Integrated proteomics and phosphoproteomics reveal perturbed regulative pathways in pancreatic ductal adenocarcinoma†
Abstract
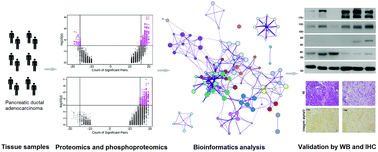
Pancreatic ductal adenocarcinoma (PDAC) has a dismal prognosis largely owing to its inefficient diagnosis, rapid progress, and tenacious drug resistance. Here, we aimed to analyze the expressive patterns of proteins and phosphorylation in PDAC tissue samples and compare them to normal pancreatic tissue to investigate the underlying mechanisms and to reveal potential protein targets for diagnosis and drug development. Liquid chromatography coupled to mass spectrometry (LC-MS) based proteomics and phosphoproteomics analyses were performed using 20 pairs of patient-derived PDAC tissue and normal pancreatic tissue samples. Protein identification and quantification were conducted using MaxQuant software. Bioinformatics analysis was used to retrieve PDAC-relevant pathways and gene ontology (GO) terms. 4985 proteins and 3643 phosphoproteins were identified with high confidence; of these, 322 proteins and 235 phosphoproteins were dysregulated in PDAC. Several pathways, including several extracellular matrix-related pathways, were found to be strongly associated with PDAC. Further, the expression levels of filamin A (FLNA), integrin alpha-V (ITGAV), thymidine phosphorylase (TYMP), medium-chain specific acyl-CoA dehydrogenase, mitochondrial (ACADM), short-chain specific acyl-CoA dehydrogenase, mitochondrial (ACADS), and acetyl-CoA acetyltransferase, mitochondrial (ACAT1) were examined through western blot and immunohistochemistry analysis, and the results confirmed the validity of the proteomics analysis results. These findings provide comprehensive insight into the dysregulated regulative networks in PDAC tissue samples at the protein and phosphorylation levels, and they provide clues for further pathological studies and drug-target development.


 Please wait while we load your content...
Please wait while we load your content...