Efficient synthesis and replication of diverse sequence libraries composed of biostable nucleic acid analogues†
Abstract
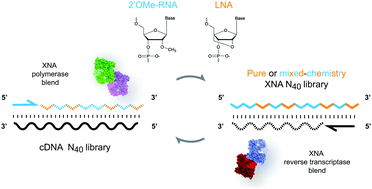
Functional nucleic acids can be evolved in vitro using cycles of selection and amplification, starting from diverse-sequence libraries, which are typically restricted to natural or partially-modified polymer chemistries. Here, we describe the efficient DNA-templated synthesis and reverse transcription of libraries entirely composed of serum nuclease resistant alternative nucleic acid chemistries validated in nucleic acid therapeutics; locked nucleic acid (LNA), 2′-O-methyl-RNA (2′OMe-RNA), or mixtures of the two. We evaluate yield and diversity of synthesised libraries and measure the aggregate error rate of a selection cycle. We find that in addition to pure 2′-O-methyl-RNA and LNA, several 2′OMe-RNA/LNA blends seem suitable and promising for discovery of biostable functional nucleic acids for biomedical applications.
- This article is part of the themed collection: Themed collection on XNA Xeno-nucleic acids


 Please wait while we load your content...
Please wait while we load your content...