Functional protein shells fabricated from the self-assembling protein sheets of prokaryotic organelles†
Abstract
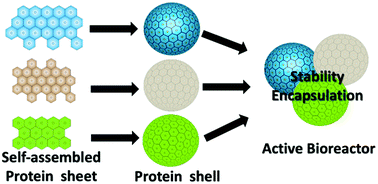
Fabricating protein compartments from protein units is challenging and limited by the use of external stimuli and crosslinkers. Here we explore the fabrication of all-protein compartments using self-assembled proteins of prokaryotic organelles. These proteins have intrinsic interacting domains which are ionic in nature, and spontaneously self-assemble into sheets when over-expressed. Using a one-step approach, we maneuvered the formation of the protein shells from the sheets without any external stimuli or crosslinker. The spontaneous self-assembly of the native protein sheets into protein shells not only preserves the native functional properties of the protein but also enhances their thermal stability compared to the sheets. We further demonstrate that these compartments can encapsulate macromolecular enzymes and, more interestingly, permit the free exchange of small molecules and substrates through their intrinsic conduit channels. The porous nature of the shell housing active enzymes and allowing movement of small molecules makes them suitable as active bioreactors. Furthermore, to extend the tunability of these protein-compartments with respect to stability, enzyme-encapsulation, and permeability, we fabricated three different compartments using three different sheet proteins, PduA/B/B′ and compared their properties. Interestingly we find that all three protein shells show similar behaviour with respect to an encapsulated diol-dehydratase enzyme and vitamin B12, which are native to the Pdu BMC system. Furthermore, for the non-native enzyme CytC, the small molecule R6G dye, doxorubicin, NR and curcumin they behave diversely. Insights from this analysis will allow us to design and develop sheet protein based synthetic active bioreactors requiring meticulous, compartmentalization in process optimization.


 Please wait while we load your content...
Please wait while we load your content...