Optimisation of 1H PMLG homonuclear decoupling at 60 kHz MAS to enable 15N–1H through-bond heteronuclear correlation solid-state NMR spectroscopy†‡
Abstract
The Lee–Goldburg condition for homonuclear decoupling in 1H magic-angle spinning (MAS) solid-state NMR sets the angle θ, corresponding to arctan of the ratio of the rf nutation frequency, ν1, to the rf offset, to be the magic angle, θm, equal to tan−1(√2) = 54.7°. At 60 kHz MAS, we report enhanced decoupling compared to MAS alone in a 1H spectrum of 15N-glycine with 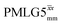 at θ = 30° for a ν1 of ∼100 kHz at a 1H Larmor frequency, ν0, of 500 MHz and 1 GHz, corresponding to a high chemical shift scaling factor (λCS) of 0.82. At 1 GHz, we also demonstrate enhanced decoupling compared to 60 kHz MAS alone for a lower ν1 of 51 kHz, i.e., a case where the nutation frequency is less than the MAS frequency, with θ = 18°, λCS = 0.92. The ratio of the rotor period to the decoupling cycle time, Ψ = τr/τc, is in the range 0.53 to 0.61. Windowed
at θ = 30° for a ν1 of ∼100 kHz at a 1H Larmor frequency, ν0, of 500 MHz and 1 GHz, corresponding to a high chemical shift scaling factor (λCS) of 0.82. At 1 GHz, we also demonstrate enhanced decoupling compared to 60 kHz MAS alone for a lower ν1 of 51 kHz, i.e., a case where the nutation frequency is less than the MAS frequency, with θ = 18°, λCS = 0.92. The ratio of the rotor period to the decoupling cycle time, Ψ = τr/τc, is in the range 0.53 to 0.61. Windowed 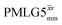 decoupling using the optimised parameters for a ν1 of ∼100 kHz also gives good performance in a 1H spin-echo experiment, enabling implementation in a 1H-detected 15N–1H cross polarisation (CP)-refocused INEPT heteronuclear correlation NMR experiment. Specifically, initial 15N transverse magnetisation as generated by 1H–15N CP is transferred back to 1H using a refocused INEPT pulse sequence employing windowed
decoupling using the optimised parameters for a ν1 of ∼100 kHz also gives good performance in a 1H spin-echo experiment, enabling implementation in a 1H-detected 15N–1H cross polarisation (CP)-refocused INEPT heteronuclear correlation NMR experiment. Specifically, initial 15N transverse magnetisation as generated by 1H–15N CP is transferred back to 1H using a refocused INEPT pulse sequence employing windowed 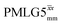 1H decoupling. Such an approach ensures the observation of through-bond N–H connectivities. For 15N-glycine, while the CP-refocused INEPT experiment has a lower sensitivity (∼50%) as compared to a double CP experiment (with a 200 μs 15N to 1H CP contact time), there is selectivity for the directly bonded NH3+ moiety, while intensity is observed for the CH21H resonances in the double CP experiment. Two-dimensional 15N–1H correlation MAS NMR spectra are presented for the dipeptide β-AspAla and the pharmaceutical cimetidine at 60 kHz MAS, both at natural isotopic abundance. For the dipeptide β-AspAla, different build-up dependence on the first spin-echo duration is observed for the NH and NH3+ moieties demonstrating that the experiment could be used to distinguish resonances for different NHx groups.
1H decoupling. Such an approach ensures the observation of through-bond N–H connectivities. For 15N-glycine, while the CP-refocused INEPT experiment has a lower sensitivity (∼50%) as compared to a double CP experiment (with a 200 μs 15N to 1H CP contact time), there is selectivity for the directly bonded NH3+ moiety, while intensity is observed for the CH21H resonances in the double CP experiment. Two-dimensional 15N–1H correlation MAS NMR spectra are presented for the dipeptide β-AspAla and the pharmaceutical cimetidine at 60 kHz MAS, both at natural isotopic abundance. For the dipeptide β-AspAla, different build-up dependence on the first spin-echo duration is observed for the NH and NH3+ moieties demonstrating that the experiment could be used to distinguish resonances for different NHx groups.


 Please wait while we load your content...
Please wait while we load your content...