Examining histone modification crosstalk using immobilized libraries established from ligation-ready nucleosomes†
Abstract
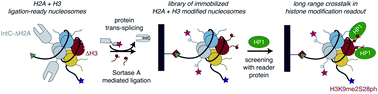
Chromatin signaling relies on a plethora of posttranslational modifications (PTM) of the histone proteins which package the long DNA molecules of our cells in reoccurring units of nucleosomes. Determining the biological function and molecular working mechanisms of different patterns of histone PTMs requires access to various chromatin substrates of defined modification status. Traditionally, these are achieved by individual reconstitution of single nucleosomes or arrays of nucleosomes in conjunction with modified histones produced by means of chemical biology. Here, we report an alternative strategy for establishing a library of differentially modified nucleosomes that bypasses the need for many individual syntheses, purification and assembly reactions by installing modified histone tails on ligation-ready, immobilized nucleosomes reconstituted in a single batch. Using the ligation-ready nucleosome strategy with sortase-mediated ligation for histone H3 and intein splicing for histone H2A, we generated libraries of up to 280 individually modified nucleosomes in 96-well plate format. Screening these libraries for the effects of patterns of PTMs onto the recruitment of a well-known chromatin factor, HP1 revealed a previously unknown long-range cross-talk between two modifications. H3S28 phosphorylation enhances recruitment of the HP1 protein to the H3K9 methylated H3-tail only in nucleosomal context. Detailed structural analysis by NMR measurements implies negative charges at position 28 to increase nucleosomal H3-tail dynamics and flexibility. Our work shows that ligation-ready nucleosomes enable unprecedented access to the ample space and complexity of histone modification patterns for the discovery and dissection of chromatin regulatory principles.


 Please wait while we load your content...
Please wait while we load your content...