PoseBusters: AI-based docking methods fail to generate physically valid poses or generalise to novel sequences†
Abstract
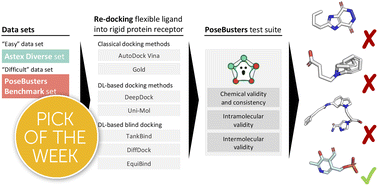
The last few years have seen the development of numerous deep learning-based protein–ligand docking methods. They offer huge promise in terms of speed and accuracy. However, despite claims of state-of-the-art performance in terms of crystallographic root-mean-square deviation (RMSD), upon closer inspection, it has become apparent that they often produce physically implausible molecular structures. It is therefore not sufficient to evaluate these methods solely by RMSD to a native binding mode. It is vital, particularly for deep learning-based methods, that they are also evaluated on steric and energetic criteria. We present PoseBusters, a Python package that performs a series of standard quality checks using the well-established cheminformatics toolkit RDKit. The PoseBusters test suite validates chemical and geometric consistency of a ligand including its stereochemistry, and the physical plausibility of intra- and intermolecular measurements such as the planarity of aromatic rings, standard bond lengths, and protein–ligand clashes. Only methods that both pass these checks and predict native-like binding modes should be classed as having “state-of-the-art” performance. We use PoseBusters to compare five deep learning-based docking methods (DeepDock, DiffDock, EquiBind, TankBind, and Uni-Mol) and two well-established standard docking methods (AutoDock Vina and CCDC Gold) with and without an additional post-prediction energy minimisation step using a molecular mechanics force field. We show that both in terms of physical plausibility and the ability to generalise to examples that are distinct from the training data, no deep learning-based method yet outperforms classical docking tools. In addition, we find that molecular mechanics force fields contain docking-relevant physics missing from deep-learning methods. PoseBusters allows practitioners to assess docking and molecular generation methods and may inspire new inductive biases still required to improve deep learning-based methods, which will help drive the development of more accurate and more realistic predictions.
- This article is part of the themed collections: 2024 Chemical Science HOT Article Collection, 2024 ChemSci Pick of the Week Collection and Most popular 2024 physical, theoretical and computational chemistry articles


 Please wait while we load your content...
Please wait while we load your content...