Themed collection Droplet-based single-cell sequencing

Single cell sequencing: An engineer and business person's perspective on how we got here
Microfluidics entrepreneur Mark Gilligan provides a perspective on the development of single-cell sequencing technologies.

Lab Chip, 2017,17, 3349-3350
https://doi.org/10.1039/C7LC90095C
InDrops and Drop-seq technologies for single-cell sequencing
Droplet-based single-cell sequencing pioneers Allon Klein and Evan Macosko summarise the growth of the technology and look ahead to future developments.

Lab Chip, 2017,17, 2540-2541
https://doi.org/10.1039/C7LC90070H
Perspective on droplet-based single-cell sequencing
Thought leader David Weitz introduces the Lab on a Chip droplet-based single-cell sequencing thematic collection.

Lab Chip, 2017,17, 2539-2539
https://doi.org/10.1039/C7LC90069D
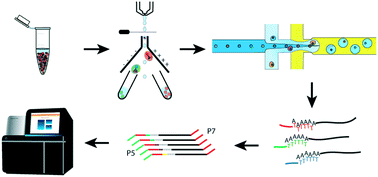
Droplet-based single cell RNAseq tools: a practical guide
A step-by-step guide for droplet-based single cell RNAseq experiments, practical considerations and technical notes.
Lab Chip, 2019,19, 1706-1727
https://doi.org/10.1039/C8LC01239C
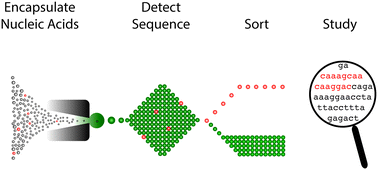
Finding a helix in a haystack: nucleic acid cytometry with droplet microfluidics
Nucleic acid cytometry using droplet microfluidics identifies and sorts single cells, virus, or free molecules based on specific “keyword” sequences.
Lab Chip, 2017,17, 2032-2045
https://doi.org/10.1039/C7LC00241F
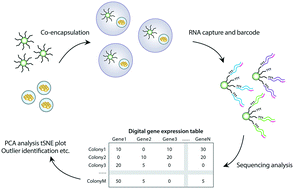
High throughput gene expression profiling of yeast colonies with microgel-culture Drop-seq
We describe isogenic colony sequencing (ICO-seq), a massively-parallel strategy to assess the gene expression profiles of large numbers of genetically distinct yeast colonies.
Lab Chip, 2019,19, 1838-1849
https://doi.org/10.1039/C9LC00084D
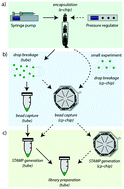
Simplified Drop-seq workflow with minimized bead loss using a bead capture and processing microfluidic chip
Single-cell RNA-sequencing (scRNA-seq) has revolutionized biomedical research by enabling the in-depth analysis of cell-to-cell heterogeneity of tissues with unprecedented resolution.
Lab Chip, 2019,19, 1610-1620
https://doi.org/10.1039/C9LC00014C
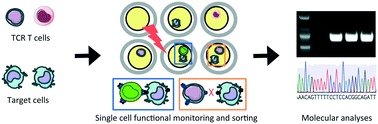
Functional TCR T cell screening using single-cell droplet microfluidics
Droplet-based single cell platform allows functional screening and sorting of desirable TCR T cells to accelerate development of adoptive T cell therapies.
Lab Chip, 2018,18, 3733-3749
https://doi.org/10.1039/C8LC00818C
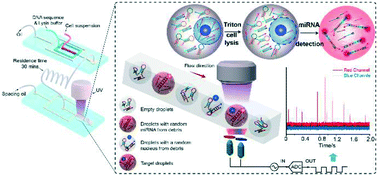
Ultrahigh-throughput droplet microfluidic device for single-cell miRNA detection with isothermal amplification
An ultrahigh-throughput single-cell miRNA assay is developed by a continuous-flow microfluidic process employing isothermal amplification to amplify the target miRNA signal.
Lab Chip, 2018,18, 1914-1920
https://doi.org/10.1039/C8LC00390D
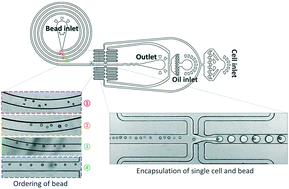
Inertial-ordering-assisted droplet microfluidics for high-throughput single-cell RNA-sequencing
We developed a modified high-throughput droplet barcoding technique for single-cell Drop-Seq via introduction of hydrodynamic ordering in a spiral microchannel.
Lab Chip, 2018,18, 775-784
https://doi.org/10.1039/C7LC01284E
About this collection
The field of droplet-based single-cell sequencing field has made increasing advances in recent years. Large numbers of studies are underway to collect and explore the new information that is now accessible with single-cell RNA-seq. Improvements to microfluidics are advancing rapidly and extensions to other sequencing methods are also being developed, enabling investigations to probe information beyond mRNA alone. This has rapidly become a burgeoning field, where microfluidic techniques are essential and where droplet-based microfluidics has enabled a major advance. The collection highlights research focussing on the interface between the technological advancements and high impact applications of droplet-based single-cell sequencing. This on-going collection is collated by Thought Leader David Weitz and the Lab on a Chip Editorial Office.
More details about the collection and how to apply can be found at rsc.li/drop-sc-seq-blog