Regulation of cell binding and entry by DNA origami mediated spatial distribution of aptamers†
Abstract
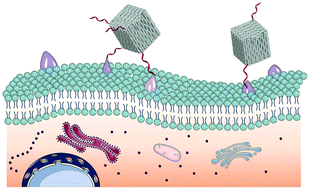
Understanding the effects of surface density and distribution of ligands on their recognition and binding is critical for the regulation of cellular behaviors. However, the correlation of spatial distribution of ligands particularly with cell binding and subsequent entry has been rarely explored. Here, we describe the use of DNA origami mediated spatial distribution of aptamers to regulate receptor ligand binding. Aptamers with tunable yet accurate density and orientation are anchored by virtue of the convenience and precision of DNA origami nanoboxes (DONs) to tailor their attachments. Cell assays demonstrate that the binding of DONs depends on both the density and orientation of aptamers, in which two adjacent aptamers exhibit the highest cellular uptake. The spatial distribution dependent uptake is further validated by utilizing two human cancer cell lines expressed with different levels of membrane receptors. Additionally, anticancer doxorubicin loaded DONs show internalization dependent proliferation inhibition of tumor cells. DNA origami mediated spatial distribution of ligands not only provides a unique method to tune cellular behaviors, but also offers new insights for the optimization of targeted drug delivery for cancer treatment.
- This article is part of the themed collection: Journal of Materials Chemistry B Emerging Investigators


 Please wait while we load your content...
Please wait while we load your content...