidazole | [Zn(Sal)2(MET)2] | 118 |
| [Cu(Sal)2(MET)2(H2O)] | 118 | |
| [Ag(Sal)(MET)] | 118 | |
| Aminocinnamic acids (n-AC) | [Zn(4-AC)2(H2O)2] | 119 |
| [Zn(4-AC)2]·H2O | 119 | |
| [Zn(3-AC)2]·2H2O | 119 | |
| [Zn(3-AC)(4-AC)] | 119 | |
| L-/DL-Arginine (Arg) | [Cu·L-Arg·(NO3)2·H2O] | 120 |
| [Cu·DL-Arg·(NO3)2·H2O] | 120 | |
| [Ag·L-Arg·NO3·0.5H2O] | 120 | |
| [Ag·DL-Arg·NO3·0.5H2O] | 120 | |
| L-/DL-Histidine (His) | [Cu·L-His·(NO3)2·H2O] | 120 |
| [Cu·DL-His·(NO3)2·H2O] | 120 | |
| [Ag·L-His·NO3] | 120 | |
| [Ag·DL-His·NO3] | 120 | |
| [Ag·(L-His)2·NO3·0.5H2O] | 120 | |
| [Ag·(DL-His)2·NO3·0.5H2O] | 120 |
The well-known disinfectant and bacteriostatic agent proflavine, a quaternary cation compound (QCC)121 also known as acridine-3,6-diamine,122 has served as a starting point for the preparation of a series of co-crystals obtained by reacting proflavine with salts and complexes of copper(I), zinc(II), silver(I), and gallium(III).109–111,123 These materials were obtained predominantly by mechanochemical grinding or by slurry methods, with the latter yielding higher-purity samples, subsequently used for antimicrobial testing against Pseudomonas aeruginosa ATCC27853, Staphylococcus aureus ATCC25923, and Escherichia coli ATCC25922.
The structure of the copper(I) chloride co-crystal proflavine·CuCl, shown in Fig. 3, consists of a (1D) polymeric chain of CuCl units accompanied by neutral proflavine molecules, arranged in a herringbone pattern.123
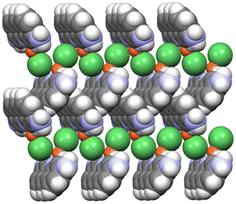 | ||
| Fig. 3 (CuCl⋯CuCl⋯)n chain motifs and herringbone packing of proflavine molecules in the proflavine·CuCl co-crystal. Reproduced from ref. 123. | ||
Antimicrobial assays performed on the proflavine·AgNO3 and proflavine·CuCl materials against S. aureus, P. aeruginosa, and E. coli reveal that both complexes outperform proflavine and the individual salts (Fig. 4). These findings support the broader concept that hybrid materials generated from inorganic salts and active molecules can yield new functional solids suitable, for instance, for antimicrobial surface coatings.
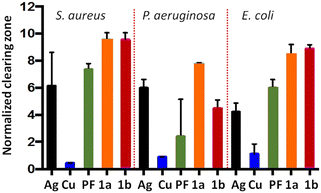 | ||
| Fig. 4 Performance of PF-based materials: normalized growth-inhibition zones from compound-impregnated disks [Ag = AgNO3, PF = proflavine, Cu = CuCl, 1a = PF·CuCl, 1b = PF·AgNO3]. Reproduced from ref. 123. | ||
Using the same rationale, analogous ZnCl2-based derivatives were synthesized.109 The product composition varied depending on the mechanochemical or solution conditions and the Zn-to-proflavine stoichiometric ratios. Two compounds were isolated and structurally characterized: ZnCl3(HPF) and the monohydrate [HPF]2[ZnCl4]·H2O, both containing proflavine in its protonated proflavinium form (HPF)+, in contrast to the neutral proflavine found in the silver and copper systems. Their structures, compared in Fig. 5, display notable stacking interactions between the proflavinium cations—an interaction likewise predominant in neutral proflavine and in the salt [HPF]Cl·2H2O.
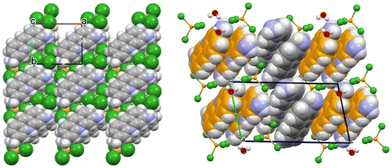 | ||
| Fig. 5 (Left) ab-Plane projection of crystalline ZnCl3(HPF), highlighting molecular stacks aligned along the c-axis. (Right) b-Axis projection of [HPF]2[ZnCl4]·H2O, illustrating the herringbone arrangement of HPF+ cation pairs. Reproduced from ref. 109. | ||
When evaluated against pathogenic indicator strains, these compounds exhibited a 50–125% enhancement in antimicrobial activity relative to AgNO3. Although the improvement was somewhat less pronounced, when compared to the simple physical mixture of the constituents, it remained significant (Fig. 6). Overall, association of proflavine in its protonated form with zinc enhances its antimicrobial effectiveness.
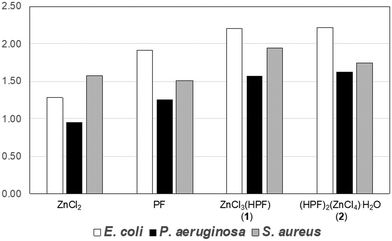 | ||
| Fig. 6 Relative antimicrobial activities of proflavine, ZnCl3(HPF), and (HPF)2(ZnCl4)·H2O normalized to the activity of AgNO3 (value of 1.00 corresponds to silver nitrate efficacy). Reproduced from ref. 109. | ||
Proflavine-based co-crystals have also been prepared using the gallium oxalate anion [Ga(ox)3]3− as a preassembled building block, giving the molecular salt [HPF]3[Ga(ox)3]·4H2O (Fig. 7).110,111
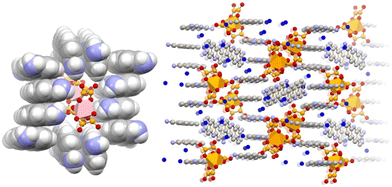 | ||
| Fig. 7 π-Stacking of [HPF]+ cations in crystalline [HPF]3[Ga(ox)3]·4H2O (left), and encapsulation of two [Ga(ox)3]3− anions within a proflavinium-cation envelope (right). Reproduced from ref. 111. | ||
The proflavinium cations envelope the gallium oxalate anions, forming stacking arrangements akin to those observed in the neutral co-crystals described earlier. Disk diffusion assays demonstrate that this gallium-containing material also displays significant antimicrobial activity. Notably, whereas [HPF]3[Ga(ox)3]·4H2O is active against all three tested strains, the precursor salt K3[Ga(ox)3] exhibits striking selectivity for P. aeruginosa, with minimal activity towards the remaining species: a remarkable behaviour, which requires further microbiological investigation.
In summary, co-crystallization of a known QCC antibacterial molecule such as proflavine with metal salts (e.g., CuCl, CuCl2, AgNO3) or preformed complexes like [Ga(ox)3]3−, via mechanochemical or slurry methods, offers a sustainable, cost-effective, and environmentally benign strategy for generating novel materials with improved antibacterial properties. All crystalline products examined thus far, namely PF·CuCl, PF·AgNO3, ZnCl3(HPF), [HPF]2[ZnCl4]·H2O, and [HPF]3[Ga(ox)3]·4H2O, show superior antibacterial performance relative to proflavine or the corresponding metal salts alone, despite containing a lower molar fraction of proflavine.
In another study, mechanochemical and solution-based approaches were also applied to generate co-crystals between the amino acids arginine or histidine (in both L- and DL- forms) and the salts Cu(NO3)2 and AgNO3. The resulting coordination polymers, [Cu·L-Arg·(NO3)2·H2O]CP, [Cu·DL-Arg·(NO3)2·H2O]CP, [Cu·L-His·(NO3)2·H2O]CP, and [Cu·DL-His·(NO3)2·H2O]CP, shown in Fig. 8, were isolated and characterized.120 Their antimicrobial profiles against P. aeruginosa, E. coli, and S. aureus indicate that the chirality of the amino acid has little, if any, influence, while the coordination polymers exhibit antimicrobial activity comparable to, or in some cases greater than, that of the metal salts.
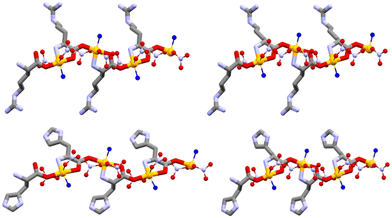 | ||
| Fig. 8 Polymeric chain structures in L-Arg·Cu (top left), DL-Arg·Cu (top right), L-His·Cu form II (bottom left), and DL-His·Cu (bottom right); Owater in blue and Cu in orange. Adapted from ref. 120. | ||
The antimicrobial activity of the AgNO3 and Cu(NO3)2 coordination polymers with the amino acids was compared to that of the corresponding solid mixtures of amino acids and salts in identical stoichiometric ratios (Fig. 9). The results show no significant difference between the coordination polymers and the simple mixtures; however, the amino acid/silver nitrate combination appears especially effective against P. aeruginosa.
 | ||
Fig. 9 Normalized antimicrobial activity from disk-diffusion assays on lysogeny agar for (A) coordination polymers formed between metal salts and amino acids and (B) physical mixtures of the corresponding metal salts and amino acids in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry. Activity is normalized to silver nitrate, with values above 1.05 indicating higher antimicrobial efficacy. Reproduced from ref. 120. 1 stoichiometry. Activity is normalized to silver nitrate, with values above 1.05 indicating higher antimicrobial efficacy. Reproduced from ref. 120. | ||
Encouraged by these findings, we recently applied the same methodology to metronidazole (MET),118 a nitroimidazole antibiotic long used against a broad spectrum of bacteria. Increasing resistance to MET shown by several pathogens underscores the urgency of developing improved formulations.124,125 Beyond metal coordination, we also sought to exploit potential synergistic effects with MET of the salicylate anion, itself known to possess antimicrobial activity.126 Mechanochemical and slurry methods enabled the preparation of the complexes [Zn(Sal)2(MET)2] form I, [Zn(Sal)2(MET)2] form II, [Cu(Sal)2(MET)2(H2O)] form I, [Cu(Sal)2(MET)2(H2O)] form II, and [Ag(Sal)(MET)], which were fully characterized.118Fig. 10 shows the different orientation of the MET ligands in the two polymorphic forms of the Zn and Cu complexes.
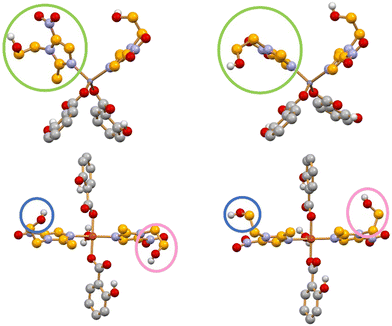 | ||
| Fig. 10 Distinct orientations of MET ligands around N–Zn bonds in [Zn(Sal)2(MET)2] forms I (top left) and II (top right), and around N–Cu bonds in [Cu(Sal)2(MET)2(H2O)] forms I (bottom left) and II (bottom right) [CMET in orange, CSal in grey; HCH atoms omitted for clarity]. Adapted from ref. 118. | ||
Given the higher stability of the form II polymorphs, these, together with [Ag(Sal)(MET)], were evaluated against four bacterial strains—Escherichia coli, Pseudomonas aeruginosa, Staphylococcus aureus, and Staphylococcus epidermidis—using minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) assays.118 Results are reported in Fig. 11.
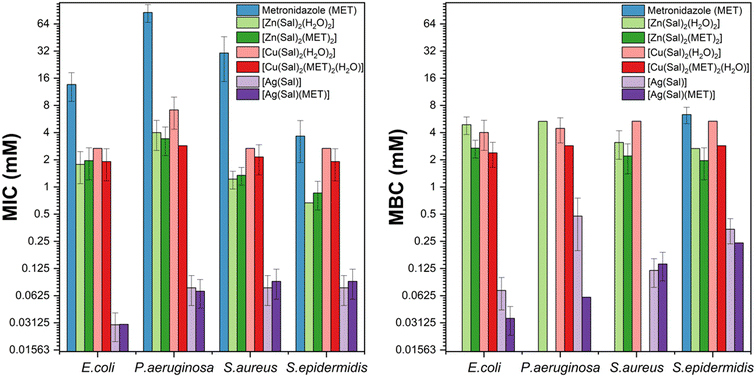 | ||
| Fig. 11 (Left) Minimum inhibitory concentration (MIC) and (right) minimum bactericidal concentration (MBC) for metronidazole metal complexes against E. coli, P. aeruginosa, S. aureus, and S. epidermidis. The y-axis is plotted on a log2 scale for better clarity. Shorter bars denote higher antimicrobial activity. Reproduced from ref. 118. | ||
The antimicrobial tests clearly show that combining metronidazole with salicylic acid and coordinating the resulting formulation to Zn(II), Cu(II), or Ag(I) significantly enhances activity against both Gram-positive and Gram-negative aerobic pathogens, with the silver complex displaying the strongest overall effect. These findings further support the central hypothesis that antimicrobial performance can be enhanced through coordination with active metal atoms.
4 Organic–inorganic co-crystals for soil enzyme inhibition
In this section, we outline our crystal-engineering strategy using inorganic and coordination compounds to address a major agro-environmental and economic challenge within the global nitrogen (N) cycle. There is a pressing demand for sustainable soil fertilization and improved crop yields with methods able to simultaneously reduce nitrogen losses from the decomposition of urea-based fertilizers.Urea remains the dominant nitrogen fertilizer worldwide, representing roughly 60% of total nitrogen fertilizer use.127–130 Industrially, urea is synthesized mainly from ammonia, produced via the energy-intensive Haber–Bosch process using nitrogen, hydrogen and carbon dioxide.131,132 However, a considerable fraction of the nitrogen supplied as urea is lost through volatilization, denitrification, and leaching, as depicted in Fig. 12.49–51,133,134 Once introduced into soil, urea undergoes rapid enzymatic hydrolysis by urease, a nickel-dependent enzyme. If not promptly absorbed, urease can convert urea into hydrogen carbonate (HCO3−) and ammonium (NH4+) at a rate up to 1015 times faster than non-enzymatic hydrolysis.135–137 Although NH4+ is a plant nutrient, the accompanying rise in pH favours the formation of gaseous ammonia, which is readily emitted into the atmosphere.138,139
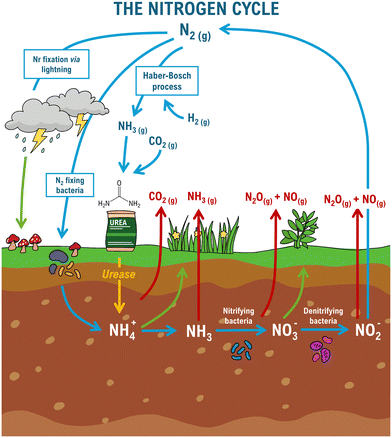 | ||
| Fig. 12 Diagram of the nitrogen cycle (blue arrows). Urea hydrolysis catalysed by urease is emphasized by a yellow arrow. Loss pathways are depicted in red, and uptake pathways in green. | ||
Urease is not the sole enzyme governing nitrogen transformations in soil. Ammonia monooxygenase (AMO), a copper-dependent enzyme present in ammonia-oxidizing microorganisms, catalyses the oxidation of ammonia to hydroxylamine (NH2OH), which is subsequently oxidized to nitrite (NO2−) and nitrate (NO3−).140 Nitrites and nitrates further serve as precursors to nitric oxide (NO) and nitrous oxide (N2O), both of which contribute to greenhouse gas emissions.141,142
Due to this cascade of reactions, nearly half of the nitrogen supplied as urea is ultimately lost, creating both economic losses and environmental damage.143,144 Two principal strategies are currently employed to mitigate these losses: (i) reducing the solubility and dissolution rate of urea, often through coating or encapsulation technologies,145,146 and (ii) inhibiting urea degradation by soil enzymes using urease inhibitors in combination with urea-based fertilizers.147,148
Several groups have investigated co-crystallization methods to advance these goals.149–152 Co-crystallization can not only modify the solubility of active ingredients, such as urea, but also allow incorporation of enzyme inhibitors as coformers within the fertilizer matrix. Recently, urea-based ionic co-crystals with inorganic compounds have shown promise in reducing urea decomposition and thus reducing ammonia release. Examples include urea·MgSO4 co-crystals and co-crystals with Ca2+ salts obtained mechanochemically from suitable minerals.153 Beyond co-crystals, inorganic salts such as Ca(NH4)2(HPO4)2·H2O and Mg(NH4)2(HPO4)2·4H2O, along with their struvite analogues Ca(NH4)(PO4)·H2O and Mg(NH4)(PO4)·6H2O, have been employed for similar applications.153,154
The co-crystallization strategy has thus been applied in the development of organic–inorganic co-crystals, designed to enhance the physicochemical properties of urea by lowering its solubility/dissolution rate, while simultaneously providing inhibition of urease and/or AMO, all while supplying nutrients to soil. Table 2 lists the compounds synthesized, isolated, characterized, and assessed for enzyme inhibition performance.
| Active molecule/ligand | Metal salt/complex, co-crystal | Reference |
|---|---|---|
| Urea (U) and thiourea (T) | U·ZnCl2·KCl | 155 |
| ZnCl2·T | 156 | |
| ZnCl2·T·U | 156 | |
| U·catechol | 157 | |
| U·L-proline·H2O | 158 | |
| U2·L-proline | 158 | |
| U·L-proline·H2O | 158 | |
| Dicyandiamide (DCD) | [Cu(DCD)2(OH2)2(NO3)2] | 159 |
| [Cu(DCD)2(OH2)Cl2]·H2O | 159 | |
| [Cu(DCD)2(OH2)2][NO3]2·2H2O | 159 | |
| [Cu(DCD)2(OH2)Cl2] | 159 | |
| Nitrapyrin (NP) | β-CD·NP | 160 |
| Bentazon (HBtz) | Na·Btz·1.75H2O | 161 |
| [β-CD·HBtz]·6H2O | 161 | |
| [γ-CD·HBtz]·8H2O | 161 | |
| Thiurams: thiram (TMTD), disulfiram (TETD), tetraisopropylthiuram disulfide (TIPTD) | β-CD·TMTD | 162 |
| (β-CD)2·TETD | 162 | |
| (β-CD)2·TIPTD | 162 | |
| Ethylenediaminetetraacetic acid (HEDTA3−) | [Bi(HEDTA)]·2H2O | 117 |
| [Bi(HEDTA)] | 117 | |
| [Bi2(HEDTA)2(μ-DL-His)2]·6H2O | 117 | |
| [Bi(HEDTA)]·Cyt·2H2O | 117 |
When urea is co-crystallized with the inorganic metal salts KCl and ZnCl2, the resulting compound forms quantitatively via a straightforward, solvent-free, and scalable procedure.155 Depending on the preparation method (Fig. 13a), the urea·ZnCl2·KCl system can be isolated in two distinct polymorphic forms.
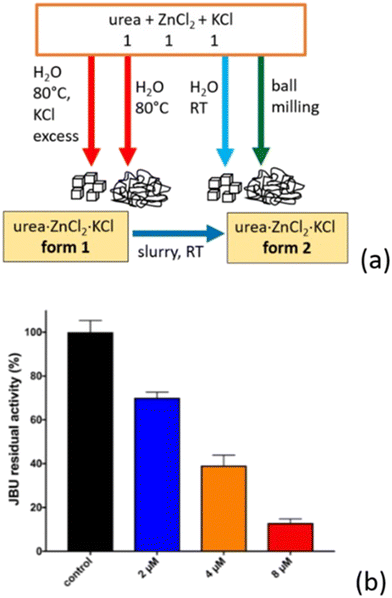 | ||
Fig. 13 (a) Polymorphic forms of urea·Zn·KCl obtained by reacting urea, ZnCl2, and KCl in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometric ratio. (b) Residual jack bean urease (JBU) activity relative to 100% control (black bar) in the presence of increasing concentrations of urea·Zn·KCl form 2 at pH 7.5. Blue, orange, and red bars correspond to 2, 4, and 8 μM inhibitor concentration, respectively. Reproduced from ref. 155. 1 stoichiometric ratio. (b) Residual jack bean urease (JBU) activity relative to 100% control (black bar) in the presence of increasing concentrations of urea·Zn·KCl form 2 at pH 7.5. Blue, orange, and red bars correspond to 2, 4, and 8 μM inhibitor concentration, respectively. Reproduced from ref. 155. | ||
The polymorph obtained from aqueous solution at 80 °C is metastable, and converts to the stable form upon slurry aging at room temperature (RT), the latter can also be obtained through ball milling or crystallization at RT.163 Because zinc(II) is a known urease inhibitor,164 the stable polymorph of urea·ZnCl2·KCl was tested against urease across increasing concentrations (Fig. 13b), revealing strong inhibition. In addition to effectively modulating urease activity, the co-crystal delivers the nutrient KCl alongside urea.
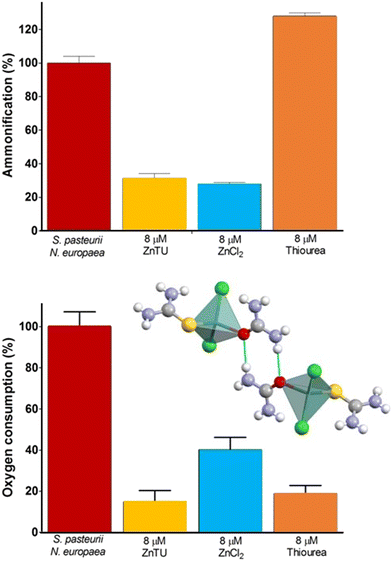
Along the same line of research, it has been shown that co-crystallization of urea and thiourea, a known inhibitor of AMO-mediated ammonia oxidation,165 with ZnCl2 yields the mixed adduct [Zn(thiourea)(urea)Cl2].156 This compound, besides supplying urea fertilizer, functions as a dual inhibitor. Experiments on bacterial cultures of S. pasteurii and N. europaea, as well as tandem cultures, showed selective suppression of ammonification and oxygen consumption attributable to Zn2+ and thiourea, respectively. By simultaneously providing nitrogen and inhibiting two key enzymes of the N cycle, [Zn(thiourea)(urea)Cl2] can improve the efficiency of nitrogen fertilization (Fig. 14).
A further illustration of how crystal-engineering strategies can be applied in agrochemical research is the in situ monitoring of the mechanochemical reaction between the AMO inhibitor dicyandiamide and copper(II) salts [CuX2, X = Cl−, NO3−], themselves urease inhibitors.159 A deeper mechanistic understanding of the reaction conditions enabled targeted synthesis of the desired adducts and provided a framework for exploring new agrochemically relevant compounds. A related approach was discussed by Baltrusaitis, Friščić et al.154
As part of our efforts to identify urease inhibitors, we also examined Bi(III) complexes.117 Although bismuth has no established biological function, Bi(III) compounds have long been used therapeutically, particularly in combination treatments for Helicobacter pylori infection.166–168 Despite their clinical efficacy, the molecular basis for the action of bismuth-based drugs remains only partially defined.169 Several Bi(III) complexes, including [Bi(HEDTA)]·2H2O (HEDTA = N-(2-hydroxyethyl)ethylenediaminetriacetic acid), [Bi(Cys)3], and ranitidine bismuth citrate, have been reported as urease inhibitors, and this activity is often cited as central to their antibacterial effects.170 However, the use of Bi(III) compounds is limited by their very low solubility in water.
To enhance the solubility of Bi(III) complexes, we selected the reported urease inhibitor [Bi(HEDTA)]·2H2O and attempted co-crystallization with (i) racemic DL-histidine, affording the conglomerate [Bi2(HEDTA)2(μ-D-His)2]·6H2O + [Bi2(HEDTA)2(μ-L-His)2]·6H2O; (ii) enantiopure L-histidine, yielding [Bi2(HEDTA)2(μ-L-His)2]·6H2O; and (iii) cytosine, forming [Bi(HEDTA)]·Cyt·2H2O (Fig. 15).117 Notably, all co-crystals improved aqueous solubility of the Bi(III) complex, ranging from 6.8 mg mL−1 for [Bi(HEDTA)]·2H2O to 36 mg mL−1 for [Bi2(HEDTA)2(μ-L-His)2]·6H2O, thus fulfilling one objective of the study.
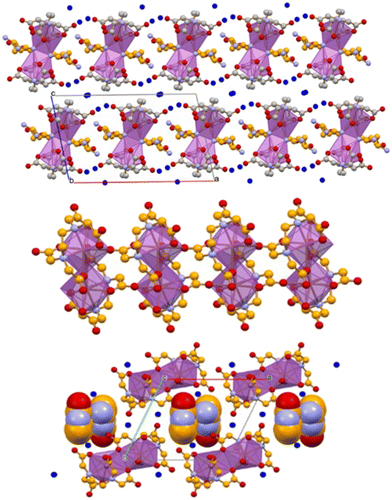 | ||
| Fig. 15 (Top) b-Axis projection of the packing in [Bi2(HEDTA)2(μ-L-His)2]·6H2O. (Middle) One-dimensional ribbons of [Bi(HEDTA)] units running parallel to the c-axis in [Bi(HEDTA)]·Cyt·2H2O. (Bottom) Side-view down the c-axis illustrating how cytosine (uppercase labels) and water molecules occupy the cavities between adjacent ribbons [bismuth coordination polyhedra in purple; histidine and cytosine C atoms in orange; water O atoms in blue; H atoms omitted]. Reproduced from ref. 117. | ||
Unexpectedly, when the co-crystals and the parent complex [Bi(HEDTA)]·2H2O were tested for urease inhibition, no activity was observed, neither in vitro against Canavalia ensiformis urease, nor in vivo against H. pylori cultures. Thus, while co-crystallization successfully increased solubility, it also raised doubts regarding the actual relevance of Bi(III) compounds in inhibiting urease in agro-environmental or clinical contexts. The inhibitory effect of Bi(III) on H. pylori growth appears to arise from interference with metabolic pathways other than urease.
5 Conclusions
In this highlight article we have collected the most significant results obtained by the Molecular Crystal Engineering group at the University of Bologna by applying crystal engineering synthetic strategies to prepare (i) new materials to be used to contrast the increasing antimicrobial resistance shown by pathogens and (ii) new materials that could be used to inhibit enzyme (urease, AMO) activity in the soil, at the same time providing the fertilizer (urea) together with nutrients or inhibitors. In order to do so, we have prepared a number of coordination compounds by applying co-crystallization strategies.The method of choice for the preparation of the co-crystals and of the coordination compounds has been the mechanochemical reaction between organic active molecules and coordination compounds in this also eliminating or reducing significantly the use of solvents.
The characterization of the products has been carried out thanks to a combination of solid-state techniques, from single-crystal to powder X-ray diffraction to differential scanning calorimetry, thermogravimetry, etc., while the efficacy of the co-crystalline products in tackling the two aforementioned issues has been tested by expert groups of researchers with complementary indispensable expertise, as well evidenced by the joint publications quoted throughout this highlight article and acknowledged below.
Author contributions
DB drafted the manuscript. All authors contributed to the writing, discussed each section and commented on the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
This is a highlight article, without new data input herein. All data can be sourced from the referenced articles and their relative supporting information (SI).Acknowledgements
Financial support from MUR, project “NICE – Nature Inspired Crystal Engineering” (PRIN2020) and the University of Bologna (RFO) is acknowledged. The PNRR DM 351/2022 PhD project (LC) is also acknowledged. All the results obtained by the Bologna's group and presented in this highlight article have been possible only thanks to the close synergistic interactions with researchers with the indispensable complementary expertise required to characterize and/or assess the properties of the various classes of compounds. We gratefully acknowledge the collaborations with the groups of Ray J. Turner (University of Calgary), Stefano L. Ciurli, Luca Mazzei and Davide Roncarati (University of Bologna), Jonas Baltrusaitis (University of Leigh), Franziska Emmerling (BAM, Berlin), Roberto Gobetto and Michele Chierotti (University of Torino). This Highlight is dedicated to Professor Resnati, celebrating a career in fluorine and noncovalent chemistry on the occasion of his 70th birthday.References
- G. R. Desiraju, Crystal Engineering: The Design of Organic Solids, Elsevier, 1989 Search PubMed.
- G. R. Desiraju, Angew. Chem., Int. Ed., 2007, 46, 8342–8356 CrossRef CAS PubMed.
- D. Braga, Chem. Commun., 2003, 2751–2754 RSC.
- D. J. Berry and J. W. Steed, Adv. Drug Delivery Rev., 2017, 117, 3–24 Search PubMed.
- D. Braga, F. Grepioni, L. Maini and M. Polito, in Molecular Networks, ed. M. W. Hosseini, Springer, Berlin, Heidelberg, 2009, pp. 87–95 Search PubMed.
- C. B. Aakeröy and D. J. Salmon, CrystEngComm, 2005, 7, 439–448 RSC.
- F. Grepioni, L. Casali, C. Fiore, L. Mazzei, R. Sun, O. Shemchuk and D. Braga, Dalton Trans., 2022, 51, 7390–7400 RSC.
- S. Aitipamula, R. Banerjee, A. K. Bansal, K. Biradha, M. L. Cheney, A. R. Choudhury, G. R. Desiraju, A. G. Dikundwar, R. Dubey, N. Duggirala, P. P. Ghogale, S. Ghosh, P. K. Goswami, N. R. Goud, R. R. K. R. Jetti, P. Karpinski, P. Kaushik, D. Kumar, V. Kumar, B. Moulton, A. Mukherjee, G. Mukherjee, A. S. Myerson, V. Puri, A. Ramanan, T. Rajamannar, C. M. Reddy, N. Rodriguez-Hornedo, R. D. Rogers, T. N. G. Row, P. Sanphui, N. Shan, G. Shete, A. Singh, C. C. Sun, J. A. Swift, R. Thaimattam, T. S. Thakur, R. Kumar Thaper, S. P. Thomas, S. Tothadi, V. R. Vangala, N. Variankaval, P. Vishweshwar, D. R. Weyna and M. J. Zaworotko, Cryst. Growth Des., 2012, 12, 2147–2152 CrossRef CAS.
- E. R. T. Tiekink, J. Vittal and M. Zaworotko, Organic Crystal Engineering: Frontiers in Crystal Engineering, John Wiley & Sons, 2010 Search PubMed.
- M. Singh, M. P. Sawarkar, M. R. Dhondale, D. R. Serrano, A. K. Agrawal and D. Kumar, Cryst. Growth Des., 2025, 25, 7852–7868 CrossRef CAS.
- D. Braga, Chem. Commun., 2023, 59, 14052–14062 RSC.
- Ö. Almarsson and M. J. Zaworotko, Chem. Commun., 2004, 1889–1896 Search PubMed.
- G. Bolla, B. Sarma and A. K. Nangia, Chem. Rev., 2022, 122, 11514–11603 CrossRef CAS PubMed.
- J. W. Steed, Trends Pharmacol. Sci., 2013, 34, 185–193 CrossRef CAS PubMed.
- M. Karimi-Jafari, L. Padrela, G. M. Walker and D. M. Croker, Cryst. Growth Des., 2018, 18, 6370–6387 Search PubMed.
- J. L. Dias, M. Lanza and S. R. S. Ferreira, Trends Food Sci. Technol., 2021, 110, 13–27 CrossRef CAS.
- A. K. Nangia and G. R. Desiraju, Angew. Chem., Int. Ed., 2019, 58, 4100–4107 CrossRef CAS PubMed.
- J. H. Williams, Crystal Engineering: How Molecules Build Solids, Morgan & Claypool Publishers, 2017 Search PubMed.
- Z. Hassan, Y. Matt, S. Begum, M. Tsotsalas and S. Bräse, Adv. Funct. Mater., 2020, 30, 1907625 CrossRef CAS.
- G. R. Desiraju, J. J. Vittal and A. Ramanan, Crystal engineering: a textbook, World Scientific, Singapore, 2011 Search PubMed.
- R.-B. Lin, Z. Zhang and B. Chen, Acc. Chem. Res., 2021, 54, 3362–3376 CrossRef CAS PubMed.
- G. Cai, P. Yan, L. Zhang, H.-C. Zhou and H.-L. Jiang, Chem. Rev., 2021, 121, 12278–12326 Search PubMed.
- D. Braga, F. Grepioni and G. R. Desiraju, Chem. Rev., 1998, 98, 1375–1406 Search PubMed.
- L. Brammer, Chem. Soc. Rev., 2004, 33, 476–489 RSC.
- Coordination Chemistry: Molecular Science of Organic–Inorganic Complexes, ed. T. Tanase and Y. Ishii, Royal Society of Chemistry, 2024 Search PubMed.
- V. F. Yusuf, N. I. Malek and S. K. Kailasa, ACS Omega, 2022, 7, 44507–44531 Search PubMed.
- X. Zhang, Z. Chen, X. Liu, S. L. Hanna, X. Wang, R. Taheri-Ledari, A. Maleki, P. Li and O. K. Farha, Chem. Soc. Rev., 2020, 49, 7406–7427 RSC.
- K. K.-W. Lo, A. W.-T. Choi and W. H.-T. Law, Dalton Trans., 2012, 41, 6021–6047 RSC.
- L. C.-C. Lee and K. K.-W. Lo, Chem. Rev., 2024, 124, 8825–9014 CrossRef CAS PubMed.
- X. Zhang, Z. Chi, Y. Zhang, S. Liu and J. Xu, J. Mater. Chem. C, 2013, 1, 3376–3390 RSC.
- L. S. Xie, G. Skorupskii and M. Dincă, Chem. Rev., 2020, 120, 8536–8580 CrossRef CAS PubMed.
- H. Liu, Y. Wang, Z. Qin, D. Liu, H. Xu, H. Dong and W. Hu, J. Phys. Chem. Lett., 2021, 12, 1612–1630 CrossRef CAS PubMed.
- A. Gualandi, M. Marchini, L. Mengozzi, H. T. Kidanu, A. Franc, P. Ceroni and P. G. Cozzi, Eur. J. Org. Chem., 2020, 2020, 1486–1490 CrossRef CAS.
- H. Madec, F. Figueiredo, K. Cariou, S. Roland, M. Sollogoub and G. Gasser, Chem. Sci., 2023, 14, 409–442 RSC.
- J. Takaya, Chem. Sci., 2021, 12, 1964–1981 RSC.
- M. Marchini, G. Bergamini, P. G. Cozzi, P. Ceroni and V. Balzani, Angew. Chem., Int. Ed., 2017, 56, 12820–12821 CrossRef CAS PubMed.
- D. Li, A. Yadav, H. Zhou, K. Roy, P. Thanasekaran and C. Lee, Glob. Chall., 2024, 8, 2300244 CrossRef PubMed.
- Nobel Prize in Chemistry 2025, https://www.nature.com/collections/cjdfehjagc, (accessed 3 November 2025) Search PubMed.
- D. Braga, F. Grepioni and L. Maini, Chem. Commun., 2010, 46, 6232–6242 RSC.
- S. R. Batten, N. R. Champness, X.-M. Chen, J. Garcia-Martinez, S. Kitagawa, L. Öhrström, M. O'Keeffe, M. P. Suh and J. Reedijk, Pure Appl. Chem., 2013, 85, 1715–1724 CrossRef CAS.
- S. R. Batten, N. R. Champness, X.-M. Chen, J. Garcia-Martinez, S. Kitagawa, L. Öhrström, M. O'Keeffe, M. P. Suh and J. Reedijk, CrystEngComm, 2012, 14, 3001–3004 RSC.
- S. R. Batten, S. M. Neville and D. R. Turner, Coordination Polymers: Design, Analysis and Application, The Royal Society of Chemistry, 2008 Search PubMed.
- K. Tan, S. Zuluaga, E. Fuentes, E. C. Mattson, J.-F. Veyan, H. Wang, J. Li, T. Thonhauser and Y. J. Chabal, Nat. Commun., 2016, 7, 13871 CrossRef CAS PubMed.
- Y. Chen, W. Lu, M. Schröder and S. Yang, Acc. Chem. Res., 2023, 56, 2569–2581 CrossRef CAS PubMed.
- Y. Gu, J.-J. Zheng, K. Otake, M. Shivanna, S. Sakaki, H. Yoshino, M. Ohba, S. Kawaguchi, Y. Wang, F. Li and S. Kitagawa, Angew. Chem., Int. Ed., 2021, 60, 11688–11694 CrossRef CAS PubMed.
- X. Liu, B. Qian, D. Zhang, M. Yu, Z. Chang and X. Bu, Coord. Chem. Rev., 2023, 476, 214921 CrossRef CAS.
- S. K. Ahmed, S. Hussein, K. Qurbani, R. H. Ibrahim, A. Fareeq, K. A. Mahmood and M. G. Mohamed, J. Med. Surg. Public Health, 2024, 2, 100081 CrossRef.
- P. Dadgostar, Infect. Drug Resist., 2019, 12, 3903–3910 CrossRef CAS PubMed.
- P. M. Vitousek, J. D. Aber, R. W. Howarth, G. E. Likens, P. A. Matson, D. W. Schindler, W. H. Schlesinger and D. G. Tilman, Ecol. Appl., 1997, 7, 737–750 Search PubMed.
- M. A. H. J. van Kessel, D. R. Speth, M. Albertsen, P. H. Nielsen, H. J. M. Op den Camp, B. Kartal, M. S. M. Jetten and S. Lücker, Nature, 2015, 528, 555–559 CrossRef CAS PubMed.
- P. G. Moe, Soil Sci. Soc. Am. J., 1967, 31, 380–382 CrossRef CAS.
- M. Q. Khan, K. Rahman, U. Ghani, A. Basharat, S. A. Qamar and M. Bilal, Case Stud. Chem. Environ. Eng., 2020, 2, 100059 CrossRef.
- N. K. Fageria and V. C. Baligar, in Advances in Agronomy, Academic Press, 2005, vol. 88, pp. 97–185 Search PubMed.
- J. A. Delgado, J. Soil Water Conserv., 2002, 57, 389–398 CrossRef.
- J.-L. Do and T. Friščić, ACS Cent. Sci., 2017, 3, 13–19 CrossRef CAS PubMed.
- E. Boldyreva, Chem. Soc. Rev., 2013, 42, 7719–7738 RSC.
- W. Jones, in Engineering Crystallography: From Molecule to Crystal to Functional Form, ed. K. J. Roberts, R. Docherty and R. Tamura, Springer Netherlands, Dordrecht, 2017, pp. 341–351 Search PubMed.
- D. Braga, L. Maini and F. Grepioni, Chem. Soc. Rev., 2013, 42, 7638–7648 RSC.
- M. Solares-Briones, G. Coyote-Dotor, J. C. Páez-Franco, M. R. Zermeño-Ortega, C. M. de la O Contreras, D. Canseco-González, A. Avila-Sorrosa, D. Morales-Morales and J. M. Germán-Acacio, Pharmaceutics, 2021, 13, 790 CrossRef CAS PubMed.
- Y. Xiao, C. Wu, X. Hu, K. Chen, L. Qi, P. Cui, L. Zhou and Q. Yin, Cryst. Growth Des., 2023, 23, 4680–4700 CrossRef CAS.
- K. Trzeciak, M. K. Dudek and M. J. Potrzebowski, Chem. – Eur. J., 2024, 30, e202402683 CrossRef CAS PubMed.
- T. Friščić, C. Mottillo and H. M. Titi, Angew. Chem., Int. Ed., 2020, 59, 1018–1029 CrossRef PubMed.
- F. Cuccu, L. De Luca, F. Delogu, E. Colacino, N. Solin, R. Mocci and A. Porcheddu, ChemSusChem, 2022, 15, e202200362 CrossRef CAS PubMed.
- S. Karki, T. Friščić and W. Jones, CrystEngComm, 2009, 11, 470–481 RSC.
- B. P. Hutchings, D. E. Crawford, L. Gao, P. Hu and S. L. James, Angew. Chem., Int. Ed., 2017, 56, 15252–15256 CrossRef CAS PubMed.
- I. R. Speight, K. J. Ardila-Fierro, J. G. Hernández, F. Emmerling, A. A. L. Michalchuk, F. García, E. Colacino and J. Mack, Nat. Rev. Methods Primers, 2025, 5, 29 CrossRef CAS.
- N. Fantozzi, J.-N. Volle, A. Porcheddu, D. Virieux, F. García and E. Colacino, Chem. Soc. Rev., 2023, 52, 6680–6714 RSC.
- K. Shankland, in International Tables for Crystallography, John Wiley & Sons, Ltd, 2019, pp. 386–394 Search PubMed.
- T. Gruene, J. J. Holstein, G. H. Clever and B. Keppler, Nat. Rev. Chem., 2021, 5, 660–668 CrossRef CAS PubMed.
- Y. Xu, S. A. Southern, P. M. J. Szell and D. L. Bryce, CrystEngComm, 2016, 18, 5236–5252 RSC.
- M. R. Chierotti and R. Gobetto, CrystEngComm, 2013, 15, 8599–8612 RSC.
- P. A. Julien and T. Friščić, Cryst. Growth Des., 2022, 22, 5726–5754 CrossRef CAS.
- G. I. Lampronti, A. A. L. Michalchuk, P. P. Mazzeo, A. M. Belenguer, J. K. M. Sanders, A. Bacchi and F. Emmerling, Nat. Commun., 2021, 12, 6134 CrossRef CAS PubMed.
- A. A. L. Michalchuk and F. Emmerling, Angew. Chem., Int. Ed., 2022, 61, e202117270 CrossRef CAS PubMed.
- F. Prestinaci, P. Pezzotti and A. Pantosti, Pathog. Global Health, 2015, 109, 309–318 CrossRef PubMed.
- K. W. K. Tang, B. C. Millar and J. E. Moore, Br. J. Biomed. Sci., 2023, 80, 11387 CrossRef PubMed.
- W. C. Reygaert, AIMS Microbiol., 2018, 4, 482–501 CAS.
- M. E. Velazquez-Meza, M. Galarde-López, B. Carrillo-Quiróz and C. M. Alpuche-Aranda, Vet. World, 2022, 15, 743–749 CAS.
- R. Laxminarayan, Lancet, 2022, 399, 606–607 CrossRef CAS PubMed.
- C. R. Steuernagel, I. Lillehagen and J. Seeberg, Global Public Health, 2024, 19, 2355318 Search PubMed.
- Nat. Commun., 2024, 15, 6198 Search PubMed.
- Global antibiotic resistance surveillance report 2025, https://www.who.int/publications/i/item/9789240116337, (accessed 24 October 2025) Search PubMed.
- C. J. L. Murray, K. S. Ikuta, F. Sharara, L. Swetschinski, G. R. Aguilar, A. Gray, C. Han, C. Bisignano, P. Rao, E. Wool, S. C. Johnson, A. J. Browne, M. G. Chipeta, F. Fell, S. Hackett, G. Haines-Woodhouse, B. H. K. Hamadani, E. A. P. Kumaran, B. McManigal, S. Achalapong, R. Agarwal, S. Akech, S. Albertson, J. Amuasi, J. Andrews, A. Aravkin, E. Ashley, F.-X. Babin, F. Bailey, S. Baker, B. Basnyat, A. Bekker, R. Bender, J. A. Berkley, A. Bethou, J. Bielicki, S. Boonkasidecha, J. Bukosia, C. Carvalheiro, C. Castañeda-Orjuela, V. Chansamouth, S. Chaurasia, S. Chiurchiù, F. Chowdhury, R. C. Donatien, A. J. Cook, B. Cooper, T. R. Cressey, E. Criollo-Mora, M. Cunningham, S. Darboe, N. P. J. Day, M. D. Luca, K. Dokova, A. Dramowski, S. J. Dunachie, T. D. Bich, T. Eckmanns, D. Eibach, A. Emami, N. Feasey, N. Fisher-Pearson, K. Forrest, C. Garcia, D. Garrett, P. Gastmeier, A. Z. Giref, R. C. Greer, V. Gupta, S. Haller, A. Haselbeck, S. I. Hay, M. Holm, S. Hopkins, Y. Hsia, K. C. Iregbu, J. Jacobs, D. Jarovsky, F. Javanmardi, A. W. J. Jenney, M. Khorana, S. Khusuwan, N. Kissoon, E. Kobeissi, T. Kostyanev, F. Krapp, R. Krumkamp, A. Kumar, H. H. Kyu, C. Lim, K. Lim, D. Limmathurotsakul, M. J. Loftus, M. Lunn, J. Ma, A. Manoharan, F. Marks, J. May, M. Mayxay, N. Mturi, T. Munera-Huertas, P. Musicha, L. A. Musila, M. M. Mussi-Pinhata, R. N. Naidu, T. Nakamura, R. Nanavati, S. Nangia, P. Newton, C. Ngoun, A. Novotney, D. Nwakanma, C. W. Obiero, T. J. Ochoa, A. Olivas-Martinez, P. Olliaro, E. Ooko, E. Ortiz-Brizuela, P. Ounchanum, G. D. Pak, J. L. Paredes, A. Y. Peleg, C. Perrone, T. Phe, K. Phommasone, N. Plakkal, A. Ponce-de-Leon, M. Raad, T. Ramdin, S. Rattanavong, A. Riddell, T. Roberts, J. V. Robotham, A. Roca, V. D. Rosenthal, K. E. Rudd, N. Russell, H. S. Sader, W. Saengchan, J. Schnall, J. A. G. Scott, S. Seekaew, M. Sharland, M. Shivamallappa, J. Sifuentes-Osornio, A. J. Simpson, N. Steenkeste, A. J. Stewardson, T. Stoeva, N. Tasak, A. Thaiprakong, G. Thwaites, C. Tigoi, C. Turner, P. Turner, H. R. van Doorn, S. Velaphi, A. Vongpradith, M. Vongsouvath, H. Vu, T. Walsh, J. L. Walson, S. Waner, T. Wangrangsimakul, P. Wannapinij, T. Wozniak, T. E. M. W. Y. Sharma, K. C. Yu, P. Zheng, B. Sartorius, A. D. Lopez, A. Stergachis, C. Moore, C. Dolecek and M. Naghavi, Lancet, 2022, 399, 629–655 CrossRef CAS PubMed.
- K. E. Arnold, G. Laing, B. J. McMahon, S. Fanning, D. J. Stekel, O. Pahl, L. Coyne, S. M. Latham and K. M. McIntyre, Lancet Planet. Health, 2024, 8, e124–e133 CrossRef PubMed.
- S. K. Gupta and R. P. Nayak, J. Pharmacol. Pharmacother., 2014, 5, 4–7 Search PubMed.
- M. Troisi, E. Marini, V. Abbiento, S. Stazzoni, E. Andreano and R. Rappuoli, Front. Microbiol., 2022, 14 DOI:10.3389/fmicb.2022.1080059.
- T. Jesudason, Lancet Microbe, 2025, 6(10) DOI:10.1016/j.lanmic.2025.101204.
- D. Sharma, I. Singh, J. Sharma, I. K. Verma and A. Ratn, J. Basic Microbiol., 2025, e70090 Search PubMed.
- I. E. Mba and E. I. Nweze, Yale J. Biol. Med., 2022, 95, 445–463 CAS.
- A. Alvani, A. Shayanfar and R. Kaviani, Chem. Pap., 2025, 79, 2679–2694 CrossRef CAS.
- M. Bashimam and H. El-Zein, Heliyon, 2022, 8, e11872 CrossRef CAS PubMed.
- A. Kumar, S. Kumar and A. Nanda, Adv. Pharm. Bull., 2018, 8, 355–363 CrossRef PubMed.
- A. K. Nangia and G. R. Desiraju, Angew. Chem., Int. Ed., 2022, 61, e202207484 CrossRef CAS PubMed.
- R. J. Turner, Microb. Biotechnol., 2017, 10, 1062–1065 CrossRef PubMed.
- A. Frei, A. D. Verderosa, A. G. Elliott, J. Zuegg and M. A. T. Blaskovich, Nat. Rev. Chem., 2023, 7, 202–224 Search PubMed.
- R. J. Turner, BioMetals, 2024, 37, 545–559 Search PubMed.
- J. A. Lemire, J. J. Harrison and R. J. Turner, Nat. Rev. Microbiol., 2013, 11, 371–384 CrossRef CAS PubMed.
- I. Luz, I. E. Stewart, N. P. Mortensen and A. J. Hickey, Chem. Commun., 2020, 56, 13339–13342 Search PubMed.
- S. Quaresma, V. André, A. Fernandes and M. T. Duarte, Inorg. Chim. Acta, 2017, 455, 309–318 CrossRef CAS.
- S. Quaresma, V. André, A. M. M. Antunes, S. M. F. Vilela, G. Amariei, A. Arenas-Vivo, R. Rosal, P. Horcajada and M. T. Duarte, Cryst. Growth Des., 2020, 20, 370–382 Search PubMed.
- H. Li and H. Sun, in Encyclopedia of Inorganic and Bioinorganic Chemistry, John Wiley & Sons, Ltd, 2022, pp. 1–19 Search PubMed.
- E. J. Anthony, E. M. Bolitho, H. E. Bridgewater, O. W. L. Carter, J. M. Donnelly, C. Imberti, E. C. Lant, F. Lermyte, R. J. Needham, M. Palau, P. J. Sadler, H. Shi, F.-X. Wang, W.-Y. Zhang and Z. Zhang, Chem. Sci., 2020, 11, 12888–12917 Search PubMed.
- Q. Peña, A. Wang, O. Zaremba, Y. Shi, H. W. Scheeren, J. M. Metselaar, F. Kiessling, R. M. Pallares, S. Wuttke and T. Lammers, Chem. Soc. Rev., 2022, 51, 2544–2582 RSC.
- D. A. Salazar-Alemán and R. J. Turner, in Microbial Metabolism of Metals and Metalloids, ed. C. J. Hurst, Springer International Publishing, Cham, 2022, pp. 77–106 Search PubMed.
- G. Gasser, Chimia, 2015, 69, 442–442 Search PubMed.
- G. Gasser and N. Metzler-Nolte, Curr. Opin. Chem. Biol., 2012, 16, 84–91 Search PubMed.
- T. Gianferrara, I. Bratsos and E. Alessio, Dalton Trans., 2009, 7588–7598 RSC.
- S. Wu, M. Wang, Z. Liu and C. Fu, Microorganisms, 2025, 13, 1570 CrossRef CAS PubMed.
- C. Fiore, O. Shemchuk, F. Grepioni, R. J. Turner and D. Braga, CrystEngComm, 2021, 23, 4494–4499 Search PubMed.
- A. Lekhan, C. Fiore, O. Shemchuk, F. Grepioni, D. Braga and R. J. Turner, ACS Appl. Bio Mater., 2022, 5, 4203–4212 CrossRef CAS PubMed.
- M. Guerrini, S. d'Agostino, F. Grepioni, D. Braga, A. Lekhan and R. J. Turner, Sci. Rep., 2022, 12, 3673 CrossRef CAS PubMed.
- O. Shemchuk, D. Braga, F. Grepioni and R. J. Turner, RSC Adv., 2020, 10, 2146–2149 RSC.
- C. Fiore, F. Antoniciello, D. Roncarati, V. Scarlato, F. Grepioni and D. Braga, Pharmaceutics, 2024, 16, 203 Search PubMed.
- O. Shemchuk, S. d'Agostino, C. Fiore, V. Sambri, S. Zannoli, F. Grepioni and D. Braga, Cryst. Growth Des., 2020, 20, 6796–6803 CrossRef CAS.
- C. Fiore, A. Baraghini, O. Shemchuk, V. Sambri, M. Morotti, F. Grepioni and D. Braga, Cryst. Growth Des., 2022, 22, 1467–1475 Search PubMed.
- R. Sun, L. Casali, R. J. Turner, D. Braga and F. Grepioni, Molecules, 2023, 28, 1244 CrossRef CAS PubMed.
- L. Contini, A. Paul, L. Mazzei, S. Ciurli, D. Roncarati, D. Braga and F. Grepioni, Dalton Trans., 2024, 53, 10553–10562 RSC.
- L. Contini, R. J. Turner and F. Grepioni, Dalton Trans., 2025, 54, 11925–11934 RSC.
- S. d'Agostino, L. Macchietti, R. J. Turner and F. Grepioni, Front. Chem., 2024, 12, 1430457 CrossRef PubMed.
- C. Fiore, A. Lekhan, S. Bordignon, M. R. Chierotti, R. Gobetto, F. Grepioni, R. J. Turner and D. Braga, Int. J. Mol. Sci., 2023, 24, 5180 CrossRef CAS PubMed.
- S. J. S. Qazi and R. J. Turner, Biochem. Biophys. Rep., 2018, 13, 129–140 Search PubMed.
- M.-E. Nedu, M. Tertis, C. Cristea and A. V. Georgescu, Diagnostics, 2020, 10, 223 CrossRef CAS PubMed.
- O. Shemchuk, D. Braga, F. Grepioni and R. J. Turner, RSC Adv., 2020, 10, 2146–2149 RSC.
- A. Smith, Br. Dent. J., 2018, 224, 403–404 CrossRef CAS PubMed.
- L. Boyanova, P. Hadzhiyski, R. Gergova and R. Markovska, Antibiotics, 2023, 12, 332 CrossRef CAS PubMed.
- A. B. Turner, E. Gerner, R. Firdaus, M. Echeverz, M. Werthén, P. Thomsen, S. Almqvist and M. Trobos, Front. Microbiol., 2022, 13, 931839 CrossRef PubMed.
- B. Z. Houlton, M. Almaraz, V. Aneja, A. T. Austin, E. Bai, K. G. Cassman, J. E. Compton, E. A. Davidson, J. W. Erisman, J. N. Galloway, B. Gu, G. Yao, L. A. Martinelli, K. Scow, W. H. Schlesinger, T. P. Tomich, C. Wang and X. Zhang, Earth's Future, 2019, 7, 865–872 CrossRef PubMed.
- R. Nieder and D. K. Benbi, Rev. Environ. Health, 2022, 37, 229–246 CrossRef CAS PubMed.
- B. L. Bodirsky, A. Popp, H. Lotze-Campen, J. P. Dietrich, S. Rolinski, I. Weindl, C. Schmitz, C. Müller, M. Bonsch, F. Humpenöder, A. Biewald and M. Stevanovic, Nat. Commun., 2014, 5, 3858 CrossRef CAS PubMed.
- P. Bacon, Nitrogen Fertilization in the Environment, CRC Press, 1995 Search PubMed.
- J. W. Erisman, M. A. Sutton, J. Galloway, Z. Klimont and W. Winiwarter, Nat. Geosci., 2008, 1, 636–639 Search PubMed.
- N. W. Krase and V. L. Gaddy, J. Ind. Eng. Chem., 1922, 14, 611–615 CrossRef CAS.
- G. L. Terman, in Advances in Agronomy, ed. N. C. Brady, Academic Press, 1980, vol. 31, pp. 189–223 Search PubMed.
- K. C. Cameron, H. J. Di and J. L. Moir, Ann. Appl. Biol., 2013, 162, 145–173 CrossRef CAS.
- B. P. Callahan, Y. Yuan and R. Wolfenden, J. Am. Chem. Soc., 2005, 127, 10828–10829 Search PubMed.
- Y. Qin and J. M. S. Cabral, Biocatal. Biotransform., 2009, 20, 1–14 Search PubMed.
- L. Mazzei, F. Musiani and S. Ciurli, J. Biol. Inorg. Chem., 2020, 25, 829–845 CrossRef CAS PubMed.
- A. K. Rosmarina, Y. M. Khanif, M. M. Hanafi, A. Hussin and K. B. A. Rahim, Environ. Sci., 2016, 16, 641–651 CAS.
- S. V. Krupa, Environ. Pollut., 2003, 124, 179–221 CrossRef CAS PubMed.
- F. Musiani, V. Broll, E. Evangelisti and S. Ciurli, J. Biol. Inorg. Chem., 2020, 25, 995–1007 CrossRef CAS PubMed.
- G. Zhu, S. Wang, Y. Li, L. Zhuang, S. Zhao, C. Wang, M. M. M. Kuypers, M. S. M. Jetten and Y. Zhu, Environ. Microbiol., 2018, 20, 1723–1738 CrossRef CAS PubMed.
- T. Al-Kanani, A. F. MacKenzie and H. Blenkhorn, Fert. Res., 1990, 23, 113–119 CrossRef CAS.
- P. Govindasamy, S. K. Muthusamy, M. Bagavathiannan, J. Mowrer, P. T. K. Jagannadham, A. Maity, H. M. Halli, G. K. Sujayananad, R. Vadivel, R. Raj, V. Pooniya, S. Babu, S. S. Rathore, L. Muralikrishnan and G. Tiwari, Front. Plant Sci., 2023, 14, 1121073 Search PubMed.
- A. Mălinaş, R. Vidican, I. Rotar, C. Mălinaş, C. M. Moldovan and M. Proorocu, Plants, 2022, 11, 217 CrossRef PubMed.
- B. Beig, M. B. K. Niazi, Z. Jahan, A. Hussain, M. H. Zia and M. T. Mehran, J. Plant Nutr., 2020, 43, 1510–1533 CrossRef CAS.
- M. Y. Naz and S. A. Sulaiman, J. Controlled Release, 2016, 225, 109–120 CrossRef CAS PubMed.
- H. Cantarella, R. Otto, J. R. Soares and A. G. de B. Silva, J. Adv. Res., 2018, 13, 19–27 CrossRef CAS PubMed.
- A. B. da Fonseca, C. Santos, A. P. P. Nunes, D. P. Oliveira, M. E. A. de Melo, T. Takayama, B. L. Mansur, T. de Jesus Fernandes, G. do Carmo Alexandrino, M. A. N. Dias and D. Guelfi, Sci. Rep., 2023, 13, 22739 CrossRef CAS PubMed.
- M. Ammar, S. Ashraf, D. A. Gonzalez-Casamachin and J. Baltrusaitis, RSC Sustainability, 2025, 3, 781–803 RSC.
- M. Eisa, M. Brondi, C. Williams, R. Hejl and J. Baltrusaitis, Sustainable Chem. Environ., 2025, 9, 100209 CrossRef CAS.
- S. S. Fatima, R. Kumar, M. I. Choudhary and S. Yousuf, IUCrJ, 2020, 7, 105–112 CrossRef CAS PubMed.
- V. Nagaraju, C. Jange, C. Wassgren and K. Ambrose, J. Environ. Chem. Eng., 2024, 12, 114308 CrossRef CAS.
- K. Honer, E. Kalfaoglu, C. Pico, J. McCann and J. Baltrusaitis, ACS Sustainable Chem. Eng., 2017, 5, 8546–8550 Search PubMed.
- P. A. Julien, L. S. Germann, H. M. Titi, M. Etter, R. E. Dinnebier, L. Sharma, J. Baltrusaitis and T. Friščić, Chem. Sci., 2020, 11, 2350–2355 RSC.
- L. Casali, L. Mazzei, O. Shemchuk, K. Honer, F. Grepioni, S. Ciurli, D. Braga and J. Baltrusaitis, Chem. Commun., 2018, 54, 7637–7640 RSC.
- L. Mazzei, V. Broll, L. Casali, M. Silva, D. Braga, F. Grepioni, J. Baltrusaitis and S. Ciurli, ACS Sustainable Chem. Eng., 2019, 7, 13369–13378 CrossRef CAS.
- L. Casali, L. Mazzei, O. Shemchuk, L. Sharma, K. Honer, F. Grepioni, S. Ciurli, D. Braga and J. Baltrusaitis, ACS Sustainable Chem. Eng., 2019, 7, 2852–2859 CrossRef CAS.
- L. Contini, D. Gogoi, A. C. Fluck, J. G. Farias, C. Williams, R. Thakuria, J. Baltrusaitis and F. Grepioni, ACS Sustainable Resour. Manage., 2025, 2(9), 1672–1680 Search PubMed.
- L. Casali, T. Feiler, M. Heilmann, D. Braga, F. Emmerling and F. Grepioni, CrystEngComm, 2022, 24, 1292–1298 RSC.
- L. Casali, V. Broll, S. Ciurli, F. Emmerling, D. Braga and F. Grepioni, Cryst. Growth Des., 2021, 21, 5792–5799 CrossRef CAS.
- A. Azzali, S. d'Agostino and F. Grepioni, ACS Sustainable Chem. Eng., 2021, 9, 12530–12539 Search PubMed.
- L. Casali, L. Mazzei, R. Sun, M. R. Chierotti, R. Gobetto, D. Braga, F. Grepioni and S. Ciurli, Cryst. Growth Des., 2022, 22, 4528–4537 Search PubMed.
- B. Sandhu, A. S. Sinha, J. Desper and C. B. Aakeröy, Chem. Commun., 2018, 54, 4657–4660 Search PubMed.
- B. Krajewska, J. Mol. Catal. B: Enzym., 2009, 59, 9–21 CrossRef CAS.
- W. K. Keener, S. A. Russell and D. J. Arp, Biochim. Biophys. Acta, 1998, 1388, 373–385 CrossRef CAS PubMed.
- Y. Cao, J. Zhang, Y. Liu, L. Zhang, L. Wang, J. Wang, Y. Qi, H. Lv, J. Liu, L. Huo, X. Wei and Y. Shi, Medicine, 2021, 100, e27923 CrossRef CAS PubMed.
- M. P. Dore, H. Lu and D. Y. Graham, Gut, 2016, 65, 870–878 CrossRef CAS PubMed.
- R. M. Zagari, S. Rabitti, L. H. Eusebi and F. Bazzoli, Eur. J. Clin. Invest., 2018, 48(1) DOI:10.1111/eci.12857.
- K. Kappaun, A. R. Piovesan, C. R. Carlini and R. Ligabue-Braun, J. Adv. Res., 2018, 13, 3–17 CrossRef CAS PubMed.
- L. Zhang, S. B. Mulrooney, A. F. K. Leung, Y. Zeng, B. B. C. Ko, R. P. Hausinger and H. Sun, BioMetals, 2006, 19, 503–511 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |