OligoTRAFTACs: A generalizable method for transcription factor degradation†
Abstract
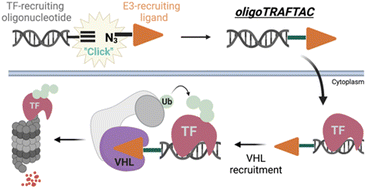
Dysregulated transcription factors (TFs) that rewire gene expression circuitry are frequently identified as key players in disease. Although several TFs have been drugged with small molecules, the majority of oncogenic TFs are not currently pharmaceutically tractable due to their paucity of ligandable pockets. The first generation of transcription factor targeting chimeras (TRAFTACs) was developed to target TFs for proteasomal degradation by exploiting their DNA binding ability. In the current study, we have developed the second generation TRAFTACs (“oligoTRAFTACs”) composed of a TF-binding oligonucleotide and an E3 ligase-recruiting ligand. Herein, we demonstrate the development of oligoTRAFTACs to induce the degradation of two oncogenic TFs, c-Myc and brachyury. In addition, we show that brachyury can be successfully degraded by oligoTRAFTACs in chordoma cell lines. Furthermore, zebrafish experiments demonstrate in vivo oligoTRAFTAC activity. Overall, our data demonstrate oligoTRAFTACs as a generalizable platform towards difficult-to-drug TFs and their degradability via the proteasomal pathway.


 Please wait while we load your content...
Please wait while we load your content...