Characterization of DNA aptamer–protein binding using fluorescence anisotropy assays in low-volume, high-efficiency plates†
Abstract
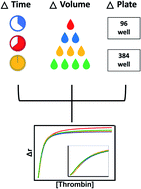
Aptamers have many useful attributes including specific binding to molecular targets. After aptamers are identified, their target binding must be characterized. Fluorescence anisotropy (FA) is one technique that can be used to characterize affinity and to optimize aptamer–target interactions. Efforts to make FA assays more efficient by reducing assay volume and time from mixing to measurement may save time and resources by minimizing consumption of costly reagents. Here, we use thrombin and two thrombin-binding aptamers as a model system to show that plate-based FA experiments can be performed in volumes as low as 2 μL per well with 20 minute incubations with minimal loss in assay precision. We demonstrate that the aptamer–thrombin interaction is best modelled with the Hill equation, indicating cooperative binding. The miniaturization of this assay has implications in drug development, as well as in the efficiency of aptamer selection workflows by allowing for higher throughput aptamer analysis.


 Please wait while we load your content...
Please wait while we load your content...