Structure–activity relationships of aniline-based squaraines for distinguishable staining and bright two-photon fluorescence bioimaging in plant cells†
Nan
Zhang
,
Haixia
Chang
,
Rong
Miao
,
Taihong
Liu
 *,
Liping
Ding
*,
Liping
Ding
 and
Yu
Fang
and
Yu
Fang
 *
*
Key Laboratory of Applied Surface and Colloid Chemistry of Ministry of Education, School of Chemistry and Chemical Engineering, Shaanxi Normal University, Xi’an 710119, P. R. China. E-mail: liuth121@snnu.edu.cn; yfang@snnu.edu.cn
First published on 16th April 2024
Abstract
An organelle-selective vision provides insights into the physiological response of plants and crops to environmental stresses in sustainable agriculture ecosystems. Biological applications often require two-photon excited fluorophores with low phototoxicity, high brightness, deep penetration, and tuneable cell entry. We obtained three aniline-based squaraines (SQs) tuned from hydrophobic to hydrophilic characteristics by modifying terminal pendant groups and substituents, and investigated their steady-state absorption and far-red-emitting fluorescence properties. The SQs exhibited two-photon absorption (2PA) ranging from 750 to 870 nm within the first biological spectral window; their structure–property relationships, corresponding to the 2PA cross sections (δ2PA), and structure differences were demonstrated. The maximum δ2PA value was ∼1220 GM at 800 nm for hydrophilic SQ3. Distinct biological staining efficiency and selective SQ bioimaging were evaluated utilizing the onion epidermal cell model. Contrary to the hydrophobic SQ1 results in the onion epidermal cell wall, amphiphilic SQ2 tagged the vacuole and nucleus and SQ3 tagged the vacuole. Distinguishable staining profiles in the roots and leaves were achieved. We believe that this study is the first to demonstrate distinct visualisation efficiency induced by the structure differences of two-photon excited SQs. Our results can help establish the versatile roles of novel near-infrared-emitting SQs in biological applications.
Introduction
Sustainable and smart agriculture ecosystems have increasingly embraced information science for detecting and quantifying bioactive components in plants and crops through a comprehensive integration of sensor networks.1–3 New sensing innovations and imaging capabilities are crucial and conducive to collecting agricultural metrics, providing insights into how plant physiology responds to environmental stresses, such as pathogen infection, nutrient levels, and pesticide accumulation, to name a few.4,5 Fluorescence techniques feature the intrinsic advantages of high sensitivity, spatial resolution, non-invasive visualization, etc. Concurrently, the strong endogenous autofluorescence of biological tissue and a limited penetration depth hinder the promotion of conventional plant staining.6 The important emerging area of whole animal and deep tissue imaging has made long wavelength and near-infrared (NIR) imaging have special significance. NIR-emitting fluorophores within the notably biological spectral window (650–1450 nm) are of ongoing interest due to the merits of deep tissue penetration, reduced light scattering, and high imaging signal-to-noise (S/N) ratio.7–9 Novel small-molecule organic fluorophores for specific staining and fixing delimit the cellular contour, thus enabling a clear distinction between extracellular and intracellular components.10 Therefore, photochemically stable and selective NIR-emitting fluorophores with enhanced brightness are highly desirable. In the past few years, two-photon absorption (2PA) and two-photon excited fluorescence (2PEF) techniques have advanced significantly in high-resolution bioimaging and photodynamic therapy due to the remarkable advantages of reduced scattering and ignorable auto-fluorescence of normal tissue. Regarding efficient two-photon excited fluorophores, their characteristics of high 2PA cross sections (δ2PA) and rational incubation play important roles in practical applications, e.g., bioimaging.11–13Compared to some small-molecule organic fluorophores, multipolar squaraines (SQs) possessing relatively sharp and intense absorption, long-wavelength emission, and outstanding photostability have been demonstrated as promising NIR-emitting candidates for broad biological applications.14–18 Long-wavelength excitation and emission of SQs have attractive advantages of deep tissue penetration, minimum photodamage, and minimum interference from auto-fluorescence in biological systems, as compared with some commercial fluorophores illustrated in Scheme S3 (ESI†). While zwitterionic SQs have the tendency to undergo H-/J-aggregation in aqueous media, which may restrict their optical applications to some extent due to the fluorescence emission quenching. Considering the comprehensive characteristics of the aniline-based SQs, there are several functionalization strategies for synthesizing novel derivatives, including the dendritic SQ rotaxanes,19 asymmetrical SQ analogues,20 functionalized dicyanomethylkene,21 substituents adjacent to the SQ core,22etc. (Scheme 1a). The quadrupolar and zwitterionic donor–acceptor–donor (D–A–D) motifs render the SQs noticeable intramolecular charge transfer (ICT) and large transition dipoles, favourable for 2PA.8,23,24 However, promising SQs for specific cell entry and the structure–property correlations are rarely examined.25–28 For instance, Ramaiah et al. examined the negligible absorption changes and considerable fluorescence enhancement of a few amphiphilic SQs, indicating their effective microencapsulation in mimic biological environments.29 Zhao et al. validated the strategy of micelle encapsulation of a monomeric SQ in aqueous media for excellent in vivo fluorescence and photoacoustic bimodal imaging.21 Belfield et al. identified that the substituents and terminal pendant groups adjacent to the SQ core affected the relevant optical properties and aggregation behaviours.30 Meanwhile, the chemical stability toward the nucleophilic attack was improved. Despite the extensive advances in SQs, problems of tedious aggregation and low chemical stability are often encountered. There are still attractive considerations of novel SQs for addressing the challenges and exploring some specific requirements.27,31,32 In this contribution, three aniline-based SQs (hydrophobic SQ1, amphiphilic SQ2, and hydrophilic SQ3, Scheme 1b) tuned from hydrophobic to hydrophilic characteristics were obtained, and their steady-state optical properties were first studied. Modifications of increased steric hindrance close to the SQ core aided in reducing their self-aggregation.22 Broadband 2PA and excellent δ2PA values were obtained based on the open-aperture Z-scan method. The maxima δ2PA values of amphiphilic SQ2 and hydrophilic SQ3 were found to be ∼1060 GM and ∼1220 GM at 800 nm, respectively. Mimicked biological applications in the plant cells were further conducted using a confocal laser scanning microscope (CLSM). Interestingly, though the SQs have similar absorption spectra in neat solvents, corresponding apparent differences were observed in the cellular trials. Rapid uptake and uniform distribution of hydrophobic SQ1 in the onion epidermal cell wall were investigated. Amphiphilic SQ2 had special affinities to the vacuole and nucleus, unlike the hydrophilic SQ3 differently attached to the former. Based on the two-photon fluorescence microscopy (2PFM), comparative staining profiles in the roots and leaves of the Arabidopsis seedlings were accomplished. Deep tissue penetration and high-resolution three-dimensional (3D) reconstruction in the plants were further demonstrated toward bioimaging applications.
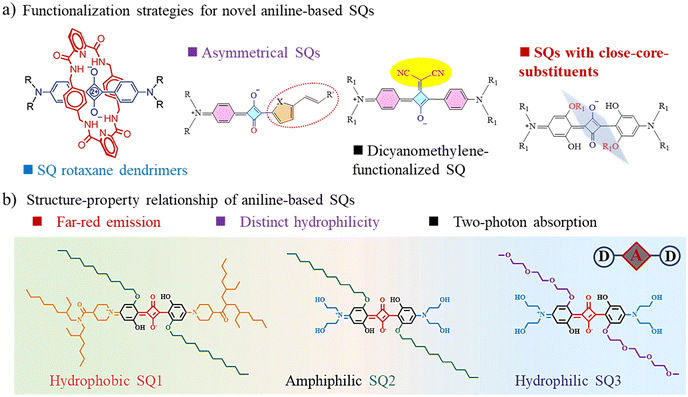 | ||
| Scheme 1 (a) Functionalization strategies for novel aniline-based SQs. (b) Molecular structures of the three investigated SQs. | ||
Results and discussion
Linear optical properties of the SQs
Steady-state photophysical properties of the hydrophobic SQ1, amphiphilic SQ2, and hydrophilic SQ3 in various solvents (Fig. S1, ESI†) were investigated and key optical parameters are summarized in Table 1. Typical UV-vis absorption maxima (λmax) of the SQs around 650 nm with significant molar extinction coefficients (ε = 1.47–2.09 × 105 M−1 cm−1 in THF) corresponded to the S0 → S1 (π–π*) transition processes. Relatively weak and short-wavelength absorption bands around 360 nm could be assigned to the S0 → S2 transition (Fig. 1a).33 Upon excitation, three SQs emitted similar reddish emission at around 670 nm and fulfilled the mirror symmetry rules (Fig. 1b).34 Negligible aggregation in organic solvents (Fig. S2, ESI†), and the fluorescence quantum yields (ΦF) ranging from 0.31 to 0.45 in the polar DMSO were characterized. Red-shifted emission maxima and low ΦF values manifested when increasing the polarities of the surrounding medium (Fig. S3, ESI†). It implied that the corresponding far-red emission was particularly beneficial for deep tissue imaging as well as improved penetration depth and image quality. In H2O, the UV-vis absorption spectra of SQ1 and SQ2 broadened significantly due to the self-aggregation. The values of full widths at half-maximum (fwhm) expanded from 39 nm of SQ3 to 185 nm of SQ2. Notably, a blue-shifted peak at 569 nm, not associated with the SQ2 monomer, could be assigned to its non-emissive H-dimer (Fig. 1c).35 We speculated that despite the structural distortion because of the close-core substituents, the squaraine skeleton still retained a certain amount of overall planarity which would facilitate the intermolecular π–π stacking interaction for aggregation.22,30 A low ΦF value of ∼0.03 and a short fluorescence lifetime of ∼2.18 ns for SQ3 in the aqueous medium could be attributed to its efficient interactions with surrounding solvent, resulting in the non-radiative decay processes (Fig. S4, ESI†).29| Comp. | Solv. | λ Abs/nm | fwhm/nm | ε/M−1 cm−1 | λ Em/nm | Δλ/nm (cm−1) | τ/ns | Φ F | λ max2PA/nm | δ max2PA/GM |
|---|---|---|---|---|---|---|---|---|---|---|
| SQ1 | THF | 642 | 33 | 1.47 × 105 | 672 | 30 (695) | 1.75 | 0.26 | 820 | 991 |
| DMSO | 653 | 37 | 1.16 × 105 | 678 | 25 (565) | 2.51 | 0.31 | — | — | |
| H2O | 650 | 79 | 2.75 × 104 | — | — | — | — | — | — | |
| SQ2 | THF | 642 | 31 | 2.09 × 105 | 675 | 33 (761) | 2.87 | 0.27 | 800 | 1060 |
| DMSO | 653 | 36 | 1.76 × 105 | 678 | 25 (565) | 2.85 | 0.32 | — | — | |
| H2O | 569/646 | 185 | 3.03 × 104/3.41 × 104 | — | — | — | — | — | — | |
| SQ3 | THF | 642 | 31 | 1.64 × 105 | 673 | 31 (717) | 3.26 | 0.31 | 800 | 1220 |
| DMSO | 654 | 36 | 1.55 × 105 | 678 | 24 (432) | 2.77 | 0.45 | — | — | |
| H2O | 640 | 39 | 1.30 × 105 | 676 | 36 (832) | 2.18 | 0.03 | — | — | |
Degenerate 2PA and 2PEF properties of the SQs
δ 2PA values reflect the two-photon activity efficiency of the fluorophores interacting with the ultrashort laser pulses. The degenerate δ2PA values of SQs were investigated in THF and acquired by a standard open-aperture Z-scan method employing the excitation of 200 femtosecond pulses.35–37 Z-scan profiles of the SQs showed intense transmission valleys indicative of typical reverse saturable absorption behaviors (Fig. 1d and Fig. S5, ESI†). The occurred intensity-dependent absorption of the incident light indicated an induced positive nonlinear absorption phenomenon (Fig. S6, ESI†).11,36 Significant δ2PA values ranging from 750 to 870 nm (Table S1, ESI†), partially consistent with the NIR biological spectral window, were obtained.38 The maximum δ2PA value was found to be ∼1220 GM at 800 nm for SQ3, a little higher than that of SQ2 (1060 GM). Interestingly, the specific δ2PA maximum for hydrophobic SQ1 (∼991 GM) was found at a longer wavelength of 820 nm (Fig. 1e). We speculated that the molecular structure and ICT character of the excited states strongly influenced the corresponding 2PA responses. Confirmed by the 2PEF strategy, the quadratic relationships with the slopes of 1.92–1.89 at 800 nm between the fluorescence intensity and the excitation power indicated their intrinsic 2PA properties (Fig. S7, ESI†). As previously discovered, one meaningful aspect of 2PA different from 1PA is the distinct selection rules for the electronic dipole transitions dependent upon the molecular symmetry. Therefore, dominant 2PA transitions close to the 1PA maxima rather than the 2-fold relation could be elucidated using the three-level energy model. It means that these large values of δ2PA for symmetrical SQs could be mainly rationalized by the large molar extinction coefficient and a resonance-enhanced mechanism (Fig. 1f).34,35,39 Extensive 2PA properties within the first NIR biological window and variable hydrophilicity are beneficial for the bioimaging applications.Comparative vision of the onion epidermal cells incubated with the distinct SQs
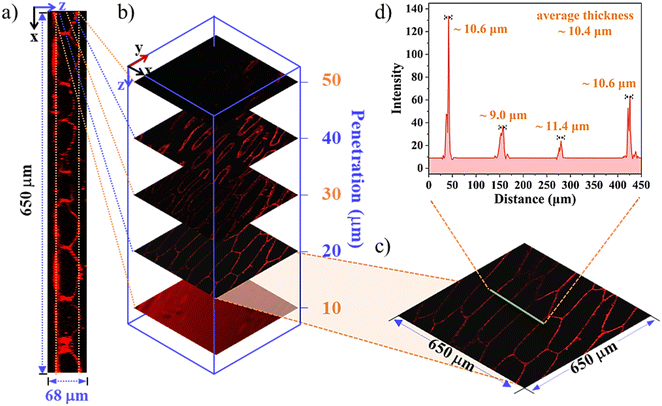
Fluorescent labelling combined with staining is an evergreen research field because of its intrinsic sensitivity, high spatiotemporal resolution, non-invasion, etc.40,41 Aiming to achieve the proof-of-concept objectives, cellular experiments of the SQs utilizing the model of onion epidermal cells were first assessed. Rapid cellular uptake of the SQs and notable differences in their internalization and organelle distribution were illustrated (Fig. 2). As shown in the CLSM images of Fig. 2a and Fig. S8 (ESI†), clear quasi-rectangular profiles of the onion epidermal cells were observable. In contrast to their low luminescence efficiency in H2O, they emitted bright and reddish fluorescence under an excitation of 635 nm. The microenvironment changes were rationalized that the SQs microencapsulated into the cell walls with high viscosity and low polarity. Even for 5 min incubation at the low concentrations of SQs, SQ1 displayed bright and strong fluorescence along the edges of the cells with noticeable superiority, supported by the control cells untreated with the SQs (Fig. S9, ESI†). We deemed that the hydrophobic SQ1 showed high retention affinity to the hydrophobic lignin domains of the cell wall.6,42 The average thickness ∼9.8 μm and cell width (∼62.7 μm) could be observed at high quality (Fig. 2a–d). The other two SQs with subtle structure changes exhibited distinct bioactive and biocompatible properties. There was distinguishable visibility of intracellular space of the cells tagged with SQ2, sequestrated in the vacuole and nucleus (Fig. 2e–h), as confirmed by the co-localization of the nuclear staining agent Hoechst 33342 (Fig. 2a, e and i, and Fig. S8, ESI†). The corresponding Pearson's correlation coefficient value of 0.825 in the cell nucleus was obtained for SQ2. The intracellular space and cell nucleus showed bright fluorescence, and a similar cell width around 70.2 μm was estimated properly. Hydrophilic SQ3 was distributed through the whole cell vacuole with bright fluorescence rather than a dark nucleus (Fig. 2i–l). The Pearson's correlation coefficients for the colocalization of SQ3 with Hoechst were found to be 0.134, demonstrating SQ3 was unable to enter the cell nucleus (Fig. 2i). Through the colour inversion processing, the cell nucleus size could be estimated to be ∼5.5 μm. To further explore the effectiveness of SQs in imaging the morphology of onion inner epidermal cells, the CLSM images in the absence and presence of NaCl solution were obtained. The outlines of cells stained with the SQs became obviously indistinct after the addition of NaCl solution (Fig. S10–S12, ESI†), proving the dehydration effect of salt on the biological cells.As 3D imaging provides a more accurate view, high-resolution 3D reconstructions were performed under conventional experimental conditions. As shown in Fig. 3, a deep penetration as large as 68 μm was achieved with a step depth of 10 μm, and the intercellular and spatial space of the onion epidermal cells could be visualized easily (Fig. 3a). Selected 2D fluorescence images of the cells became less uniform as the orthoslice depth increased (Fig. 3b). Analysing the periodic cytoskeleton grids in the selected zone of 650 × 650 μm (Fig. 3c), it confirmed the specific affinity of SQ1 to the cell wall, and the average wall thickness (n = 4) was estimated to be ∼10.4 μm (Fig. 3d). Similar 3D reconstruction of the amphiphilic SQ2 (Fig. S13, ESI†) and hydrophilic SQ3 (Fig. S14, ESI†) was also observed, supporting their specific bioactive properties to the vacuole and nucleus as mentioned above. These results indicated that alterations in the cell size and staining distribution could further function as plant health markers and responses to environmental stimuli at the cellular level. The in vivo imaging trials in onions were also conducted and the overall better results of SQ2 and SQ3 than SQ1 were investigated properly (Fig. S15, ESI†), indicating that SQ2 and SQ3 with better water hydrophilicity permeated efficiently throughout the onions.43
Uptake and translocation observations of the SQs in plants
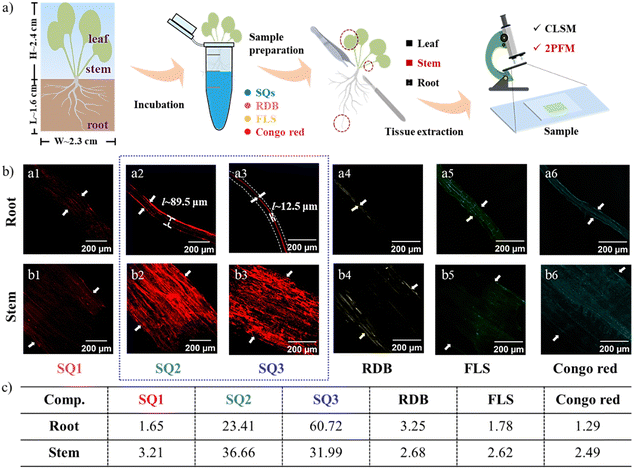
Inspired by the intriguing results in the onion epidermal cell model, we developed a staining protocol for the plants underlying various cellular events. The original Arabidopsis plants were obtained commercially and cultivated in water. Around the 14th day, a height of ∼2.4 cm and a root length of ∼1.6 cm were measured properly. By incubating the plant root in the aqueous solution of the fluorophores, the SQs could access stems and leaves via the biological translocation system (Fig. 4a). In contrast, the Arabidopsis root and stem incubated with SQ1 showed inhomogeneous and low staining characteristics (Fig. 4b). The S/N ratios were calculated to be 1.65 and 3.21, respectively. We deemed that the Arabidopsis seedling roots and stems were inactive for the uptake of SQ1. It means that good biocompatibility should be a crucial and necessary property for biological applications. The substituents of aliphatic alcohols and glycol units render SQ2 and SQ3 good solubility in aqueous medium and improve fluorescence intensity and cellular uptake. As anticipated, the uptakes of SQ2 and SQ3 could be identified in vivo due to the strong and reddish emissions under 635 nm excitation. The S/N ratios enhanced to be 23.41 in roots and 36.66 in stems. Concurring with the onion epidermal cell results, SQ2 tagged the cortex in roots (width ∼ 89.5 μm). SQ3 was witnessed and labelled on the root xylem (S/N ∼ 60.72) with a width of 12.5 μm.44 The primary xylem is an important vascular tissue in plants responsible for continuously transporting water from the roots to the leaves. SQ2 and SQ3 also stained the stems (S/N ∼ 31.99), which could be explained by a much higher lignification process in the stem relative to other zones.45 To observe the staining more intuitively, seedling bioimaging under different incubation times (1–30 min) was also recorded (Fig. S16 and S17, ESI†). The longer the incubation time, the clearer the imaging efficiency (Fig. S18, ESI†), clarifying the rapid SQ uptake within 5 min of incubation. Concerning the efficacy of the SQs compared with some commercial fluorophores under the same experiment conditions, the samples stained with the small-molecule organic fluorophores like rhodamine B (RDB), fluorescein sodium (FLS), and congo red produced relatively nonspecific staining and low fluorescence brightness (Fig. 4b and Fig. S16–S19, ESI†). Their S/N ratios were evaluated in the levels of 1.29–3.25, significantly lower than those of SQ2 and SQ3. These advantages of tissue selectivity and 2PEF characteristics would provide new insights for applying SQ2 and SQ3 in monitoring plant growth and development by visualization.In vivo 2PFM trials and high-resolution 3D reconstruction
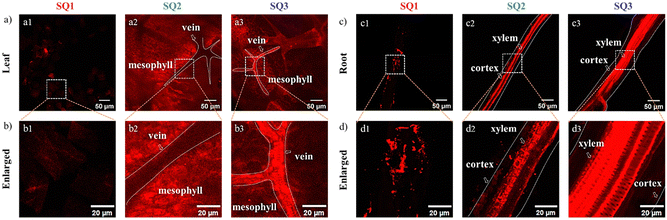
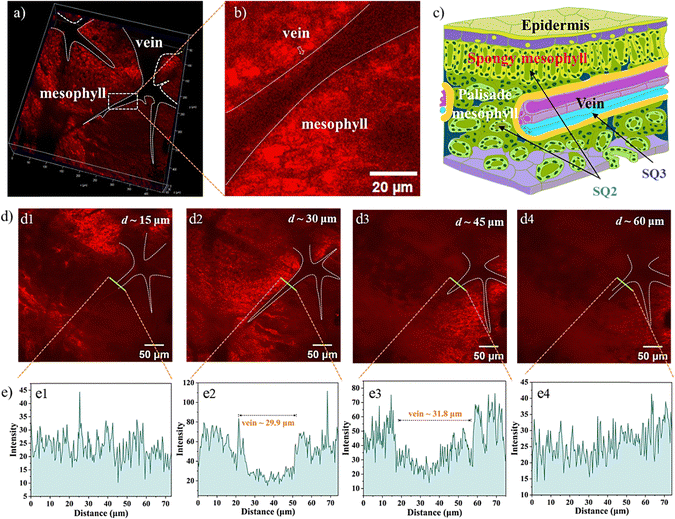
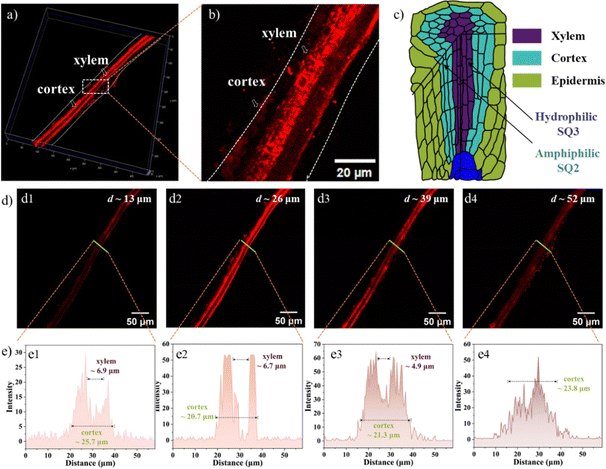
To evaluate NIR-emitting labels and monitor biological translocation, the Arabidopsis seedling incubated with the SQs was evaluated utilizing 2PFM. Initially, the excitation wavelength-dependent brightness values in the leaves were recorded, supporting the optimized excitation wavelength of the SQs above 780 nm for 2PEF (Fig. S20, ESI†). Thus, excited by an 800 nm pulse laser, bright 2PEF images of the Arabidopsis leaves and roots could be achieved (Fig. 5). First, the SQ1 could be absorbed by the plant tissues, and uneven reddish luminescence in leaves and roots was observed. The unsatisfactory staining results of SQ1 might be due to its hydrophobic character. The better imaging efficiencies of SQ2 and SQ3 than that of SQ1 were consistent with the above investigations. The leaf is a complex framework of epidermal cells. Inner leaf tissue consists of spongy mesophyll, palisade mesophyll, veins, etc. The root, responsible for uptake of water and minerals from the soil, consists of the tissues of xylem, phloem, cortex, epidermis, etc.46 Fluorescence brightness of amphiphilic SQ2 selectively accumulated in the mesophyll tissue (Fig. 5a2) and the cortex in the root (Fig. 5c2) at the subcellular level.47 Arabidopsis root tissue was also stained using SQ2, demonstrating unique tissue selectivity attributed to its water solubility and lignin specificity. In contrast, hydrophilic SQ3 accumulated in the leaf vein (Fig. 5a3) and the xylem tissue in roots (Fig. 5c3). These distinguishable structure–activity relationships inspired us to study their staining efficiency in detail below.Visualizing biological activities at high spatiotemporal resolution is essential for unravelling fundamental biological processes and requires better and brighter fluorophores. The impressive translocation pathways of the SQs prompted us to reconstruct high-resolution 3D rendering images (a planar size of 400 × 400 μm and vertical depths of 72–108 μm). The contrast plot of SQ2 showed the profiles and distribution of veins, which were not brilliant. The contrast results showed the special feasibility of SQ2 to the leaf mesophyll tissue and proved the translocation from root to leaf in the present trials through the vascular system (Fig. 6a).48 A diagram of the Arabidopsis leaf architecture and cell landscape showed the strong staining of SQ2 to the leaf mesophyll (Fig. 6c). The orthoslice images at the selected depths indicated strong and reddish fluorescence from the x- and y-directions (Fig. 6d). One thing should be noted that SQ2 was detectable at the whole z-direction, indicating the translocation of SQ2 from the leaf mesophyll into the epidermis. To further investigate the translocation pathway of SQ2, fluorescence intensity analysis of the transverse tissue sections of the leaf was conducted. Fig. 6e illustrates the distribution histograms generated from the fluorescence intensity in the selected zone. The vein widths were evaluated to be ∼30 μm at the centre depth. Bright fluorescence was found in different regions of the tissue section, including the spongy mesophyll and palisade mesophyll of the leaf. Through the converse color difference, a vein close to the middle of the leaf could also be identified.
In the 3D images of the root incubated with SQ2, the orthoslice images at the selected depths indicated strong and reddish fluorescence from the x- and y-directions (Fig. 7a and d). We know that the apoplastic vascular system, functioning as a microfluidic network throughout the plant, consists of the xylem and phloem tissues.46 A diagram of the Arabidopsis root architecture and cell landscape indicated the strong staining of SQ2 to the cortex due to its lignin specificity, not found in the epidermis and xylem zones (Fig. 7c). Fig. 7e illustrates the distribution histograms generated from the fluorescence intensity in the selected zone. The xylem widths were evaluated to be ∼6.8 μm, and the root cortex varied from 20.7 to 25.7 μm along with the centre penetration depths.
3D reconstruction of the SQ3 staining revealed the deep penetration ∼108 μm in the leaves (Fig. S21, ESI†). Following different pathways, hydrophilic SQ3 was extensively absorbed by Arabidopsis roots, transported to the Arabidopsis leaves through the vascular bundles, and accumulated mainly in the veins compared with the mesophyll system. Associated with the Arabidopsis root incubated with SQ3, the fluorescence focused on the primary xylem tissue (Fig. S22, ESI†). The comparative performance of the present three SQs for cellular imaging with the previously reported SQs illustrates that more desirable imaging results can be obtained by tuning the molecular structures (Table S2, ESI†). These results might also contribute to a better understanding of the accumulation mechanisms of plants to small molecules, indicating its potential in further biological applications.
Conclusions
In summary, three two-photon absorbing SQs were developed, and the regulation effects of different substituents were investigated comprehensively. The steady-state optical properties induced by the subtle structure differences were manifested properly. Utilizing the open-aperture Z-scan and 2PEF strategies, broadband 2PA within the NIR spectral window was obtained. Amphiphilic SQ2 and hydrophilic SQ3 exhibited impressive δ2PA values of ∼1060 and ∼1220 GM at 800 nm, respectively. Potential biological applications in plant cells were further evaluated based on the images of the CLSM and 2PFM. Though the SQs exhibited similar optical properties in solution media, apparent differences were observed in the cellular uptake due to their unique tissue affinity. Rapid uptake and uniform distribution of hydrophobic SQ1 in the cell walls were investigated systematically. At the cellular level, amphiphilic SQ2 had a special affinity to the cell vacuole and nucleus, in contrast to hydrophilic SQ3 for the cell vacuole. Deep tissue labelling and high-resolution 3D reconstruction in plant cells were further accompanied. We deem that this is the first demonstration of distinct labelling of plant cells induced by the subtle structure differences of the far-red-emitting SQs. The promising advantages of spatial resolution, sensitivity, and reduced endogenous autofluorescence highlight the capabilities of far-red-emitting SQs for 2PEF and live bioimaging applications at the subcellular level in plant systems.Author contributions
The manuscript was written through contributions of all authors. N. Zhang performed all experimental works and data analyses and wrote the original draft. H. Chang analysed the data. R. Miao and L. Ding analysed the data and corrected the manuscript. T. Liu analysed the data and wrote the manuscript. Y. Fang supervised the project and corrected the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Acknowledgements
The authors acknowledge the funding from the National Natural Science Foundation of China (22172097), the Innovation Capability Support Program of Shaanxi (2021TD-18), the 111 Project (B14041), and the China Agriculture Research System (CARS-27).References
- N. R. Scott, H. Chen and H. Cui, J. Agric. Food Chem., 2018, 66, 6451–6456 CrossRef CAS PubMed.
- J. Zhao, K. Liu, R. Wang, T. Liu, Z. Wu, L. Ding and Y. Fang, ACS Appl. Mater. Interfaces, 2022, 14, 53323–53330 CrossRef CAS PubMed.
- M. Mandrone, I. Chiocchio, L. Barbanti, P. Tomasi, M. Tacchini and F. Poli, J. Agric. Food Chem., 2021, 69, 1132–1145 CrossRef CAS PubMed.
- K. Khambatta, A. Hollings, G. Sauzier, L. M. V. P. Sanglard, A. R. Klein, M. J. Tobin, J. Vongsvivut, M. R. Gibberd, A. D. Payne, F. Naim and M. J. Hackett, Adv. Sci., 2021, 8, 2101902 CrossRef CAS PubMed.
- E. Voke, R. L. Pinals, N. S. Goh and M. P. Landry, ACS Sens., 2021, 6, 2802–2814 CrossRef CAS PubMed.
- A. Nicol, K. Wang, R. T. K. Kwok, Z. Song, N. Li and B. Z. Tang, ACS Appl. Mater. Interfaces, 2017, 9, 28298–28304 CrossRef CAS PubMed.
- S. Bourke, F. Donà, Y. Teijeiro Gonzalez, B. Qazi Chaudhry, M. Panamarova, E. Mackay, P. S. Zammit, L. A. Dailey, U. S. Eggert, K. Suhling and M. A. Green, ACS Appl. Polym. Mater., 2022, 4, 8193–8202 CrossRef CAS PubMed.
- S. Pascal, S. David, C. Andraud and O. Maury, Chem. Soc. Rev., 2021, 50, 6613–6658 RSC.
- V. J. Pansare, S. Hejazi, W. J. Faenza and R. K. Prud’homme, Chem. Mater., 2012, 24, 812–827 CrossRef CAS PubMed.
- M. Collot, R. Kreder, A. L. Tatarets, L. D. Patsenker, Y. Mely and A. S. Klymchenko, Chem. Commun., 2015, 51, 17136–17139 RSC.
- W. Feng, K. Liu, J. Zang, J. Xu, H. Peng, L. Ding, T. Liu and Y. Fang, J. Phys. Chem. B, 2021, 125, 11540–11547 CrossRef CAS PubMed.
- G. Wang, K.-Y. Pu, X. Zhang, K. Li, L. Wang, L. Cai, D. Ding, Y.-H. Lai and B. Liu, Chem. Mater., 2011, 23, 4428–4434 CrossRef CAS.
- N. Fan, P. Li, Y. Zhou, C. Wu, X. Wang, Z. Liu and B. Tang, ACS Sens., 2022, 7, 71–81 CrossRef CAS PubMed.
- M. V. Bondar, S. Faryadras, N. Munera, H.-J. Chang, M. Uddin, K. D. Belfield, O. D. Kachkovsky, E. W. Van Stryland and D. J. Hagan, J. Phys. Chem. B, 2022, 126, 3897–3907 CrossRef CAS PubMed.
- G. Xia and H. Wang, J. Photochem. Photobiol., C, 2017, 31, 84–113 CrossRef CAS.
- K. Ilina, W. M. MacCuaig, M. Laramie, J. N. Jeouty, L. R. McNally and M. Henary, Bioconjugate Chem., 2020, 31, 194–213 CrossRef CAS PubMed.
- P. Sun, Q. Wu, X. Sun, H. Miao, W. Deng, W. Zhang, Q. Fan and W. Huang, Chem. Commun., 2018, 54, 13395–13398 RSC.
- H.-J. Chang, M. V. Bondar, T. Liu, X. Liu, S. Singh, K. D. Belfield, A. Sheely, A. E. Masunov, D. J. Hagan and E. W. Van Stryland, ACS Omega, 2019, 4, 14669–14679 CrossRef CAS PubMed.
- S. Xiao, N. Fu, K. Peckham and B. D. Smith, Org. Lett., 2010, 12, 140–143 CrossRef CAS PubMed.
- D. Yang, L. Yang, Y. Huang, Y. Jiao, T. Igarashi, Y. Chen, Z. Lu, X. Pu, H. Sasabe and J. Kido, ACS Appl. Mater. Interfaces, 2015, 7, 13675–13684 CrossRef CAS PubMed.
- S. Sreejith, J. Joseph, M. Lin, N. V. Menon, P. Borah, H. J. Ng, Y. X. Loong, Y. Kang, S. W.-K. Yu and Y. Zhao, ACS Nano, 2015, 9, 5695–5704 CrossRef CAS PubMed.
- T. Liu, X. Liu, W. Wang, Z. Luo, M. Liu, S. Zou, C. Sissa, A. Painelli, Y. Zhang, M. Vengris, M. V. Bondar, D. J. Hagan, E. W. Van Stryland, Y. Fang and K. D. Belfield, J. Phys. Chem. C, 2018, 122, 3994–4008 CrossRef CAS.
- H. M. Kim and B. R. Cho, Chem. Rev., 2015, 115, 5014–5055 CrossRef CAS PubMed.
- L. Guo and M. S. Wong, Adv. Mater., 2014, 26, 5400–5428 CrossRef CAS PubMed.
- J. Karpenko, A. S. Klymchenko, S. Gioria, R. Kreder, I. Shulov, P. Villa, Y. Mély, M. Hibert and D. Bonnet, Chem. Commun., 2015, 51, 2960–2963 RSC.
- K. T. Fam, M. Collot and A. S. Klymchenko, Chem. Sci., 2020, 11, 8240–8248 RSC.
- T. S. Jarvis, F. M. Roland, K. M. Dubiak, P. W. Huber and B. D. Smith, J. Mater. Chem. B, 2018, 6, 4963–4971 RSC.
- G. Renno, F. Cardano, V. Ilieva, G. Viscardi and A. Fin, Eur. J. Org. Chem., 2022, e202200833 CrossRef CAS.
- K. T. Arun and D. Ramaiah, J. Phys. Chem. A, 2005, 109, 5571–5578 CrossRef CAS PubMed.
- T. Liu, X. Liu, Y. Zhang, M. V. Bondar, Y. Fang and K. D. Belfield, Eur. J. Org. Chem., 2018, 4095–4102 CrossRef CAS.
- D. Yao, Y. Wang, R. Zou, K. Bian, P. Liu, S. Shen, W. Yang, B. Zhang and D. Wang, ACS Appl. Mater. Interfaces, 2020, 12, 4276–4284 CrossRef CAS PubMed.
- Y. Zhang, X. Yue, B. Kim, S. Yao, M. V. Bondar and K. D. Belfield, ACS Appl. Mater. Interfaces, 2013, 5, 8710–8717 CrossRef CAS PubMed.
- C.-L. Sun, S.-K. Lv, Y.-P. Liu, Q. Liao, H.-L. Zhang, H. Fu and J. Yao, J. Mater. Chem. C, 2017, 5, 1224–1230 RSC.
- T. Liu, M. V. Bondar, K. D. Belfield, D. Anderson, A. E. Masunov, D. J. Hagan and E. W. Van Stryland, J. Phys. Chem. C, 2016, 120, 11099–11110 CrossRef CAS.
- N. Zhang, L. Liu, H. Chang, K. Liu, T. Liu, L. Ding and Y. Fang, J. Phys. Chem. Lett., 2023, 14, 7283–7289 CrossRef CAS PubMed.
- W. Feng, K. Liu, J. Zang, G. Wang, R. Miao, L. Ding, T. Liu, J. Kong and Y. Fang, ACS Appl. Mater. Interfaces, 2021, 13, 28985–28995 CrossRef CAS PubMed.
- L. Liu, S. Li, N. Zhang, Q. Shi, K. Liu, T. Liu, Z. Huang, L. Ding and Y. Fang, J. Phys. Chem. B, 2023, 127, 10171–10178 CrossRef CAS PubMed.
- E. D. Onal and K. Guven, J. Phys. Chem. C, 2017, 121, 684–690 CrossRef CAS.
- J. Zang, W. Feng, X. Chang, K. Liu, H. Peng, L. Ding, T. Liu and Y. Fang, Dyes Pigm., 2021, 193, 109487 CrossRef CAS.
- J. Wang, C. Li, Y. Cui, Q. Wang, J. Ye, J. Yang, Z. Liu, S. Zhang, Y. Fu and J. Xu, Nanoscale, 2023, 15, 11026–11037 RSC.
- L. Wang, Y. Zhang, Y. Han, Q. Zhang, Z. Wen, H. Li, S. Sun, X. Chen and Y. Xu, J. Mater. Chem. B, 2021, 9, 2001–2009 RSC.
- T. S. Jarvis, F. M. Roland, K. M. Dubiak, P. W. Huber and B. D. Smith, J. Mater. Chem. B, 2018, 6, 4963–4971 RSC.
- S. Guo, W. Deng, S. Wu, P. Sun and Y. Liu, J. Mater. Chem. B, 2023, 11, 4389–4395 RSC.
- X. Zeng, X. Ma, J. Dong, B. Li, S. Hua Liu, J. Yin and G.-F. Yang, Angew. Chem., Int. Ed., 2023, 62, e202312618 CrossRef CAS PubMed.
- J. Chen, B. Liu, Z. Yang, J. Qu, H. Xun, R. Dou, X. Gao and L. Wang, Environ. Sci.: Nano, 2018, 5, 2672–2685 RSC.
- G. Dufil, I. Bernacka-Wojcik, A. Armada-Moreira and E. Stavrinidou, Chem. Rev., 2022, 122, 4847–4883 CrossRef CAS PubMed.
- F. Chang, L. Kuang, C. Huang, W. Jane, Y. Hung, Y. C. Hsing and C. Mou, J. Mater. Chem. B, 2013, 1, 5279–5287 RSC.
- W. Li, Y. Zheng, H. Zhang, Z. Liu, W. Su, S. Chen, Y. Liu, J. Zhuang and B. Lei, ACS Appl. Mater. Interfaces, 2016, 8, 19939–19945 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Synthesis procedures, characterization details, optical spectra, microscopy images, and NMR and HRMS spectra. See DOI: https://doi.org/10.1039/d4tb00400k |
| This journal is © The Royal Society of Chemistry 2024 |