Themed collection Integrative Approaches for Signalling and Metabolic Networks

Front cover

Inside front cover

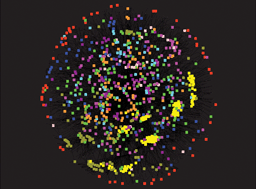
Integrative approaches for signalling and metabolic networks
Guest editors Vassily Hatzimanikatis and Julio Saez-Rodriguez introduce the Integrative Approaches for Signalling and Metabolic Networks themed issue of Integrative Biology.
Integr. Biol., 2015,7, 844-845
https://doi.org/10.1039/C5IB90030A
Genome scale models of yeast: towards standardized evaluation and consistent omic integration
We review genome scale models of yeast, how are they typically evaluated, and how can they be integrated with omic data.

Integr. Biol., 2015,7, 846-858
https://doi.org/10.1039/C5IB00083A
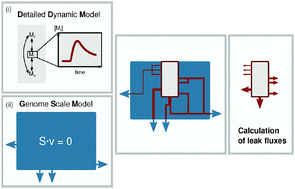
Dynamic metabolic models in context: biomass backtracking
Biomass backtracking supports the integration of dynamic metabolic models into a genome scale metabolic model and provides exact drain reactions.
Integr. Biol., 2015,7, 940-951
https://doi.org/10.1039/C5IB00050E
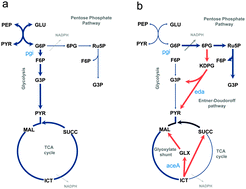
Sub-optimal phenotypes of double-knockout mutants of Escherichia coli depend on the order of gene deletions
Metabolic properties of the double-knockout strain E. coli ΔaceA Δpgi are shown to depend on the order of gene inactivation.
Integr. Biol., 2015,7, 930-939
https://doi.org/10.1039/C5IB00096C
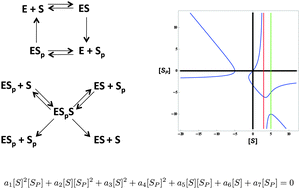
Invariants reveal multiple forms of robustness in bifunctional enzyme systems
High-throughput algebraic calculations reveal that bifunctional enzymes can implement many forms of robust control.
Integr. Biol., 2015,7, 883-894
https://doi.org/10.1039/C5IB00009B
Investigating Moorella thermoacetica metabolism with a genome-scale constraint-based metabolic model
Moorella thermoacetica is a strictly anaerobic, endospore-forming, and metabolically versatile acetogenic bacterium capable of conserving energy by both autotrophic (acetogenesis) and heterotrophic (homoacetogenesis) modes of metabolism.
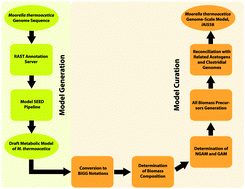
Integr. Biol., 2015,7, 869-882
https://doi.org/10.1039/C5IB00095E
Identification of drug-specific pathways based on gene expression data: application to drug induced lung injury
An Integer Linear Programming (ILP) formulation is introduced to model the modes of action of lung toxic drugs based on gene expression data and prior knowledge of protein connectivity.
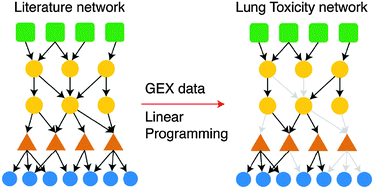
Integr. Biol., 2015,7, 904-920
https://doi.org/10.1039/C4IB00294F
Predicting genetic interactions from Boolean models of biological networks
The network representation of the cell fate decision model (Calzone et al., 2010) is used to generate a genetic interaction network for the apoptosis phenotype. Most genetic interactions are epistatic, single nonmonotonic, and additive (Drees et al., 2005).
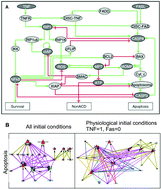
Integr. Biol., 2015,7, 921-929
https://doi.org/10.1039/C5IB00029G
An entropy-like index of bifurcational robustness for metabolic systems
Natural and synthetic metabolic pathways need to retain stability when faced against random changes in gene expression levels and kinetic parameters.
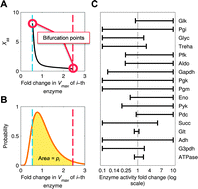
Integr. Biol., 2015,7, 895-903
https://doi.org/10.1039/C4IB00257A
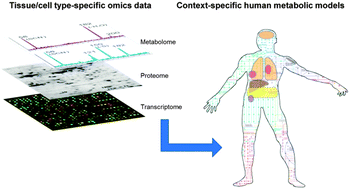
Reconstruction of genome-scale human metabolic models using omics data
Genome-scale metabolic models of human cells and tissues can be reconstructed using omics data for systematic and personalized medicine.
Integr. Biol., 2015,7, 859-868
https://doi.org/10.1039/C5IB00002E
About this collection
Living cells are composed of chemical and biophysical processes that are integrated into networks. The complex behaviour of life emerges from the function of these networks. Signal transduction and metabolism are two central cellular processes and their function is important for almost every problem in basic biology, medical and biotechnological applications. The study of these processes in the context of their network structure and function lies at the heart of integrative biology. This themed issue guest edited by Vassily Hatzimanikatis and Julio Saez-Rodriguez features original research and review articles on the experimental study and computational analysis of signal transduction and metabolism.