Discovering protein–protein interaction stabilisers by native mass spectrometry†
Abstract
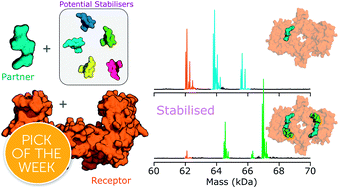
Protein–protein interactions (PPIs) are key therapeutic targets. Most PPI-targeting drugs in the clinic inhibit these important interactions; however, stabilising PPIs is an attractive alternative in cases where a PPI is disrupted in a disease state. The discovery of novel PPI stabilisers has been hindered due to the lack of tools available to monitor PPI stabilisation. Moreover, for PPI stabilisation to be detected, both the stoichiometry of binding and the shift this has on the binding equilibria need to be monitored simultaneously. Here, we show the power of native mass spectrometry (MS) in the rapid search for PPI stabilisers. To demonstrate its capability, we focussed on three PPIs between the eukaryotic regulatory protein 14-3-3σ and its binding partners estrogen receptor ERα, the tumour suppressor p53, and the kinase LRRK2, whose interactions upon the addition of a small molecule, fusicoccin A, are differentially stabilised. Within a single measurement the stoichiometry and binding equilibria between 14-3-3 and each of its binding partners was evident. Upon addition of the fusicoccin A stabiliser, a dramatic shift in binding equilibria was observed with the 14-3-3:ERα complex compared with the 14-3-3:p53 and 14-3-3:LRRK2 complexes. Our results highlight how native MS can not only distinguish the ability of stabilisers to modulate PPIs, but also give important insights into the dynamics of ternary complex formation. Finally, we show how native MS can be used as a screening tool to search for PPI stabilisers, highlighting its potential role as a primary screening technology in the hunt for novel therapeutic PPI stabilisers.
- This article is part of the themed collections: Most popular 2021 chemical biology articles, 2021 ChemSci Pick of the Week Collection and 2021 Chemical Science HOT Article Collection


 Please wait while we load your content...
Please wait while we load your content...