Automated affinity selection for rapid discovery of peptide binders†
Abstract
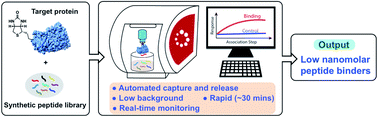
In-solution affinity selection (AS) of large synthetic peptide libraries affords identification of binders to protein targets through access to an expanded chemical space. Standard affinity selection methods, however, can be time-consuming, low-throughput, or provide hits that display low selectivity to the target. Here we report an automated bio-layer interferometry (BLI)-assisted affinity selection platform. When coupled with tandem mass spectrometry (MS), this method enables both rapid de novo discovery and affinity maturation of known peptide binders with high selectivity. The BLI-assisted AS-MS technology also features real-time monitoring of the peptide binding during the library selection process, a feature unattainable by current selection approaches. We show the utility of the BLI AS-MS platform toward rapid identification of novel nanomolar (dissociation constant, KD < 50 nM) non-canonical binders to the leukemia-associated oncogenic protein menin. To our knowledge, this is the first application of BLI to the affinity selection of synthetic peptide libraries. We believe our approach can significantly accelerate the use of synthetic peptidomimetic libraries in drug discovery.


 Please wait while we load your content...
Please wait while we load your content...