Liquid crystal ordering of nucleic acids†
Abstract
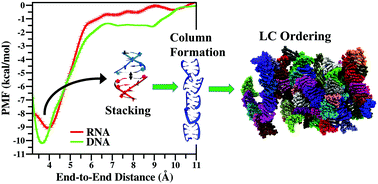
Several analytical calculations and computer simulations propose that cylindrical monodispersive rods having an aspect ratio (ratio of length to diameter) greater than 4 can exhibit liquid crystal (LC) ordering. But, recent experiments demonstrated the signature of LC ordering in systems of 4- to 20-base pair (bp) long nucleic acids (NAs) that do not satisfy the shape anisotropy criterion. Mechanisms of end-to-end adhesion and stacking have been proposed to explain this phenomenon. In this study, using all-atom molecular dynamics (MD) simulation, we explicitly verify the end-to-end stacking of double-stranded RNA (dsRNA) and demonstrate the LC ordering at the microscopic level. Using umbrella sampling (US) calculation, we quantify the potential of mean force (PMF) between two dsRNAs for various reaction coordinates (RCs) and compare our results with previously reported PMFs for double-stranded DNA (dsDNA). The PMF profiles demonstrate the anisotropic nature of inter-NA interaction. We find that, like dsDNA, dsRNA also prefers to stack on top of each other while repelling sideways, leading to the formation of supra-molecular-columns that undergo LC ordering at high NA volume fraction (ϕ). We also demonstrate and quantify the nematic ordering of the RNAs using several hundred nanosecond-long MD simulations that remain almost invariant for different initial configurations and under different external physiological conditions.


 Please wait while we load your content...
Please wait while we load your content...